A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Trewhella J,
Vachette P,
Bierma J,
Blanchet C,
Brookes E,
Chakravarthy S,
Chatzimagas L,
Cleveland T,
Cowieson N,
Crossett B,
Duff A,
Franke D,
Gabel F,
Gillilan R,
Graewert M,
Grishaev A,
Guss J,
Hammel M,
Hopkins J,
Huang Q,
Hub J,
Hura G,
Irving T,
Jeffries C,
Jeong C,
Kirby N,
Krueger S,
Martel A,
Matsui T,
Li N,
Pérez J,
Porcar L,
Prangé T,
Rajkovic I,
Rocco M,
Rosenberg D,
Ryan T,
Seifert S,
Sekiguchi H,
Svergun D,
Teixeira S,
Thureau A,
Weiss T,
Whitten A,
Wood K,
Zuo X
Acta Crystallographica Section D Structural Biology
78(11)
(2022 Oct 20)
|
|
|
|
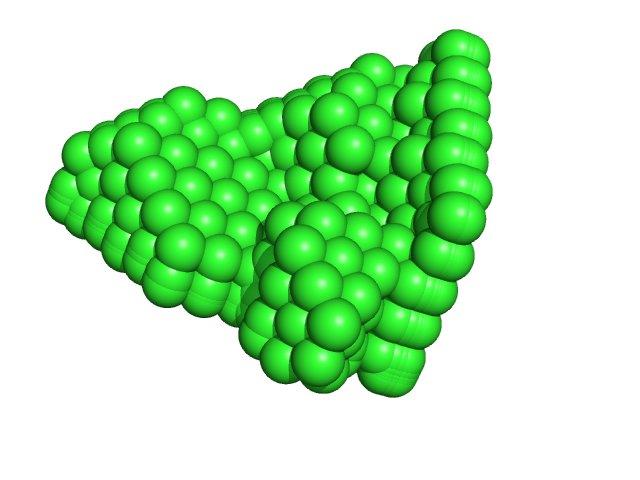

|
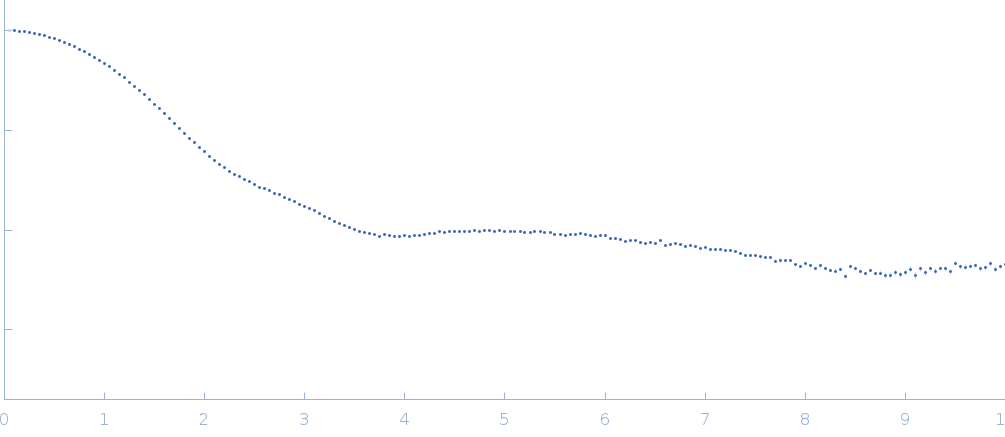
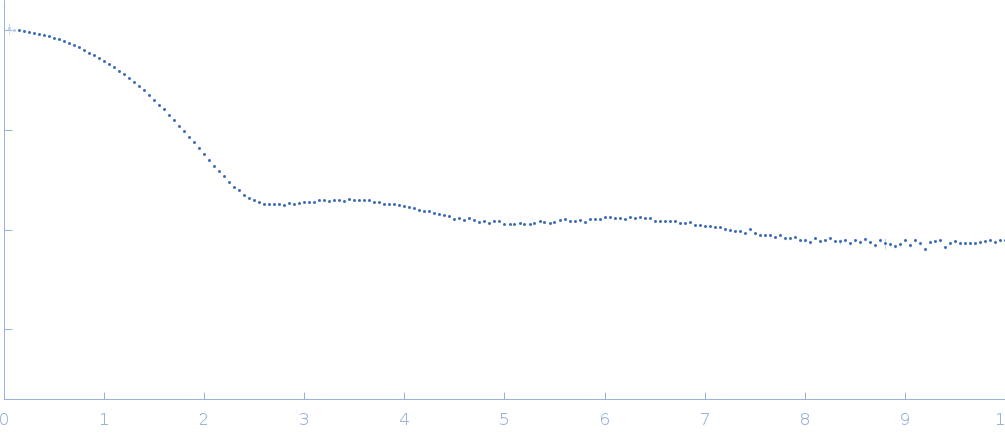
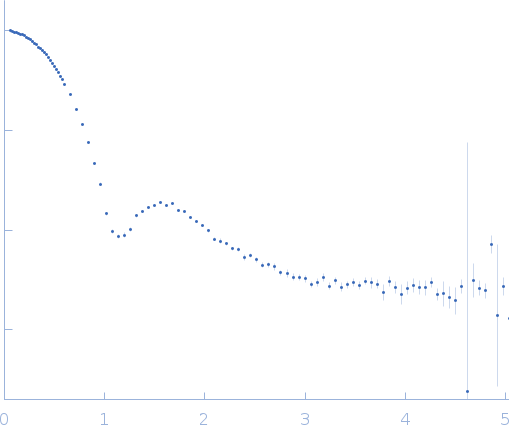
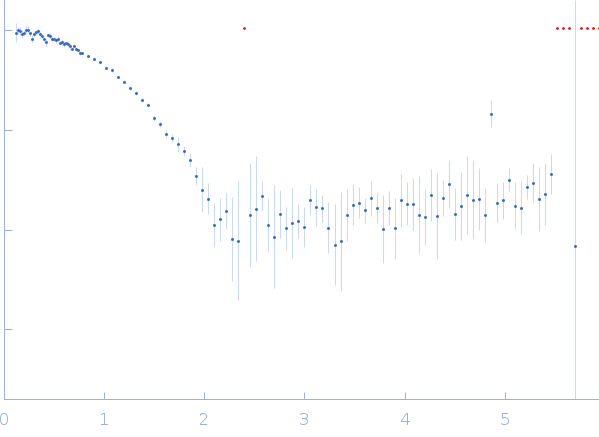
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.9 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
|
|
|
|
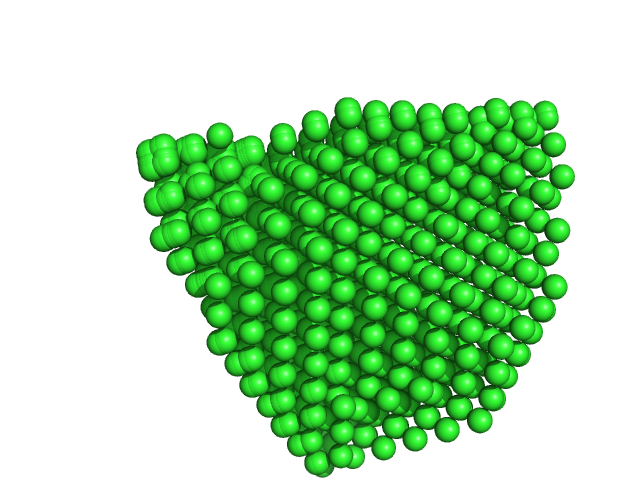

|
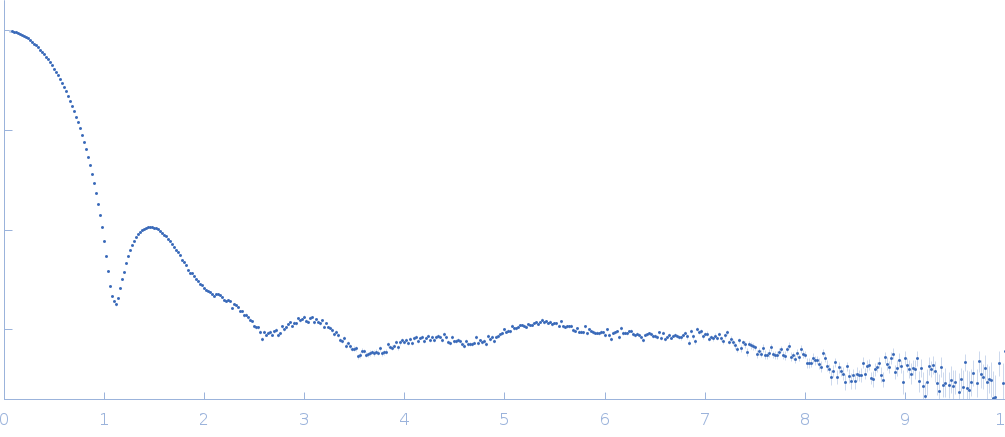
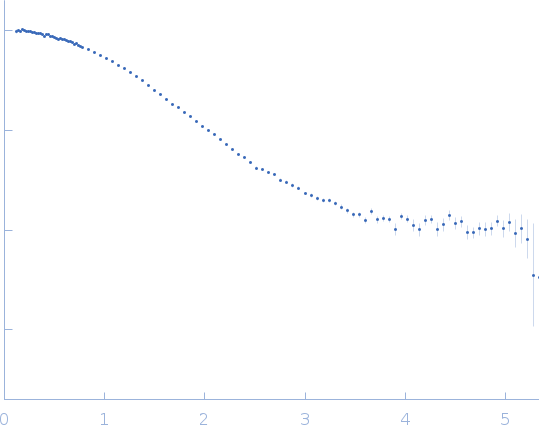
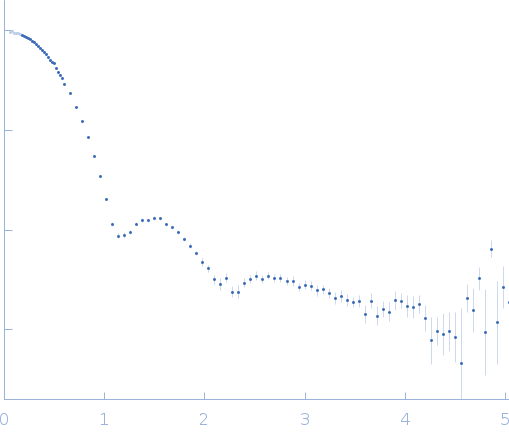
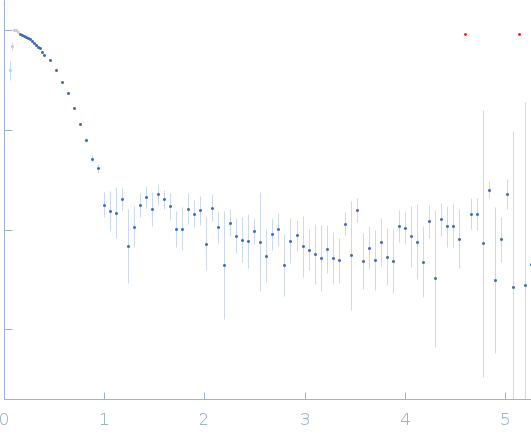
| Sample: |
Uricase tetramer, 136 kDa Aspergillus flavus protein
|
| Buffer: |
100 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
220 |
nm3 |
|
|
|
|
|
|

|
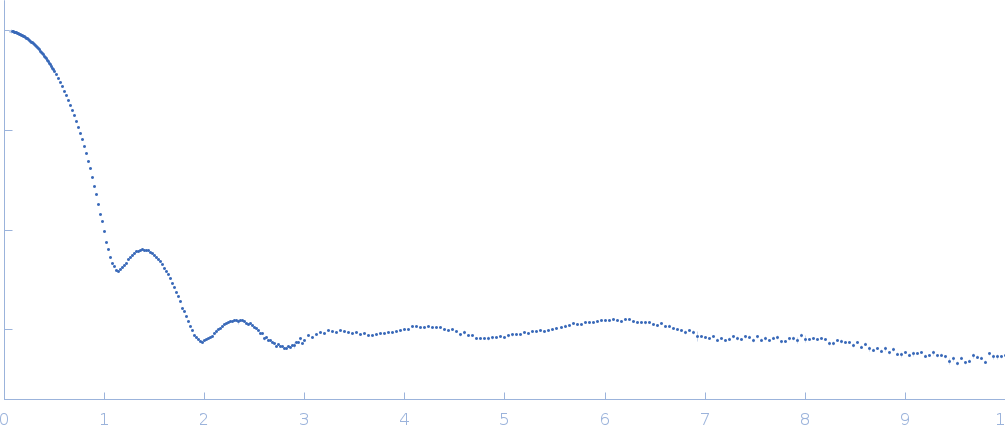
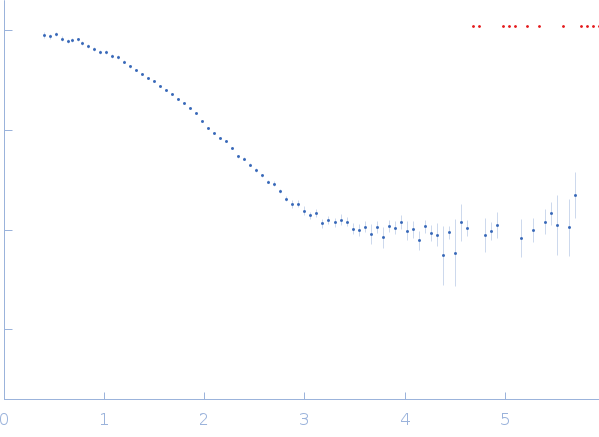
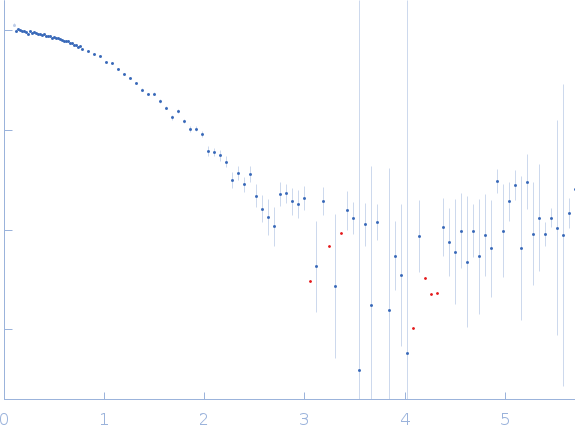
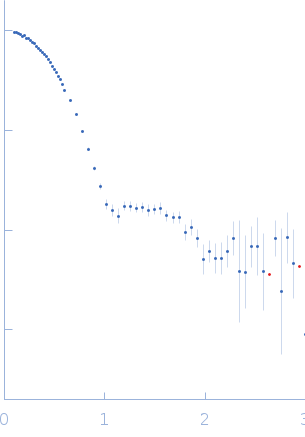
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
ConsensusBuffer_50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
243 |
nm3 |
|
|
|
|
|
|

|
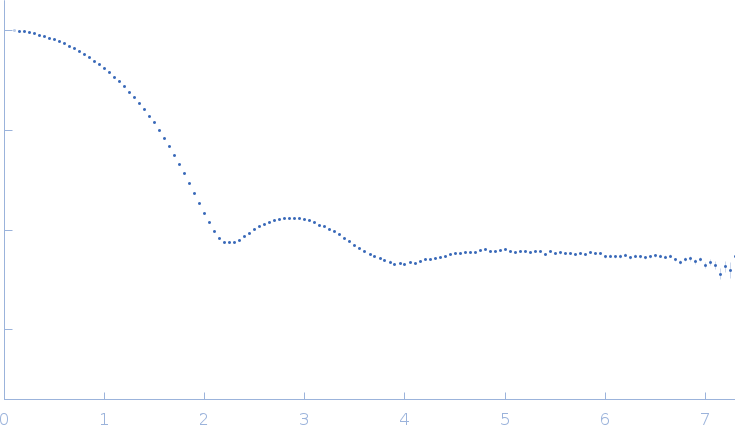
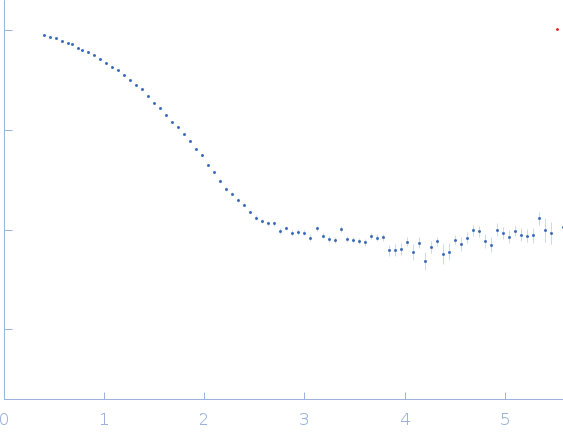
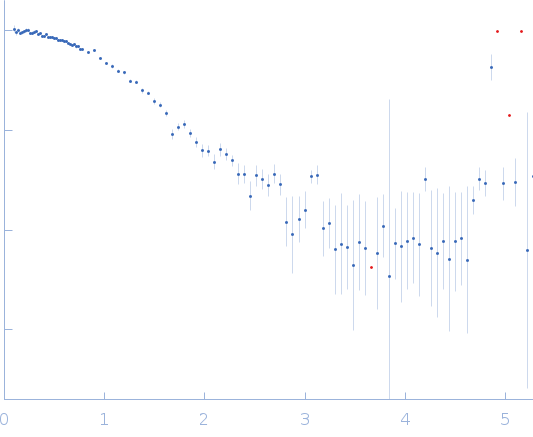
| Sample: |
Endo-1,4-beta-xylanase monomer, 21 kDa Trichoderma longibrachiatum protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.1 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
50 mM sodium citrate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.8 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
|
|
|
|
|
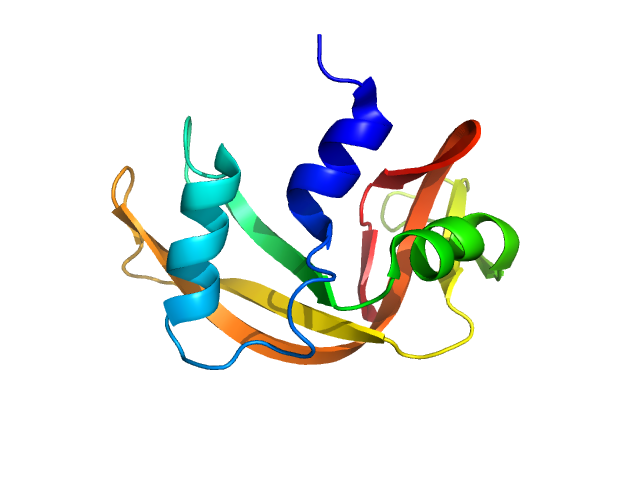
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.4 |
nm |
|
|
|
|
|
|
|
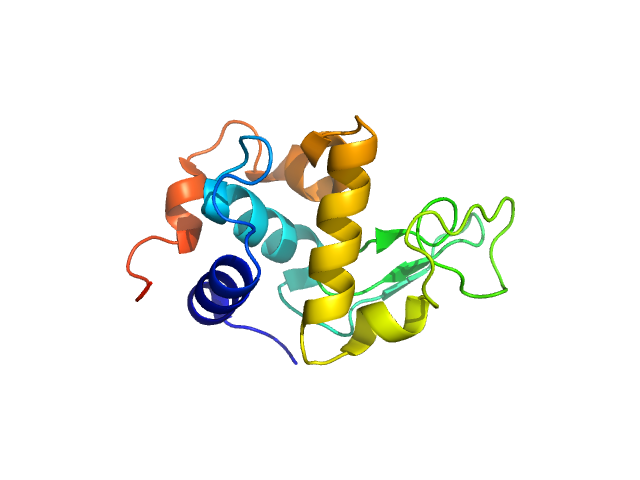
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
50 mM sodium citrate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.2 |
nm |
| Dmax |
3.8 |
nm |
|
|
|
|
|
|
|
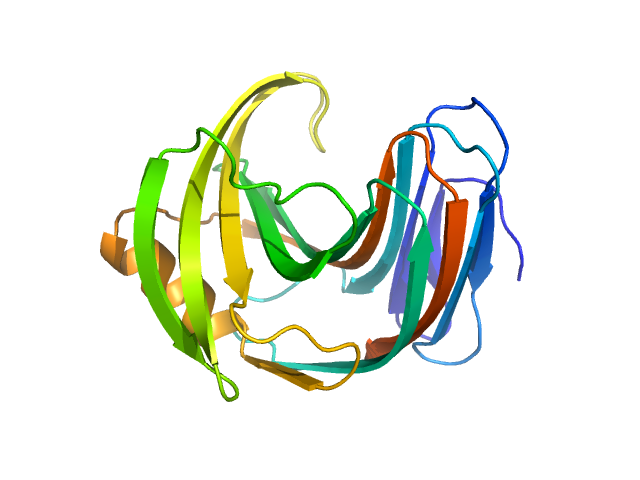
| Sample: |
Endo-1,4-beta-xylanase monomer, 21 kDa Trichoderma longibrachiatum protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.4 |
nm |
|
|
|
|
|
|
|
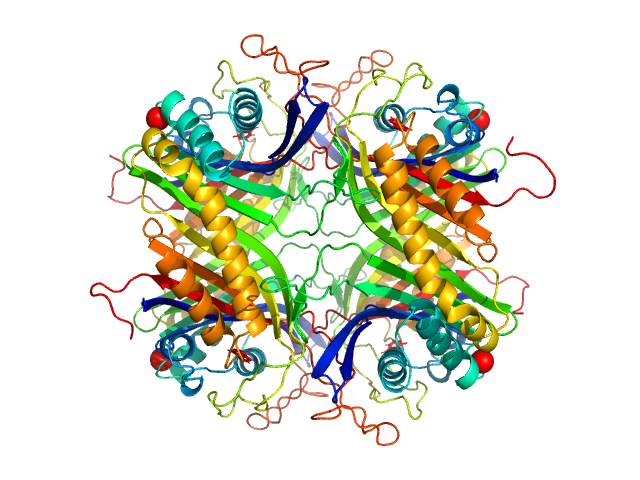
| Sample: |
Uricase tetramer, 136 kDa Aspergillus flavus protein
|
| Buffer: |
100 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.3 |
nm |
|
|
|
|
|
|
|
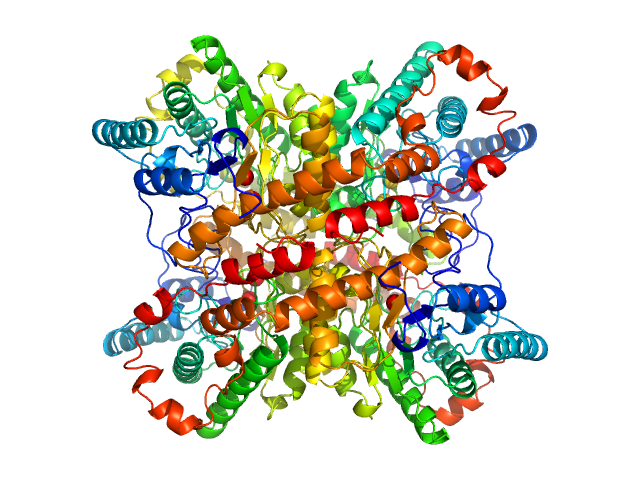
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.5 |
nm |
|
|
|
|
|
|
|
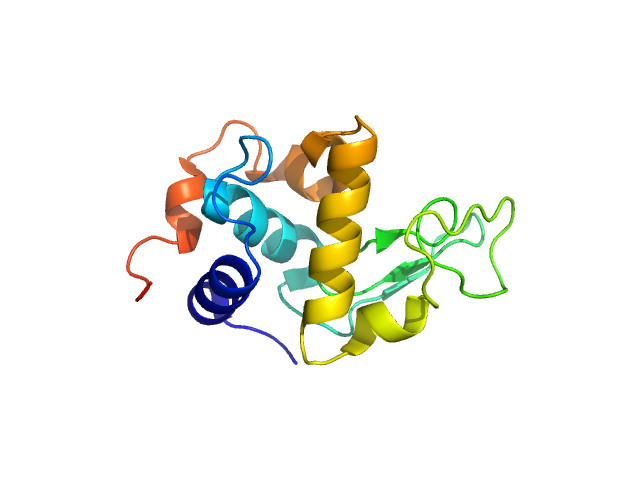
| Sample: |
Lysozyme C monomer, 14 kDa Gallus gallus protein
|
| Buffer: |
50 mM sodium citrate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 22
|
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.8 |
nm |
|
|
|
|
|
|
|
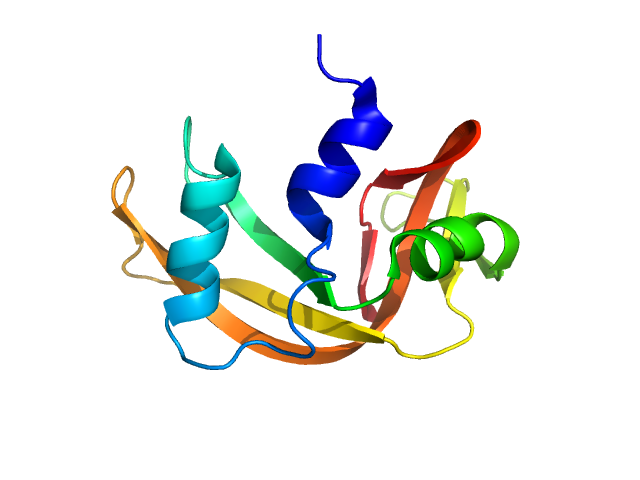
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 22
|
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.1 |
nm |
|
|
|
|
|
|
|
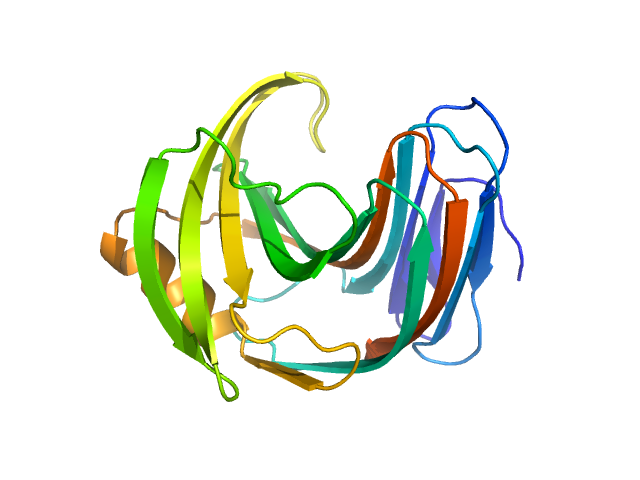
| Sample: |
Endo-1,4-beta-xylanase monomer, 21 kDa Trichoderma longibrachiatum protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 22
|
|
| RgGuinier |
1.6 |
nm |
| Dmax |
4.3 |
nm |
|
|
|
|
|
|
|
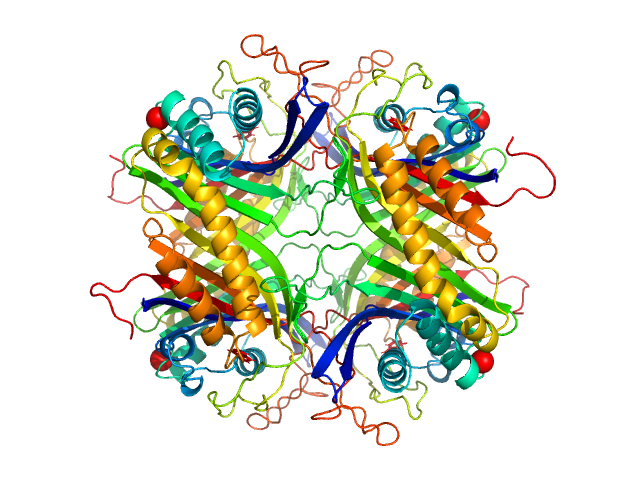
| Sample: |
Uricase tetramer, 136 kDa Aspergillus flavus protein
|
| Buffer: |
100 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 24
|
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.1 |
nm |
|
|
|
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 24
|
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.7 |
nm |
|
|