|
|
|
|
|
| Sample: |
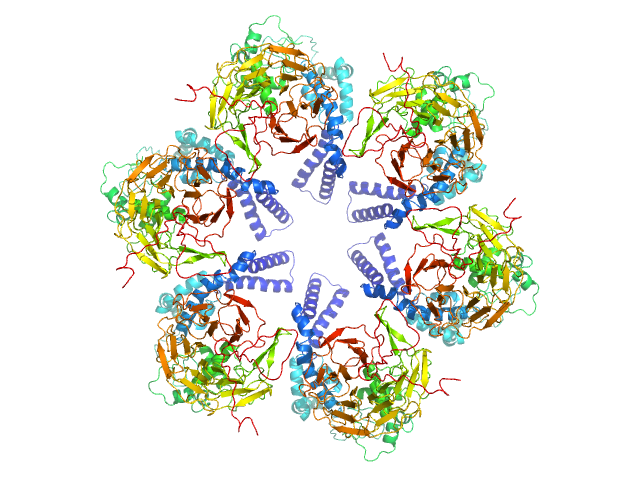
Kelch protein K13 hexamer, 396 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2020 Jul 25
|
Plasmodium falciparum Kelch13 and its artemisinin-resistant mutants assemble as hexamers in solution: a SAXS data driven modeling study.
FEBS J (2022)
Goel N, Dhiman K, Kalidas N, Mukhopadhyay A, Ashish F, Bhattacharjee S
|
| RgGuinier |
6.4 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
1630 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
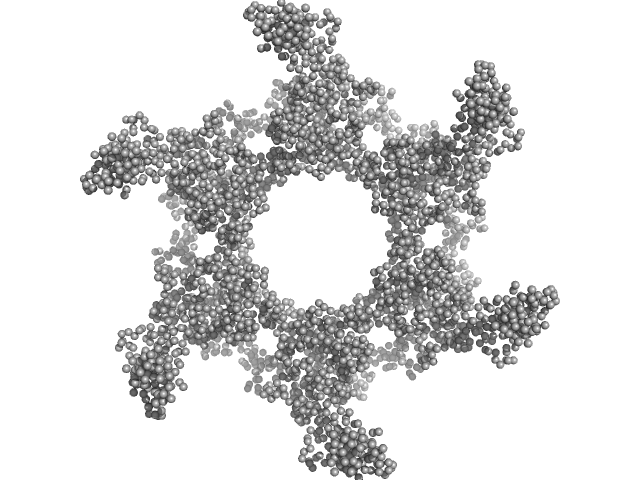
Kelch protein K13 (Truncated Kelch13-R539T, artemisinin-resistant mutation) hexamer, 396 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2020 Jul 25
|
Plasmodium falciparum Kelch13 and its artemisinin-resistant mutants assemble as hexamers in solution: a SAXS data driven modeling study.
FEBS J (2022)
Goel N, Dhiman K, Kalidas N, Mukhopadhyay A, Ashish F, Bhattacharjee S
|
| RgGuinier |
6.6 |
nm |
| Dmax |
20.5 |
nm |
| VolumePorod |
1600 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
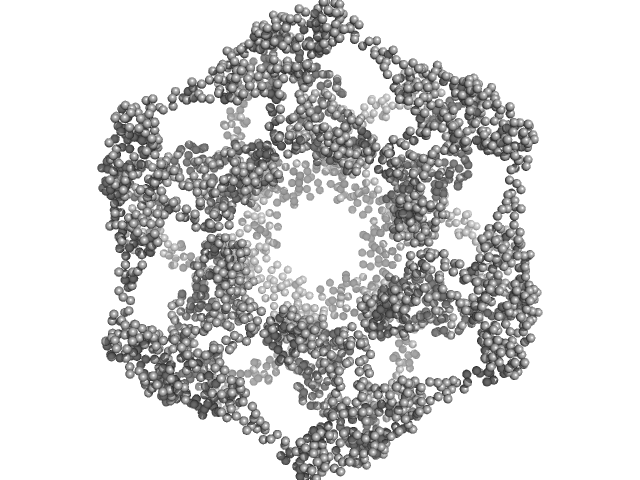
Kelch protein K13 (Truncated Kelch13-C580Y ) hexamer, 397 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
Phosphate Buffer Saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2020 Jul 28
|
Plasmodium falciparum Kelch13 and its artemisinin-resistant mutants assemble as hexamers in solution: a SAXS data driven modeling study.
FEBS J (2022)
Goel N, Dhiman K, Kalidas N, Mukhopadhyay A, Ashish F, Bhattacharjee S
|
| RgGuinier |
6.4 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
1800 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
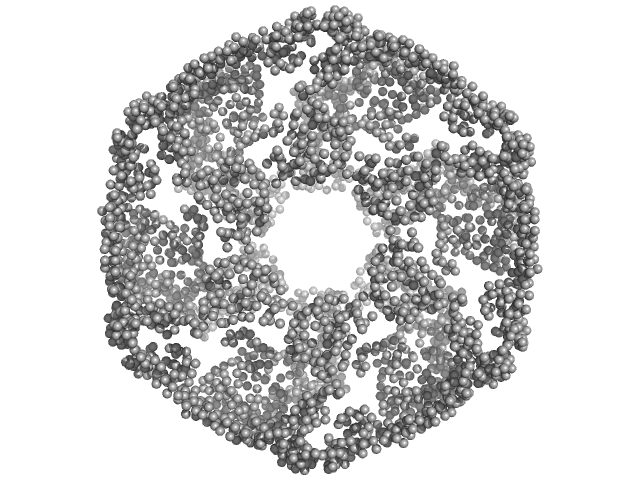
Kelch protein K13 (Truncated Kelch13-A578S) hexamer, 396 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
Phosphate Buffer Saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2020 Jul 29
|
Plasmodium falciparum Kelch13 and its artemisinin-resistant mutants assemble as hexamers in solution: a SAXS data driven modeling study.
FEBS J (2022)
Goel N, Dhiman K, Kalidas N, Mukhopadhyay A, Ashish F, Bhattacharjee S
|
| RgGuinier |
6.1 |
nm |
| Dmax |
16.8 |
nm |
| VolumePorod |
1700 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Arrestin-3 fused to extracellular signal-regulated kinase 2 monomer, 89 kDa Bos taurus/Rattus norvegicus protein
|
| Buffer: |
20 mM MOPS pH 7.5, 150 mM NaCl, 1 mM TCEP, and 5% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Mar 23
|
The two non-visual arrestins engage ERK2 differently.
J Mol Biol :167465 (2022)
Perry-Hauser NA, Bennett Hopkins J, Zhuo Y, Zheng C, Perez I, Schultz KM, Vishnivetskiy SA, Kaya AI, Sharma P, Dalby KN, Young Chung K, Klug CS, Gurevich VV, Iverson TM
|
| RgGuinier |
3.6 |
nm |
| Dmax |
14.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
Arrestin-3 fused to extracellular signal-regulated kinase 2 dimer, 178 kDa Bos taurus/Rattus norvegicus protein
|
| Buffer: |
20 mM MOPS pH 7.5, 150 mM NaCl, 1 mM TCEP, and 5% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Mar 23
|
The two non-visual arrestins engage ERK2 differently.
J Mol Biol :167465 (2022)
Perry-Hauser NA, Bennett Hopkins J, Zhuo Y, Zheng C, Perez I, Schultz KM, Vishnivetskiy SA, Kaya AI, Sharma P, Dalby KN, Young Chung K, Klug CS, Gurevich VV, Iverson TM
|
| RgGuinier |
4.3 |
nm |
| Dmax |
16.0 |
nm |
|
|
|
|
|
|
|
| Sample: |
Unconventional myosin-X component dimer, 15 kDa Homo sapiens protein
|
| Buffer: |
HEPES, 5% glycerol, 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Mar 16
|
K
-edge anomalous SAXS for protein solution structure modeling
Acta Crystallographica Section D Structural Biology 78(2) (2022)
Virk K, Yonezawa K, Choukate K, Singh L, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
24 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Unconventional myosin-X dimer, 15 kDa Homo sapiens protein
|
| Buffer: |
HEPES, 5% glycerol, 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Mar 16
|
K
-edge anomalous SAXS for protein solution structure modeling
Acta Crystallographica Section D Structural Biology 78(2) (2022)
Virk K, Yonezawa K, Choukate K, Singh L, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Unconventional myosin-X component dimer, 15 kDa Homo sapiens protein
|
| Buffer: |
HEPES, 5% glycerol, 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Mar 16
|
K
-edge anomalous SAXS for protein solution structure modeling
Acta Crystallographica Section D Structural Biology 78(2) (2022)
Virk K, Yonezawa K, Choukate K, Singh L, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Unconventional myosin-X component dimer, 15 kDa Homo sapiens protein
|
| Buffer: |
HEPES, 5% glycerol, 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at BL-15A2, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Mar 16
|
K
-edge anomalous SAXS for protein solution structure modeling
Acta Crystallographica Section D Structural Biology 78(2) (2022)
Virk K, Yonezawa K, Choukate K, Singh L, Shimizu N, Chaudhuri B
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|