|
|
|
|

|
| Sample: |
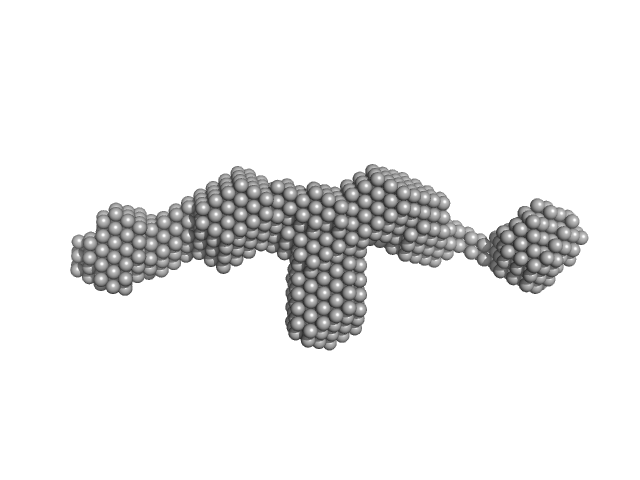
Fibrillin-1 PF3 monomer, 78 kDa Homo sapiens protein
|
| Buffer: |
Tris buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Dec 12
|
Latent TGFβ complexes are transglutaminase cross-linked to fibrillin to facilitate TGFβ activation.
Matrix Biol (2022)
Lockhart-Cairns MP, Cain SA, Dajani R, Steer R, Thomson J, Alanazi YF, Kielty CM, Baldock C
|
| RgGuinier |
5.4 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
141 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Latent-transforming growth factor beta-binding protein 1 monomer, 44 kDa Homo sapiens protein
|
| Buffer: |
Tris buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Dec 12
|
Latent TGFβ complexes are transglutaminase cross-linked to fibrillin to facilitate TGFβ activation.
Matrix Biol (2022)
Lockhart-Cairns MP, Cain SA, Dajani R, Steer R, Thomson J, Alanazi YF, Kielty CM, Baldock C
|
| RgGuinier |
4.7 |
nm |
| Dmax |
20.5 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
|
|
|
|

|
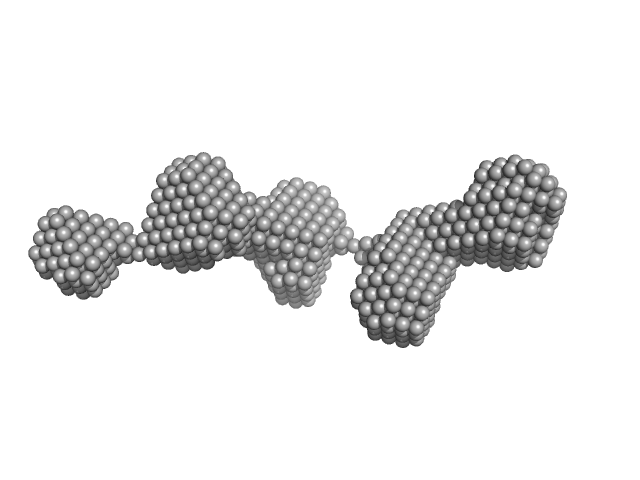
| Sample: |
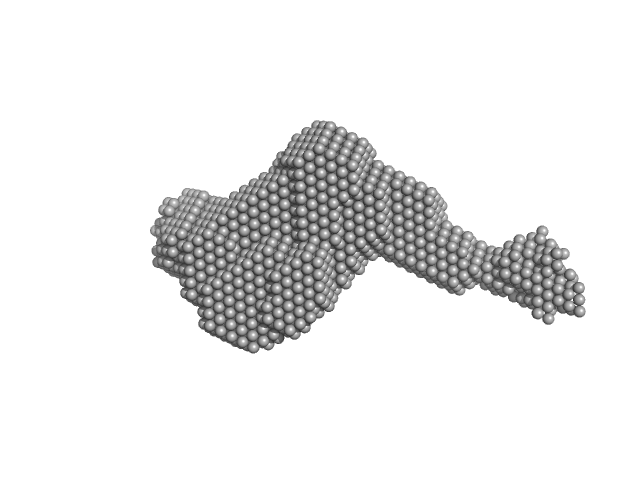
Latent-transforming growth factor beta-binding protein 1 monomer, 44 kDa Homo sapiens protein
Fibrillin-1 PF3 monomer, 78 kDa Homo sapiens protein
|
| Buffer: |
10 mM Hepes, 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jul 20
|
Latent TGFβ complexes are transglutaminase cross-linked to fibrillin to facilitate TGFβ activation.
Matrix Biol (2022)
Lockhart-Cairns MP, Cain SA, Dajani R, Steer R, Thomson J, Alanazi YF, Kielty CM, Baldock C
|
| RgGuinier |
7.4 |
nm |
| Dmax |
29.6 |
nm |
| VolumePorod |
759 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
7.8 |
nm |
| Dmax |
32.0 |
nm |
| VolumePorod |
1280 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
7.2 |
nm |
| Dmax |
29.5 |
nm |
| VolumePorod |
1270 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM EDTA, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
9.5 |
nm |
| Dmax |
40.0 |
nm |
| VolumePorod |
1370 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 (S2585C) dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
7.1 |
nm |
| Dmax |
29.5 |
nm |
| VolumePorod |
1260 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 (S2585C) dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
7.0 |
nm |
| Dmax |
29.5 |
nm |
| VolumePorod |
1270 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Teneurin-4 (S2585C) dimer, 537 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM EDTA, pH: 7.8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Dec 16
|
Teneurin4 dimer structures reveal a calcium‐stabilized compact conformation supporting homomeric trans‐interactions
The EMBO Journal (2022)
Meijer D, Frias C, Beugelink J, Deurloo Y, Janssen B
|
| RgGuinier |
7.8 |
nm |
| Dmax |
31.0 |
nm |
| VolumePorod |
1390 |
nm3 |
|
|
|
|
|
|
|
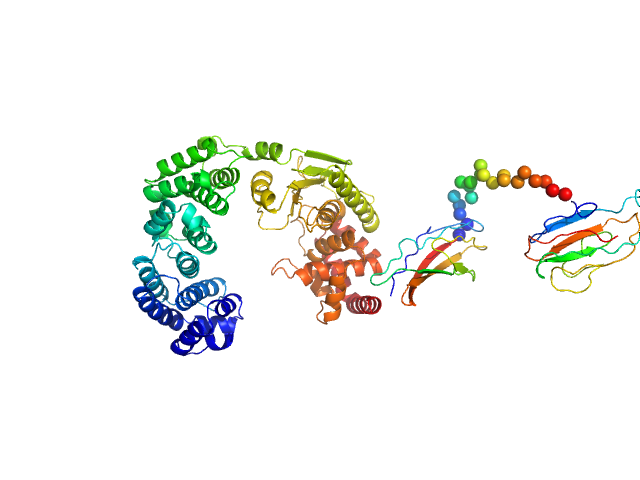
| Sample: |
Vibrio collagenase VhaC monomer, 90 kDa Vibrio harveyi protein
|
| Buffer: |
10 mM Tris-HCl, 100 mM NaCl, pH: 8
|
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2021 Jan 1
|
Structure of Vibrio collagenase VhaC provides insight into the mechanism of bacterial collagenolysis.
Nat Commun 13(1):566 (2022)
Wang Y, Wang P, Cao HY, Ding HT, Su HN, Liu SC, Liu G, Zhang X, Li CY, Peng M, Li F, Li S, Chen Y, Chen XL, Zhang YZ
|
| RgGuinier |
4.3 |
nm |
| Dmax |
17.6 |
nm |
| VolumePorod |
148 |
nm3 |
|
|