|
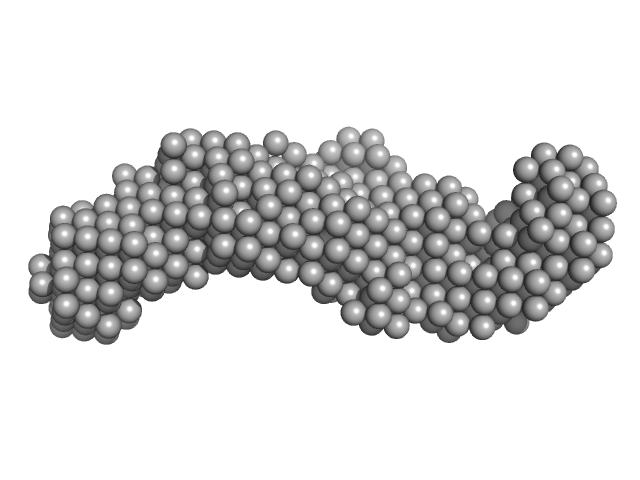
Small angle X-ray scattering measurements (SAXS) were performed at the SAXS2 beamline located at the Brazilian Synchrotron Light Laboratory (LNLS), Campinas, São Paulo, Brazil. The X-ray scattering data (I(s) vs s, where s = 4πsinθ/λ and 2θ is the scattering angle; λ = 0.1488 nm) were acquired using a two-dimension position-sensitive MARR-CCD detector and a sample-to-detector distance of ~1000 mm, corresponding to the q-range 0 < q < 0.35. In order to checking for protein aggregation or concentration-dependent effects, samples were prepared at various protein concentrations (2 mg/mL and 3 mg/mL) in 25 mM Tris-HCl (pH 7.5) buffer, containing 150 mM NaCl, 2 mM EDTA and 1 mM β-mercaptoethanol. Samples were exposed to X-ray at different time frames (5 x 30 seconds, and 180 seconds) to investigate for X-ray damage. No effects of protein concentration or X-ray damage were detected. The model depicts the averaged spatial representation of the protein (DAMFILT occupancy and volume-corrected bead model).
|
|
Co-chaperone p23
(Pfp23B)
|
| Mol. type |
|
Protein |
| Organism |
|
Plasmodium falciparum |
| Olig. state |
|
Monomer |
| Mon. MW |
|
30.9 kDa |
| |
| UniProt |
|
Q8IKU1
|
| Sequence |
|
FASTA |
| |
|
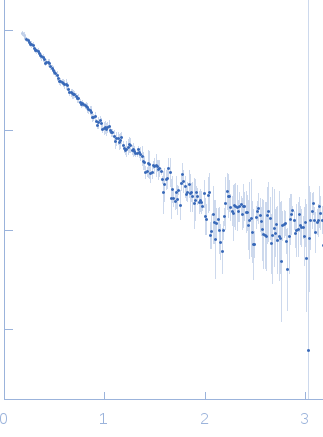 s, nm-1
s, nm-1
