|
|
|
|
|
| Sample: |
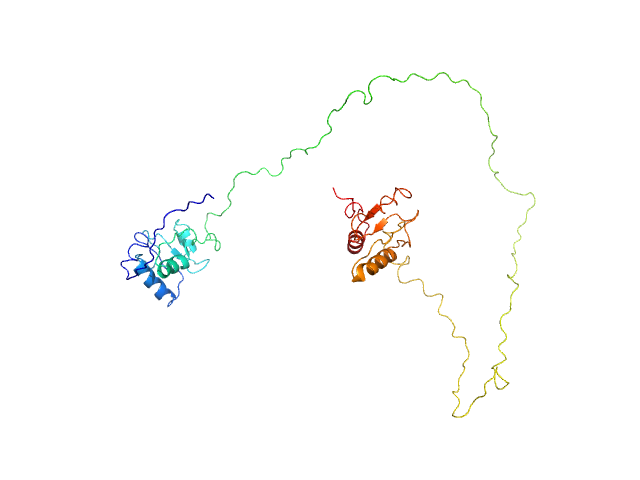
DNA repair protein XRCC1ΔN dimer, 76 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% glycerol, 2 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2011 Jul 15
|
An atypical BRCT-BRCT interaction with the XRCC1 scaffold protein compacts human DNA Ligase IIIα within a flexible DNA repair complex.
Nucleic Acids Res (2020)
Hammel M, Rashid I, Sverzhinsky A, Pourfarjam Y, Tsai MS, Ellenberger T, Pascal JM, Kim IK, Tainer JA, Tomkinson AE
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
170 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
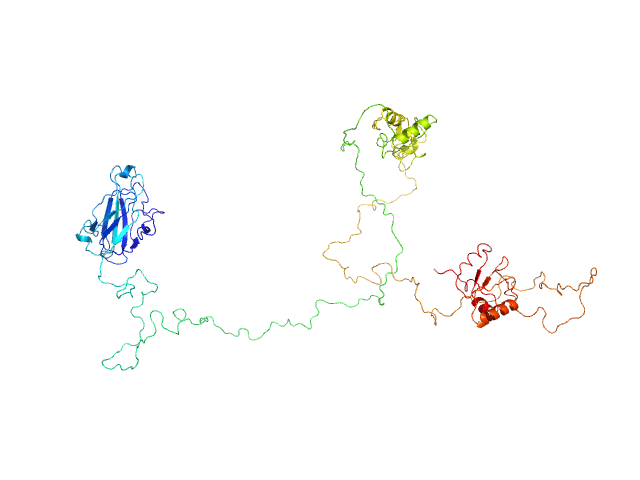
DNA repair protein XRCC1 , 69 kDa Homo sapiens protein
|
| Buffer: |
200 mM NaCl, 20 mM Tris-HCl, pH 7.5, 2% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jun 19
|
An atypical BRCT-BRCT interaction with the XRCC1 scaffold protein compacts human DNA Ligase IIIα within a flexible DNA repair complex.
Nucleic Acids Res (2020)
Hammel M, Rashid I, Sverzhinsky A, Pourfarjam Y, Tsai MS, Ellenberger T, Pascal JM, Kim IK, Tainer JA, Tomkinson AE
|
| RgGuinier |
6.3 |
nm |
| Dmax |
26.9 |
nm |
| VolumePorod |
330 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
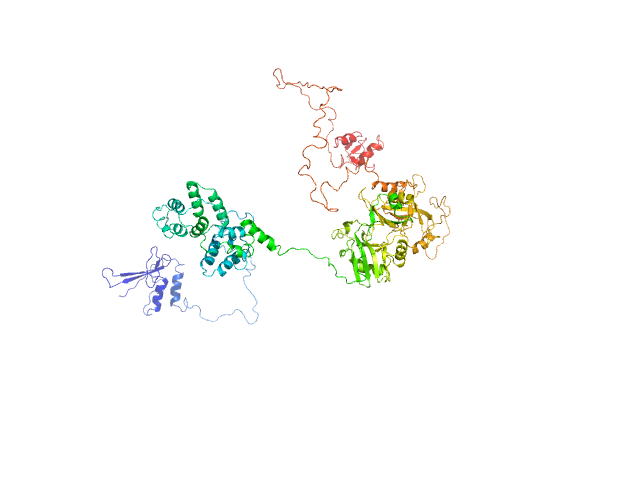
DNA ligase 3 (DNA ligase III alpha) , Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% glycerol, 2 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2011 Mar 31
|
An atypical BRCT-BRCT interaction with the XRCC1 scaffold protein compacts human DNA Ligase IIIα within a flexible DNA repair complex.
Nucleic Acids Res (2020)
Hammel M, Rashid I, Sverzhinsky A, Pourfarjam Y, Tsai MS, Ellenberger T, Pascal JM, Kim IK, Tainer JA, Tomkinson AE
|
| RgGuinier |
6.1 |
nm |
| Dmax |
21.0 |
nm |
| VolumePorod |
240 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
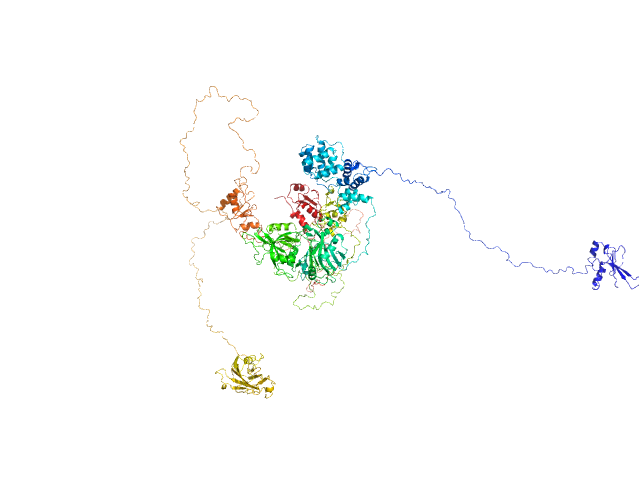
DNA repair protein XRCC1 , 69 kDa Homo sapiens protein
DNA ligase 3 (DNA ligase III alpha) , Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, pH 7.5, 100 mM NaCl, 5 mM MgCl₂, 0.2 mM PMSF, 1 mM benzamidine, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Aug 18
|
An atypical BRCT-BRCT interaction with the XRCC1 scaffold protein compacts human DNA Ligase IIIα within a flexible DNA repair complex.
Nucleic Acids Res (2020)
Hammel M, Rashid I, Sverzhinsky A, Pourfarjam Y, Tsai MS, Ellenberger T, Pascal JM, Kim IK, Tainer JA, Tomkinson AE
|
| RgGuinier |
6.2 |
nm |
| Dmax |
27.9 |
nm |
| VolumePorod |
675 |
nm3 |
|
|
|
|
|
|
|
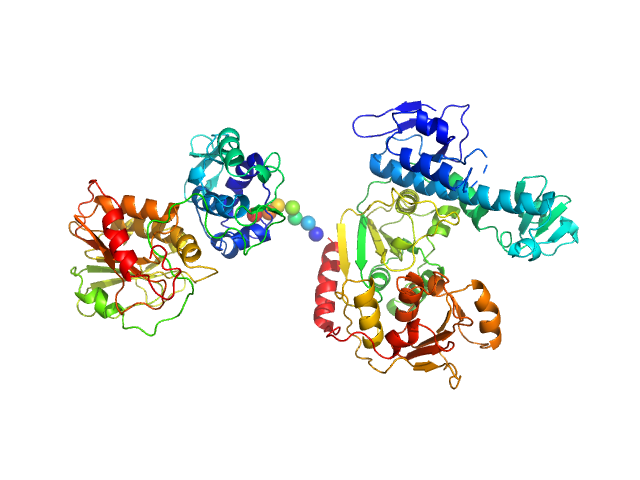
| Sample: |
NsP2 protein monomer, 90 kDa Chikungunya virus protein
|
| Buffer: |
20 mM Hepes pH 7.4, 150 mM NaCl, 1 mM DTT, 5% glycerol, pH: 7.4
|
| Experiment: |
SAXS
data collected at 23A, Taiwan Photon Source, NSRRC on 2019 Aug 10
|
Inter-domain Flexibility of Chikungunya Virus nsP2 Helicase-Protease Differentially Influences Viral RNA Replication and Infectivity.
J Virol (2020)
Law YS, Wang S, Tan YB, Shih O, Utt A, Goh WY, Lian BJ, Chen MW, Jeng US, Merits A, Luo D
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
134 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
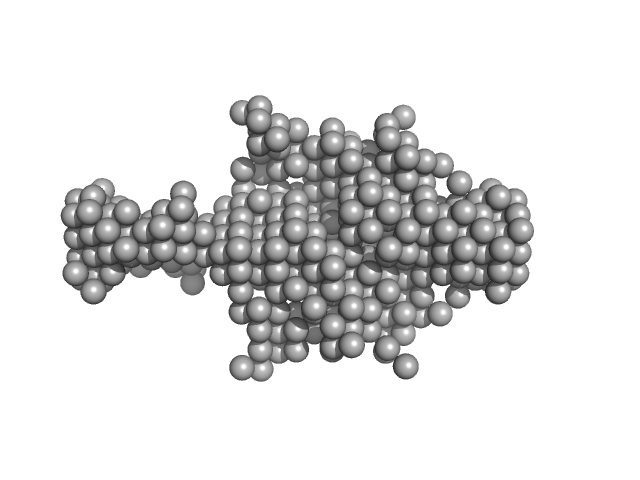
Evolutionarily conserved signaling intermediate in Toll pathway, mitochondrial dimer, 44 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 7
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.9 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
|
|
|
|
|
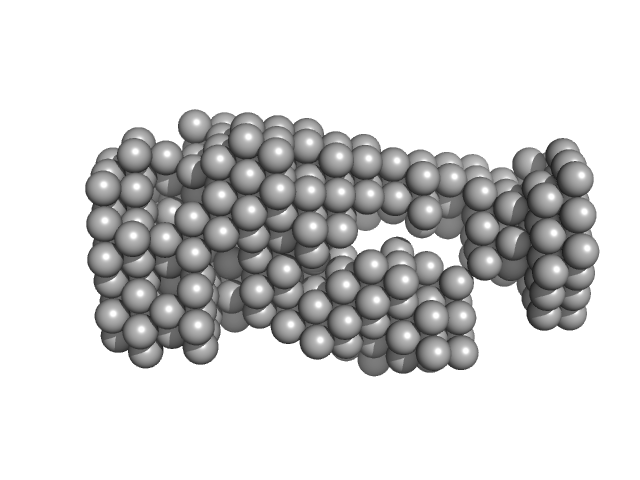
| Sample: |
Complex I intermediate-associated protein 30, mitochondrial monomer, 35 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 29
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.6 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
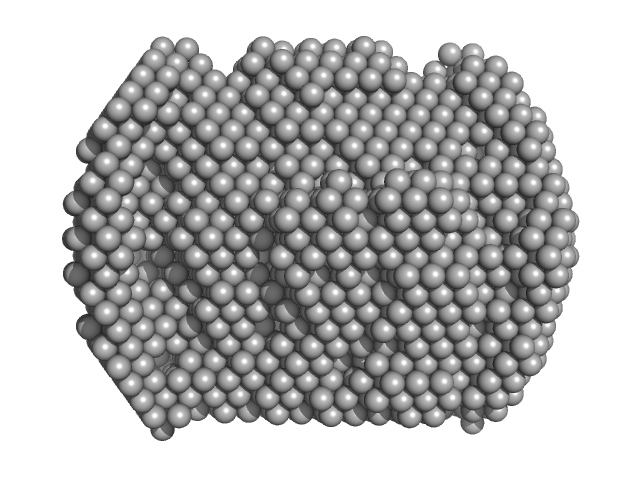
Complex I assembly factor ACAD9, mitochondrial dimer, 132 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 May 28
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.6 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
217 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
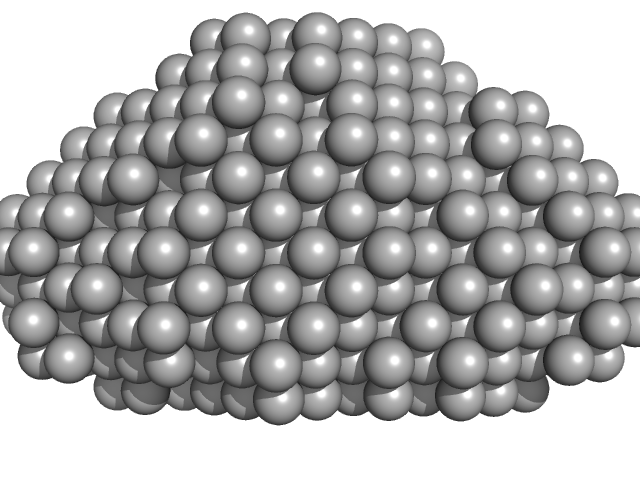
Complex I assembly factor ACAD9-VLCAD chimera dimer, 131 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Nov 30
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
217 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
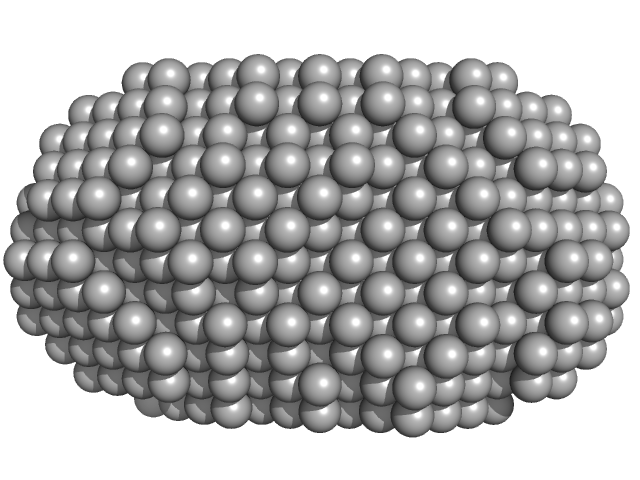
Very long-chain specific acyl-CoA dehydrogenase, mitochondrial dimer, 126 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 2
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
215 |
nm3 |
|
|