|
|
|
|

|
| Sample: |
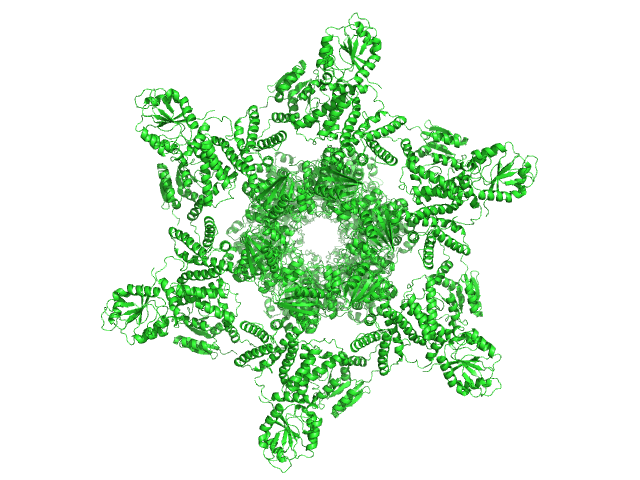
Circadian clock protein KaiA dodecamer, 393 kDa Synechococcus elongatus (strain … protein
Circadian clock protein KaiB hexamer, 71 kDa Synechococcus elongatus (strain … protein
Circadian clock protein kinase KaiC (S431D mutant) hexamer, 357 kDa Synechococcus elongatus (strain … protein
|
| Buffer: |
50 mM sodium phosphate buffer, 150 mM NaCl, 5 mM MgCl2, 0.5 mM EDTA, 1 mM ATP, 1 mM DTT, 50 mM arginine, 50 mM glutamine, pH: 7.8
|
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2017 Dec 8
|
Overall structure of fully assembled cyanobacterial KaiABC circadian clock complex by an integrated experimental-computational approach.
Commun Biol 5(1):184 (2022)
Yunoki Y, Matsumoto A, Morishima K, Martel A, Porcar L, Sato N, Yogo R, Tominaga T, Inoue R, Yagi-Utsumi M, Okuda A, Shimizu M, Urade R, Terauchi K, Kono H, Yagi H, Kato K, Sugiyama M
|
| RgGuinier |
7.0 |
nm |
| Dmax |
24.5 |
nm |
| VolumePorod |
1650 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
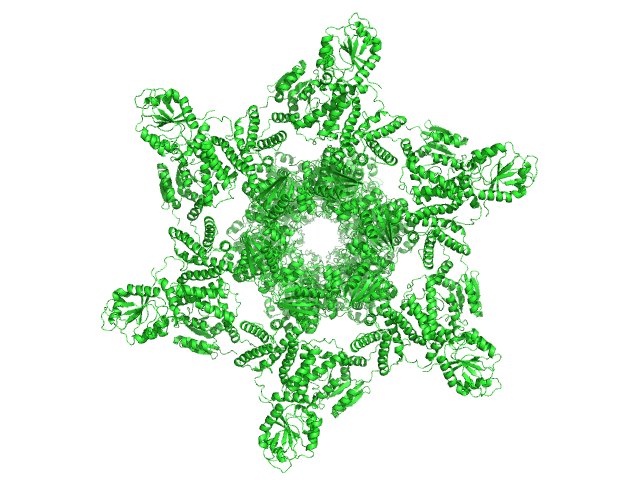
75% deuterated Circadian clock protein KaiB hexamer, 71 kDa Synechococcus elongatus (strain … protein
Circadian clock protein KaiA dodecamer, 393 kDa Synechococcus elongatus (strain … protein
75% deuterated Circadian clock protein kinase KaiC (S431D mutant) hexamer, 357 kDa Synechococcus elongatus (strain … protein
|
| Buffer: |
50 mM sodium phosphate buffer, 150 mm NaCl, 5 mM MgCl2, 0.5 mM EDTA, 1 mM ATP, 1 mM DTT, 50 mM arginine, 50 mM glutamine, in 100% D2O, pH: 7.8
|
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2018 Sep 19
|
Overall structure of fully assembled cyanobacterial KaiABC circadian clock complex by an integrated experimental-computational approach.
Commun Biol 5(1):184 (2022)
Yunoki Y, Matsumoto A, Morishima K, Martel A, Porcar L, Sato N, Yogo R, Tominaga T, Inoue R, Yagi-Utsumi M, Okuda A, Shimizu M, Urade R, Terauchi K, Kono H, Yagi H, Kato K, Sugiyama M
|
| RgGuinier |
7.8 |
nm |
| Dmax |
25.6 |
nm |
| VolumePorod |
1620 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
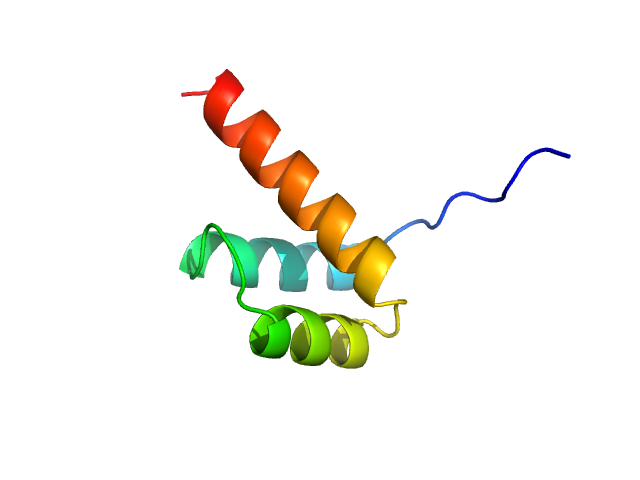
Cone-rod homeobox protein monomer, 7 kDa Homo sapiens protein
|
| Buffer: |
50 mM sodium phosphate,100 mM NaCl, and 5 mM imidazole, pH: 7
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Jun 15
|
Structural and functional analysis of the human Cone-rod homeobox transcription factor.
Proteins (2022)
Clanor PB, Buchholz C, Hayes JE, Friedman MA, White AM, Enke RA, Berndsen CE
|
| RgGuinier |
1.2 |
nm |
| Dmax |
4.0 |
nm |
| VolumePorod |
10 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
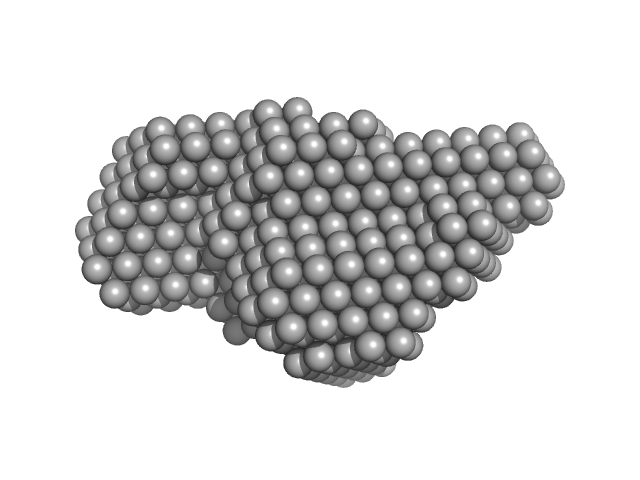
Accessory colonization factor monomer, 160 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM citrate-phosphate buffer, 200 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 27
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari...
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
244 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
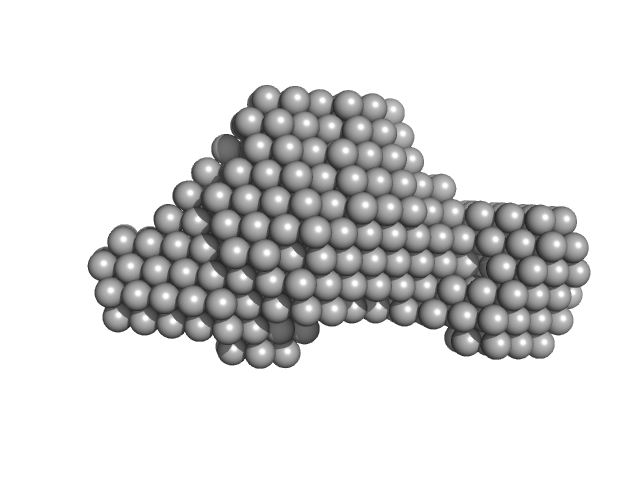
Accessory colonization factor monomer, 160 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM citrate-phosphate buffer, 200 mM NaCl, pH: 4.4
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 27
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari...
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
248 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
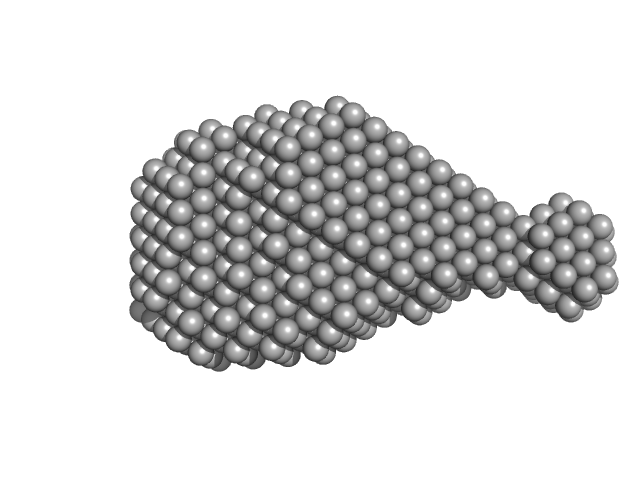
Accessory colonization factor monomer, 17 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 200 mM NaCl, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Apr 21
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari...
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
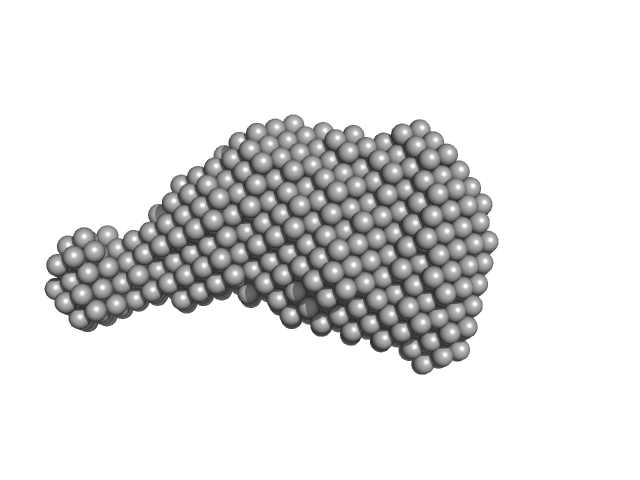
Accessory colonization factor monomer, 23 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 200 mM NaCl, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Apr 21
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari...
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
42 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
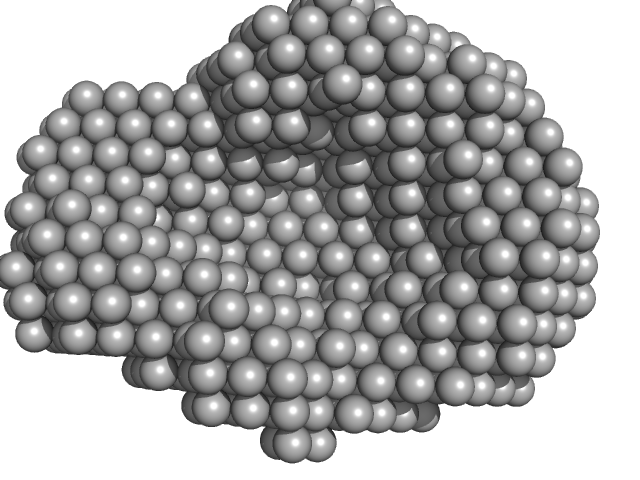
Accessory colonization factor monomer, 121 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 200 mM NaCl, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Apr 21
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari...
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
171 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Thioredoxin domain-containing protein trimer, 73 kDa Caulobacter vibrioides (strain … protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2019 Aug 21
|
The suppressor of copper sensitivity protein C from Caulobacter crescentus
is a trimeric disulfide isomerase that binds copper(I) with subpicomolar affinity
Acta Crystallographica Section D Structural Biology 78(3):337-352 (2022)
Petit G, Hong Y, Djoko K, Whitten A, Furlong E, McCoy A, Gulbis J, Totsika M, Martin J, Halili M
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
97 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
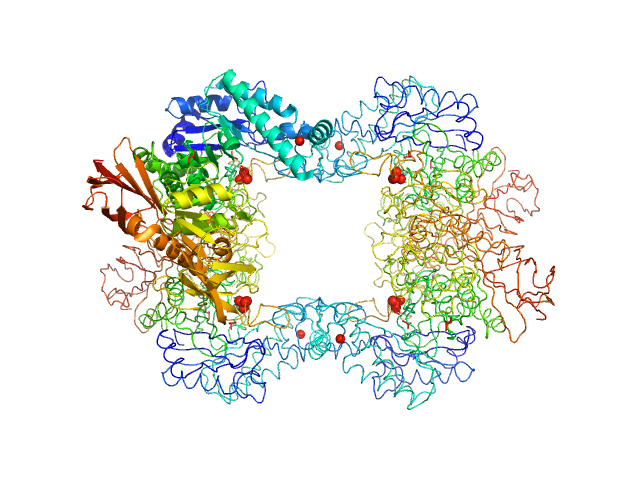
Alpha-L-fucosidase tetramer, 297 kDa Paenibacillus thiaminolyticus (Bacillus … protein
|
| Buffer: |
50 mM potassium phosphate, pH: 7.4
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2019 Jul 16
|
The first structure–function study of GH151 α‐ l
‐fucosidase uncovers new oligomerization pattern, active site complementation, and selective substrate specificity
The FEBS Journal (2022)
Koval'ová T, Kovaľ T, Stránský J, Kolenko P, Dušková J, Švecová L, Vodičková P, Spiwok V, Benešová E, Lipovová P, Dohnálek J
|
| RgGuinier |
4.2 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
418 |
nm3 |
|
|