|
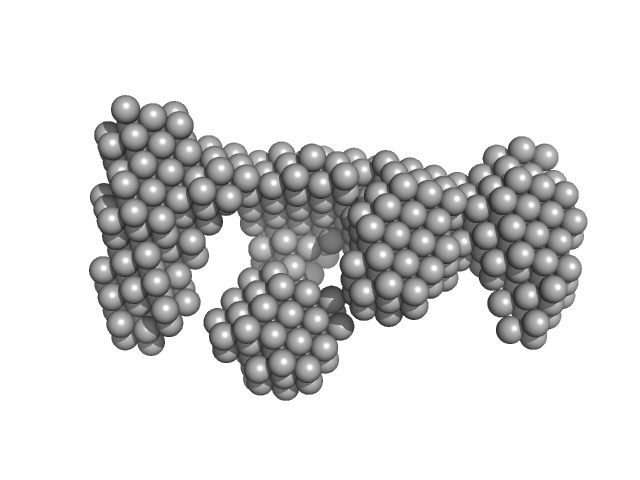
Synchrotron SAXS data from solutions of Mycobacterium tuberculosis DNA LigaseA with nicked DNA in 50 mM Tris-HCl 200 mM NaCl 2mM β-mercaptoethanol, pH 8 were collected on the BM29 beam line at the ESRF storage ring (Grenoble, France) using a Pilatus 1M detector at a sample-detector distance of 1.1 m and at a wavelength of λ = 0.081 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). One solute concentration of 2.50 mg/ml was measured at 10°C. 10 successive 1 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
|
|
DNA ligase A
|
| Mol. type |
|
Protein |
| Organism |
|
Mycobacterium tuberculosis |
| Olig. state |
|
Monomer |
| Mon. MW |
|
76.1 kDa |
| |
| UniProt |
|
P9WNV1
|
| Sequence |
|
FASTA |
| |
|
Nicked DNA
|
| Mol. type |
|
DNA |
| Olig. state |
|
Dimer |
| Mon. MW |
|
8.1 kDa |
| Sequence |
|
FASTA |
| |
|
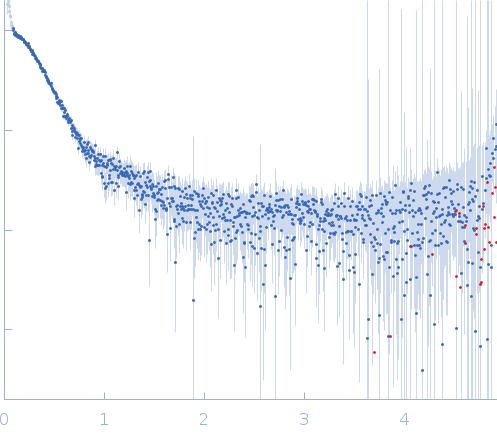 s, nm-1
s, nm-1
