|
SAXS data from solutions of SARS-CoV-2 main protease in 50 mM Tris, 1 mM DTT, 1 mM EDTA, pH 7.4 were collected using a Rigaku BioSAXS-2000 instrument (University of British Columbia, Vancouver, Canada) equipped with a HyPix-3000 detector at a wavelength of λ = 0.154 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 1.2 and 9.6 mg/ml were measured at 6°C. 12 successive 300 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The low angle data collected at lower concentrations were extrapolated to infinite dilution and merged with the higher concentration data to yield the final composite scattering curve.
Sample detector distance = UNKNOWN
|
|
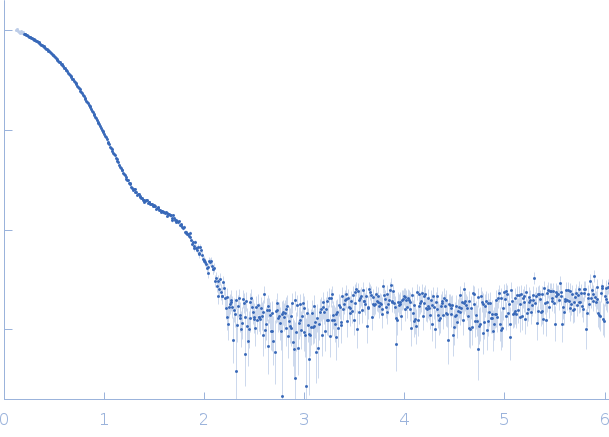 s, nm-1
s, nm-1