Chaotic advection mixer for capturing transient states of diverse biological macromolecular systems with time-resolved small-angle X-ray scattering
Zielinski K,
Katz A,
Calvey G,
Pabit S,
Milano S,
Aplin C,
San Emeterio J,
Cerione R,
Pollack L
IUCrJ
10(3):363-375
(2023 Apr 28)
|
|
|
|
|
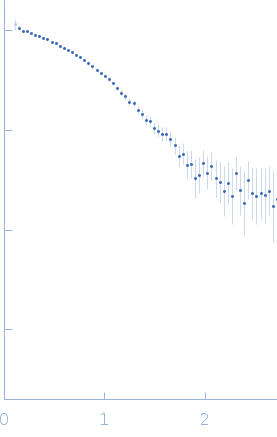
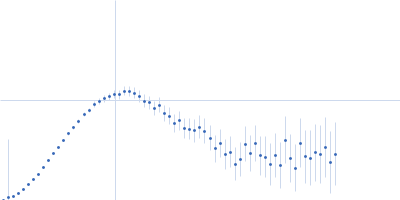
| Sample: |
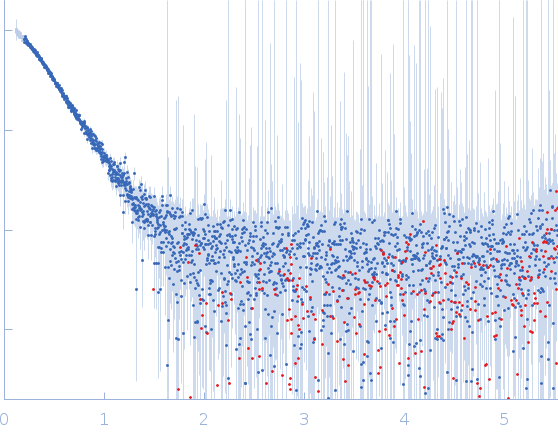
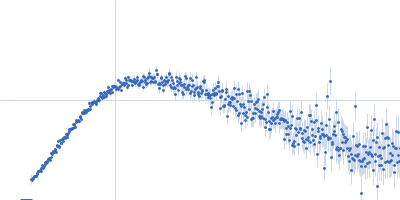
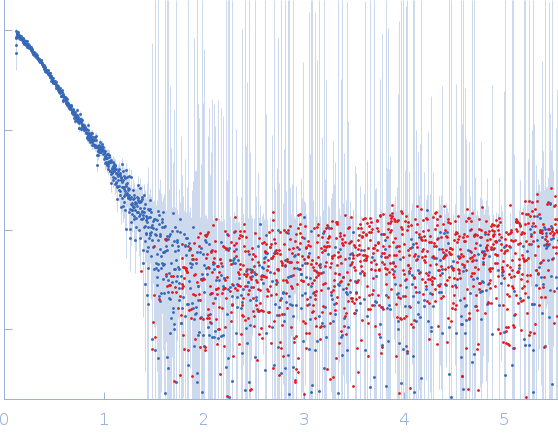
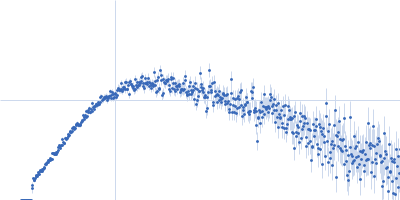
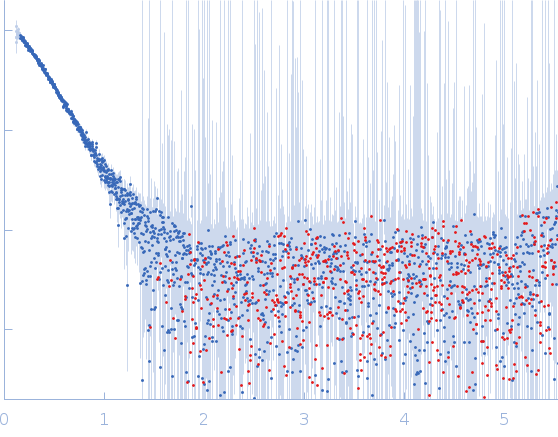
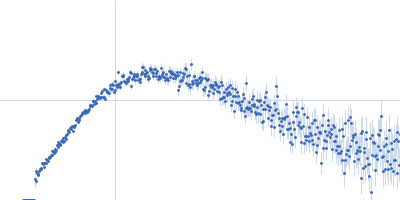
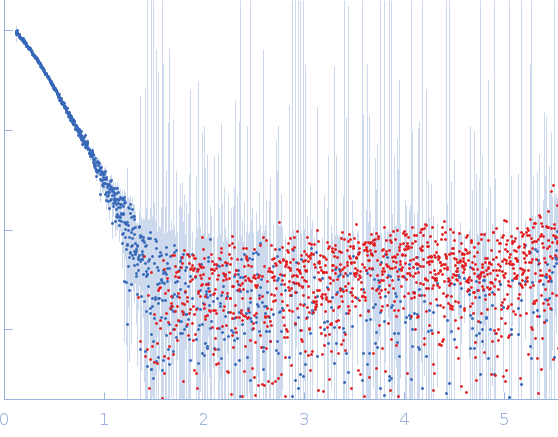
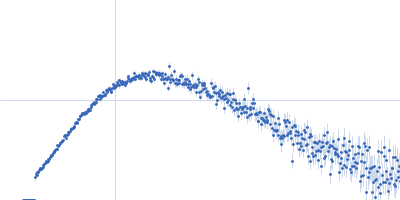
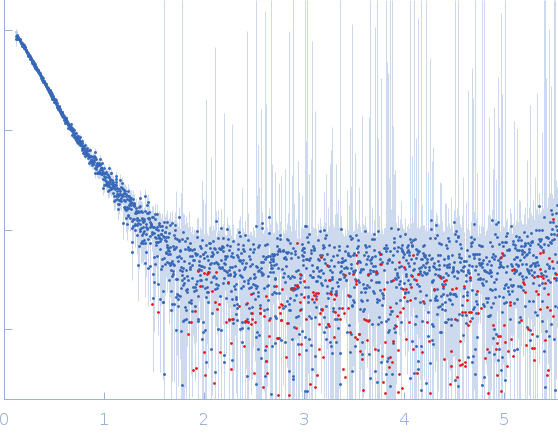
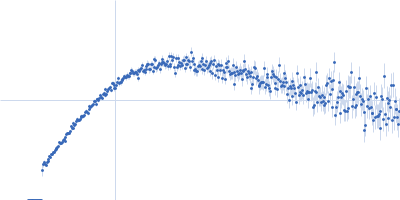
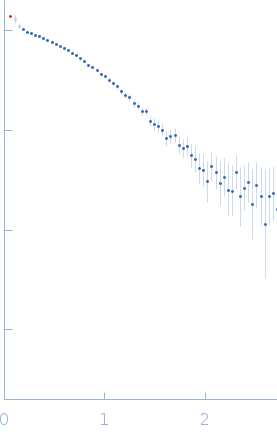
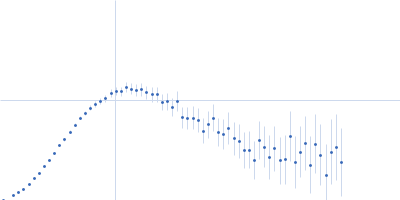
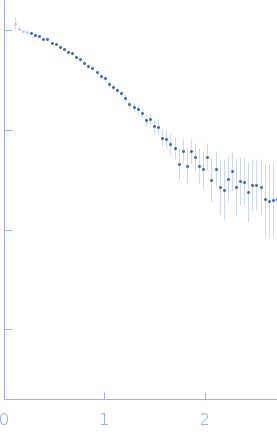
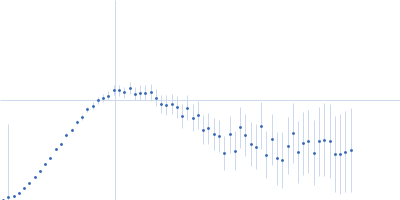
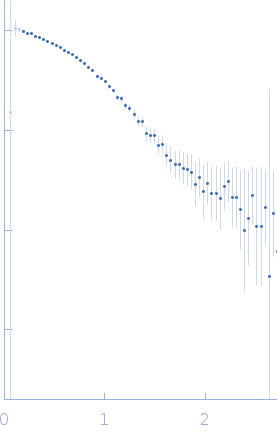
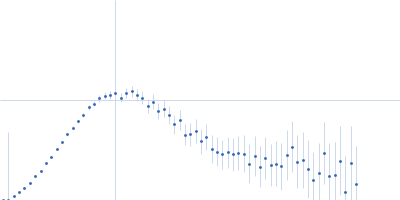
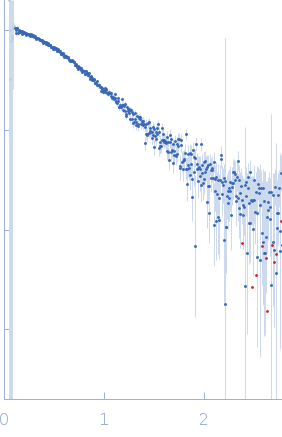
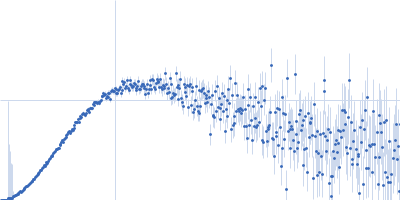
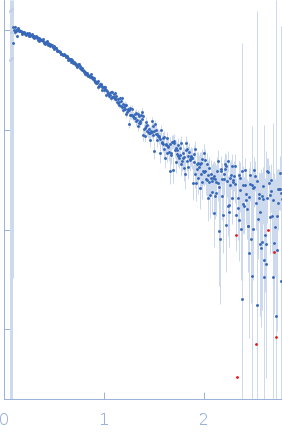
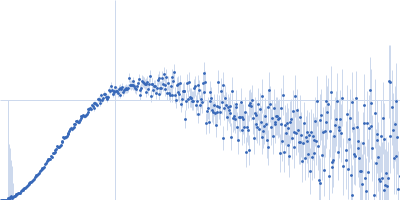
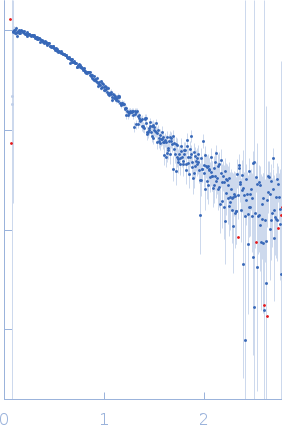
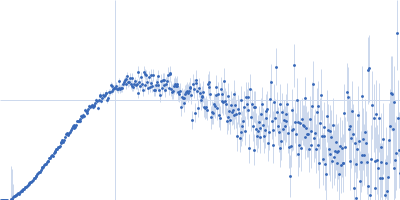
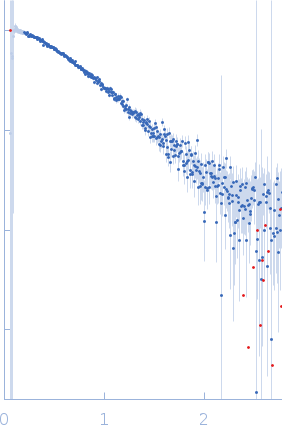
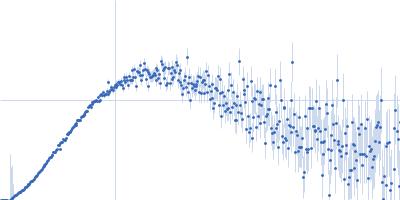
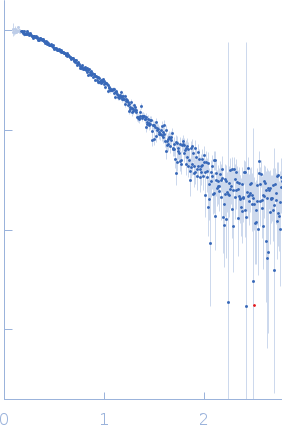
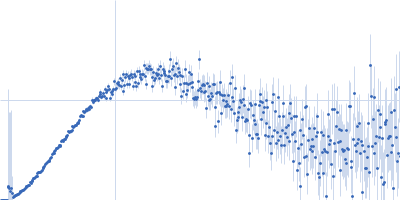
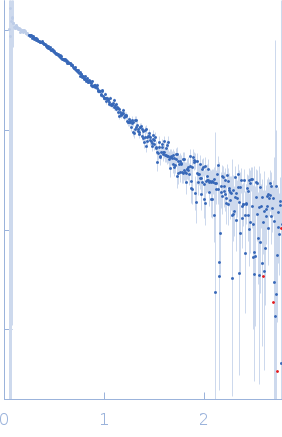
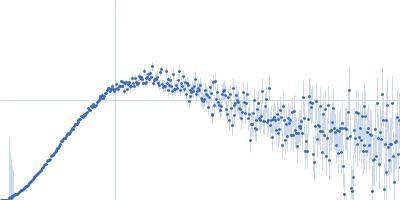
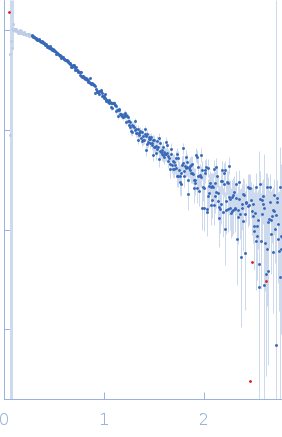
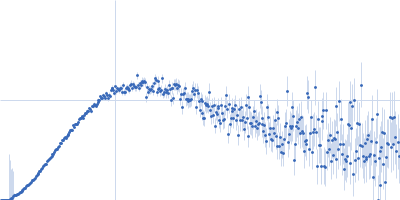
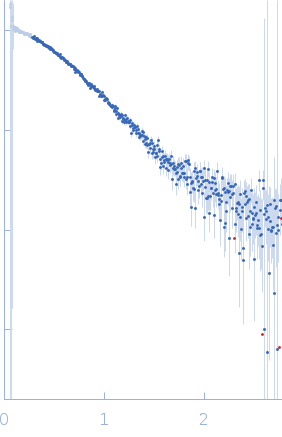
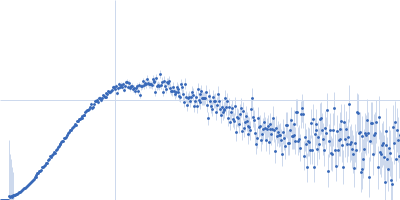
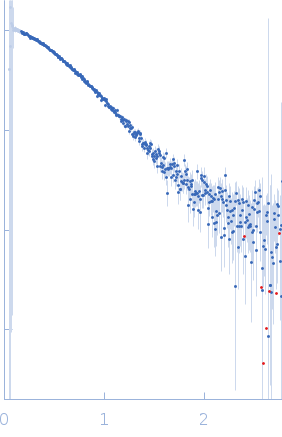
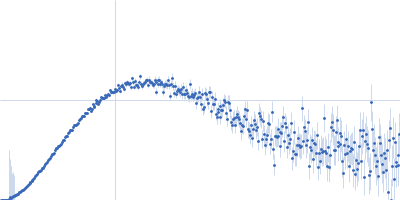
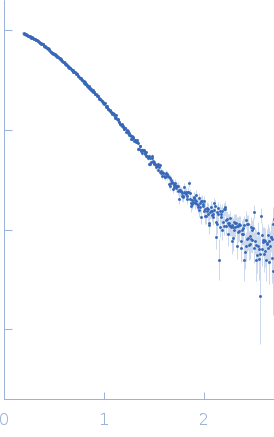
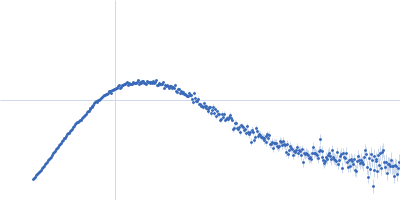
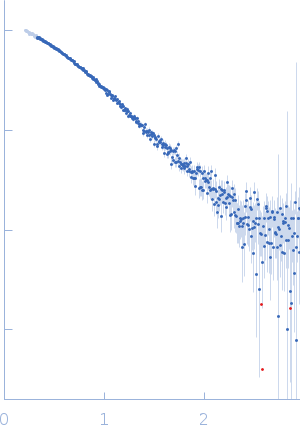
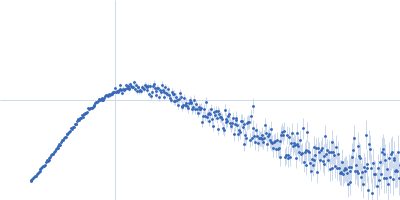
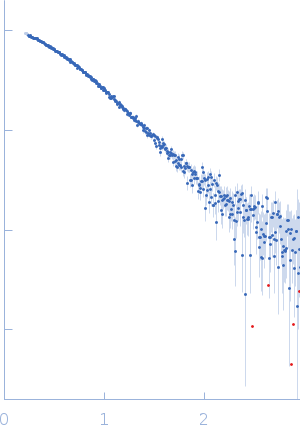
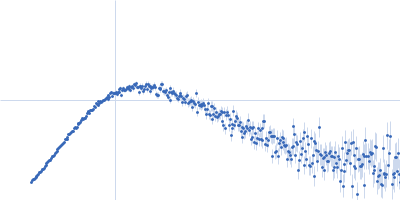
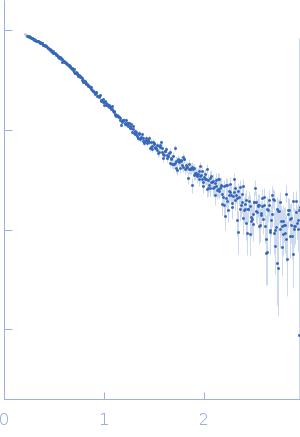
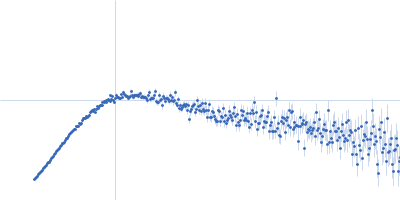
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.1 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
150 |
nm3 |
|
|
|
|
|
|
|
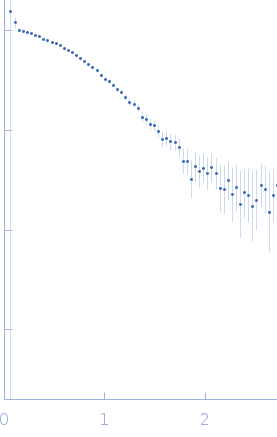
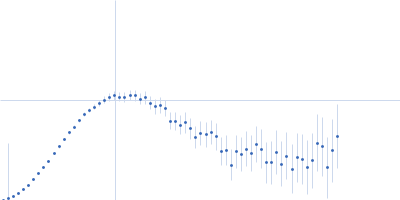
| Sample: |
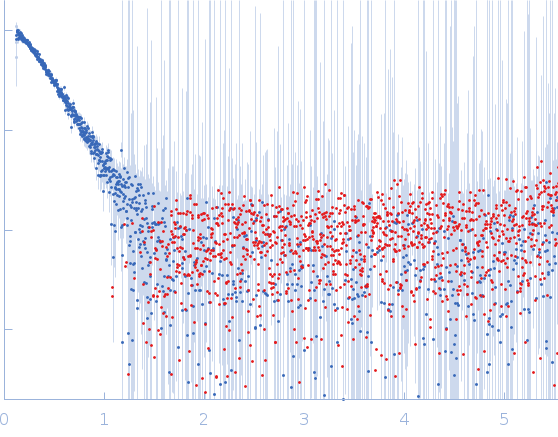
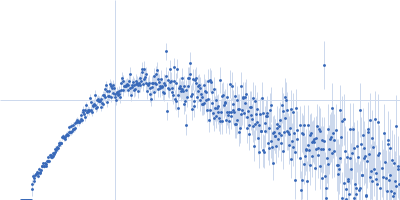
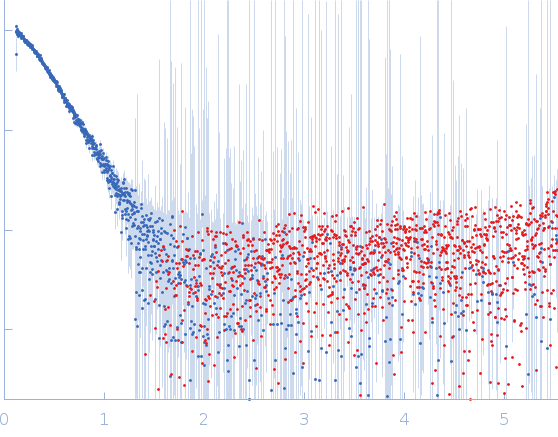
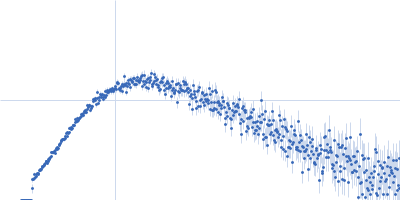
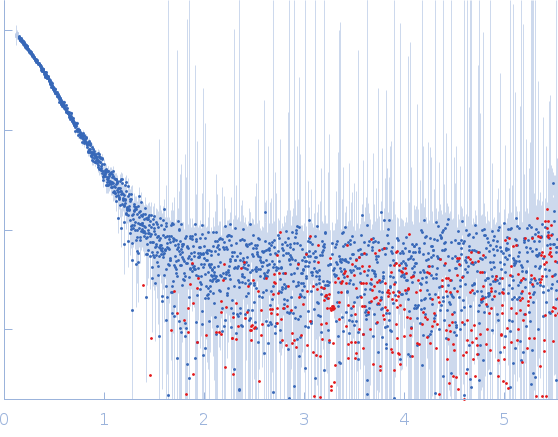
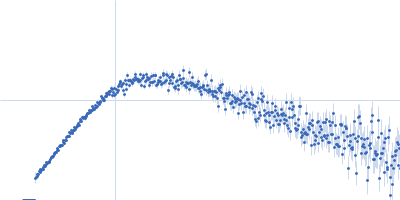
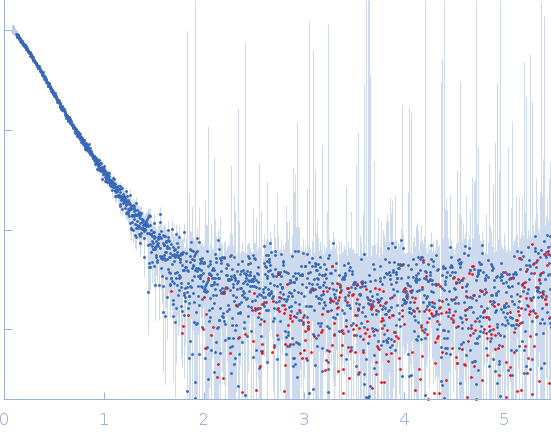
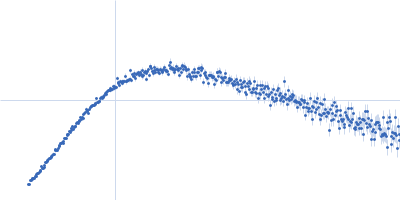
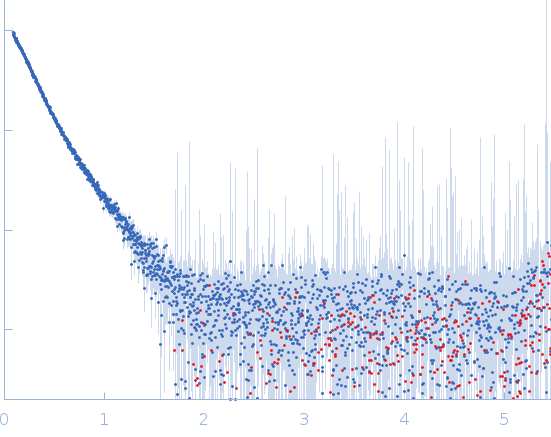
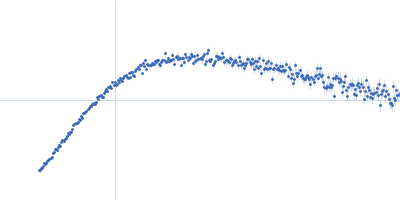
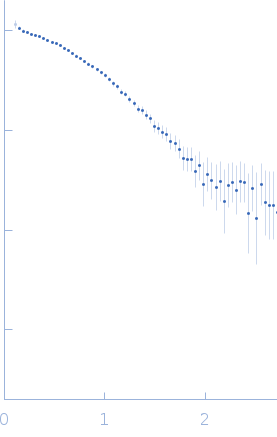
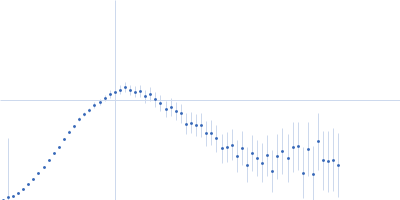
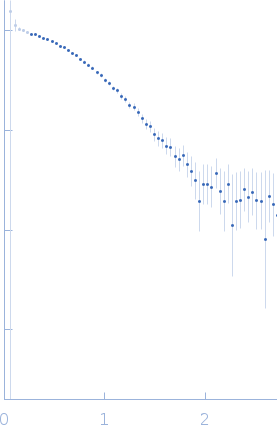
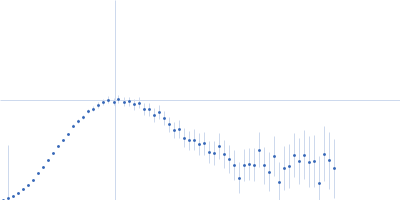
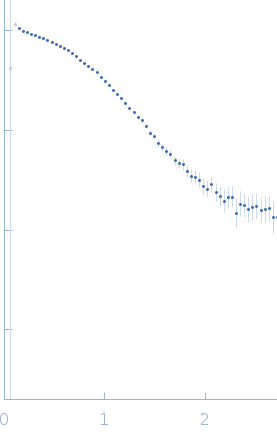
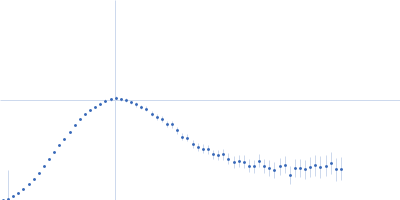
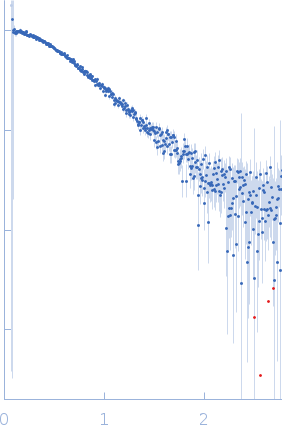
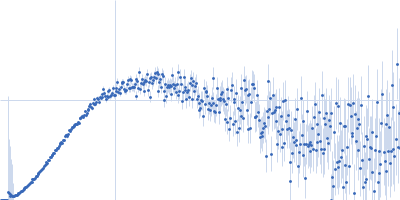
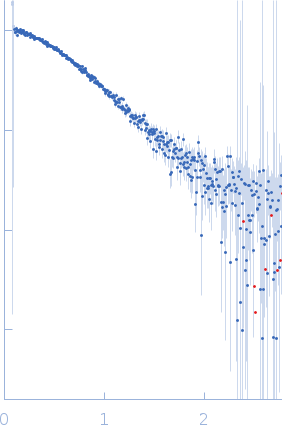
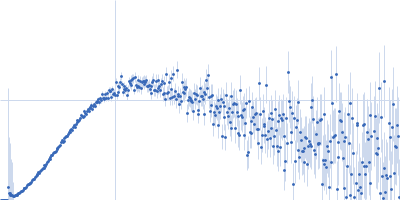
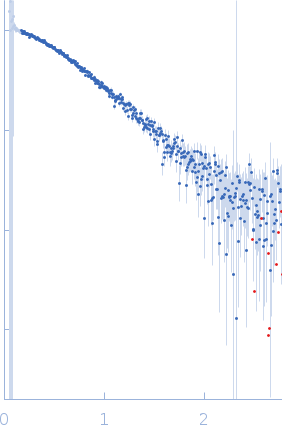
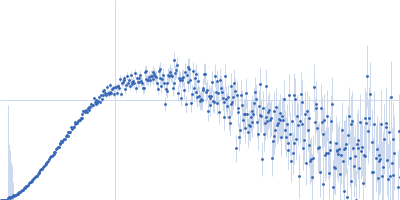
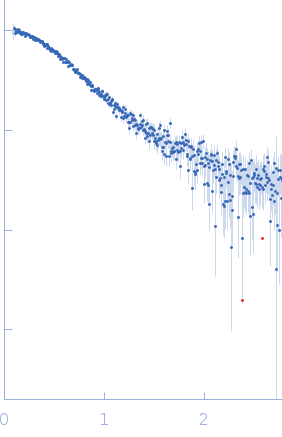
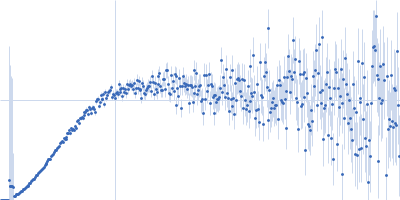
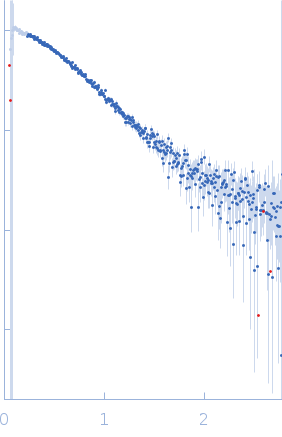
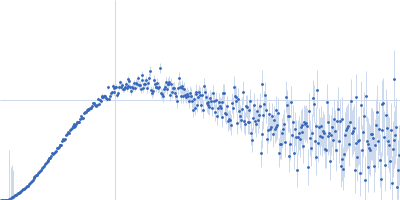
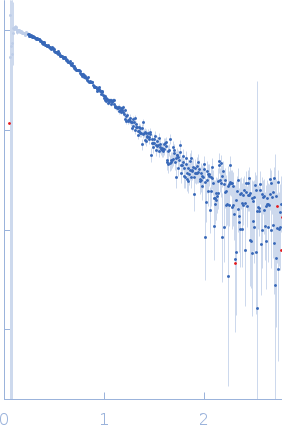
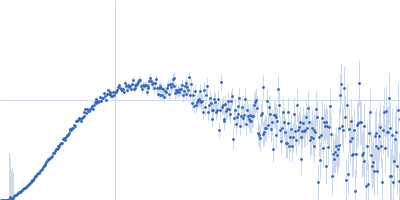
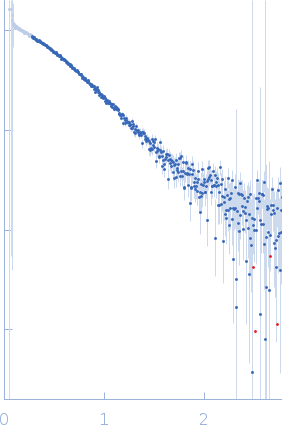
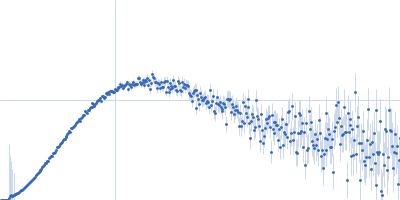
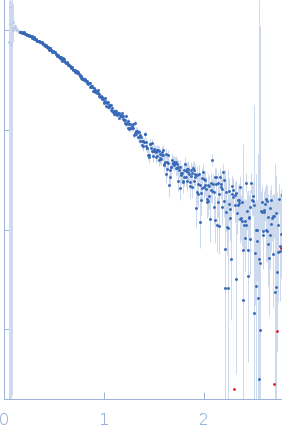
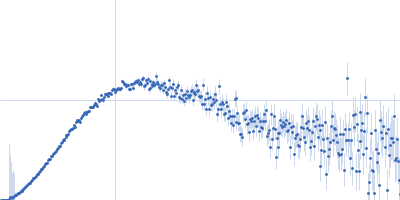
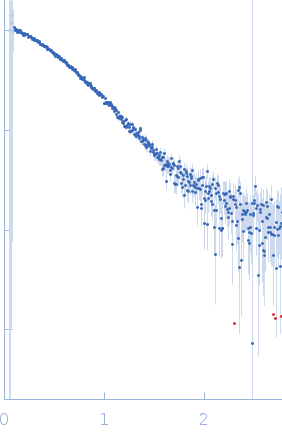
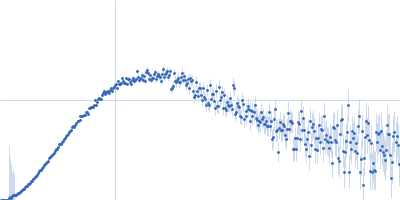
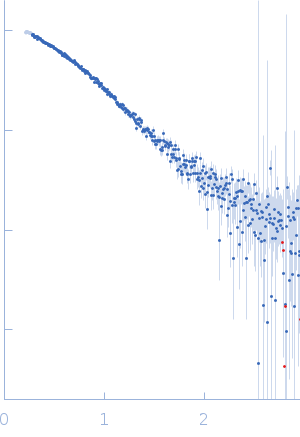
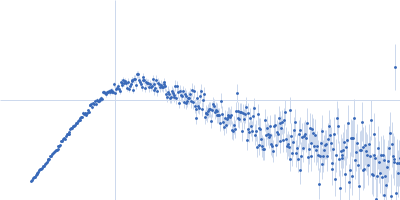
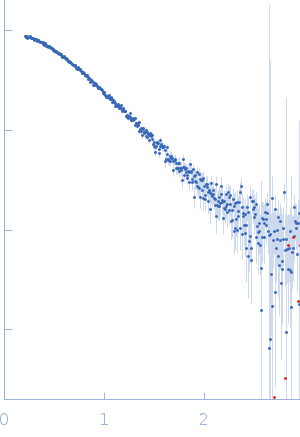
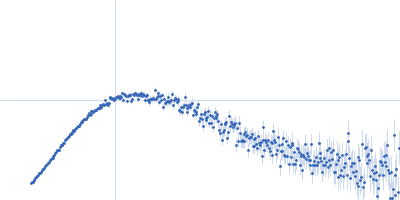
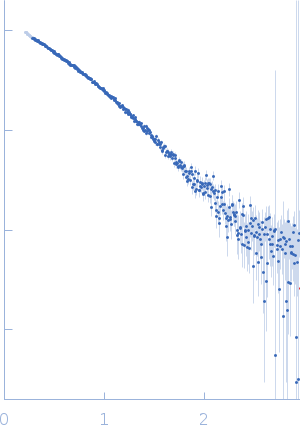
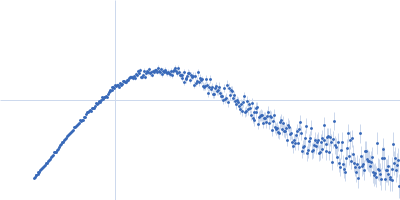
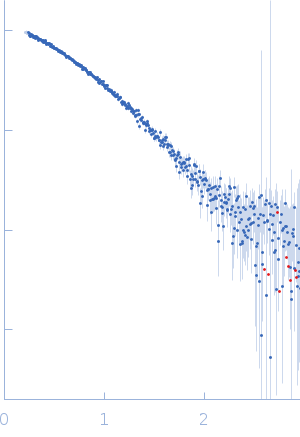
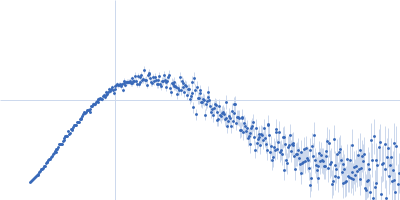
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.1 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
170 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.1 |
nm |
| Dmax |
18.0 |
nm |
| VolumePorod |
155 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.1 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.5 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
200 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.6 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
215 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
4.6 |
nm |
| Dmax |
28.0 |
nm |
| VolumePorod |
230 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Sep 29
|
|
| RgGuinier |
5.0 |
nm |
| Dmax |
27.0 |
nm |
| VolumePorod |
241 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Nov 19
|
|
| RgGuinier |
5.5 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
300 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Protein-glutamine gamma-glutamyltransferase 2, 77 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 10% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2021 Sep 29
|
|
| RgGuinier |
6.9 |
nm |
| Dmax |
31.5 |
nm |
| VolumePorod |
490 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.0 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Serine protease 1 monomer, 26 kDa Bos taurus protein
Pancreatic trypsin inhibitor monomer, 11 kDa Bos taurus protein
|
| Buffer: |
20 mM TRIS, 40 mM KCl, 20 mM CaCl2, pH: 7 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.3 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.3 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
34 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
33 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
37 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.5 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2018 May 9
|
|
| RgGuinier |
2.6 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM Na-MOPSO, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2017 Nov 19
|
|
| RgGuinier |
2.6 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
58 nucleotide RNA L11-binding domain from E. coli 23S rRNA monomer, 19 kDa Escherichia coli RNA
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.4 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
34 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
50S ribosomal protein L11 monomer, 16 kDa Thermus thermophilus protein
|
| Buffer: |
10 mM sodium cacodylate, 100 mM KCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2016 Dec 9
|
|
| RgGuinier |
2.2 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
27 |
nm3 |
|
|