|
Synchrotron SAXS data from solutions of cyclic GMP-AMP synthase (cGAS) with 2'3'-cGAMP in 20 mM HEPES, pH 7.4 were measured using size-exclusion chromatography SAXS (SEC-SAXS) on the EMBL P12 beam line at the PETRA III storage ring (Hamburg, Germany) using a Pilatus 2M detector at a sample-detector distance of 3 m and at a wavelength of λ = 0.124 nm (l(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). SEC-SAXS was performed at 20 °C using the following parameters: Column: XXXX ; Flow rate: 0.5 mL/min; Total acquisition time: 3600 x 1 s frames; Sample injection concentration: 9 mg/mL; Injection volume: 75 μL. The data obtained through the sample elution peak were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted and the individual subtracted data sets were scaled and averaged to generate the scattering profile displayed in this entry.
SEC column = UNKNOWN. Sample injection volume = UNKNOWN. Flow rate = UNKNOWN
|
|
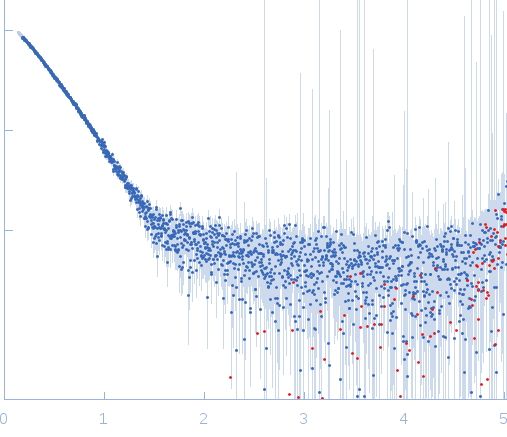 s, nm-1
s, nm-1
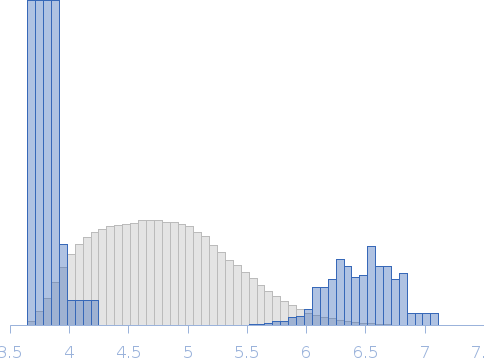 Rg, nm
Rg, nm