Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure.
Korasick DA,
Singh H,
Pemberton TA,
Luo M,
Dhatwalia R,
Tanner JJ
FEBS J
284(18):3029-3049
(2017 Sep)
|
|
|
|

|
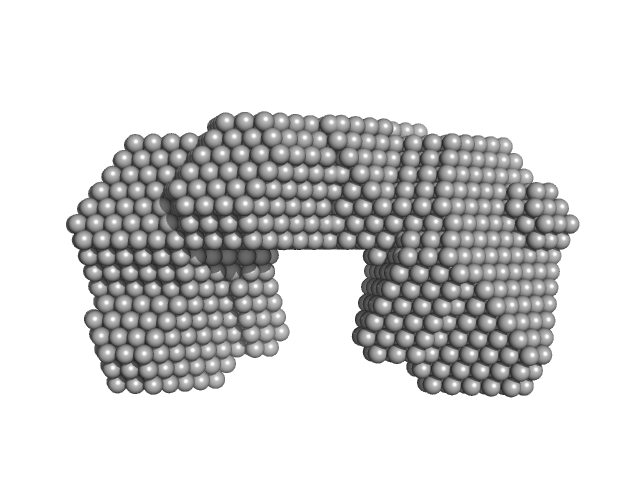
| Sample: |
Bifunctional protein PutA dimer, 219 kDa Bdellovibrio bacteriovorus protein
|
| Buffer: |
50 mM Tris, 125 mM NaCl, 1 mM EDTA, and 1 mM tris(3-hydroxypropyl)phosphine (THP) at pH 7.5,, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
287 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 229 kDa Desulfovibrio vulgaris protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
293 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 238 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 20
|
|
| RgGuinier |
4.6 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
291 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Proline dehydrogenase tetramer, 430 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
|
| RgGuinier |
5.3 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
541 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Proline dehydrogenase tetramer, 430 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 12
|
|
| RgGuinier |
5.2 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
553 |
nm3 |
|
|
|
|
|
|

|
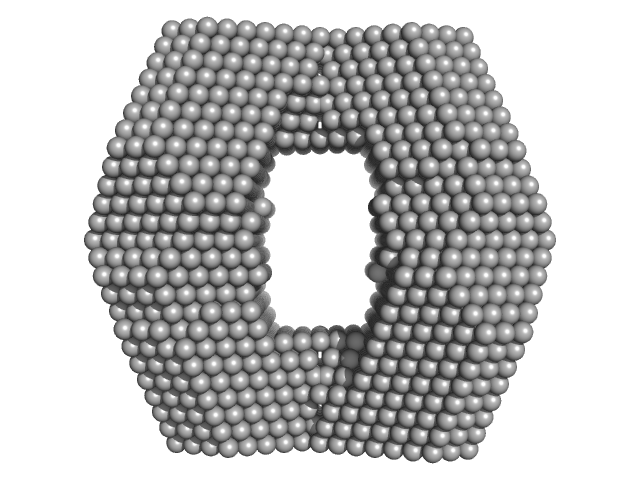
| Sample: |
Proline dehydrogenase tetramer, 430 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 12
|
|
| RgGuinier |
5.2 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
560 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 238 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 20
|
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
297 |
nm3 |
|
|
|
|
|
|

|
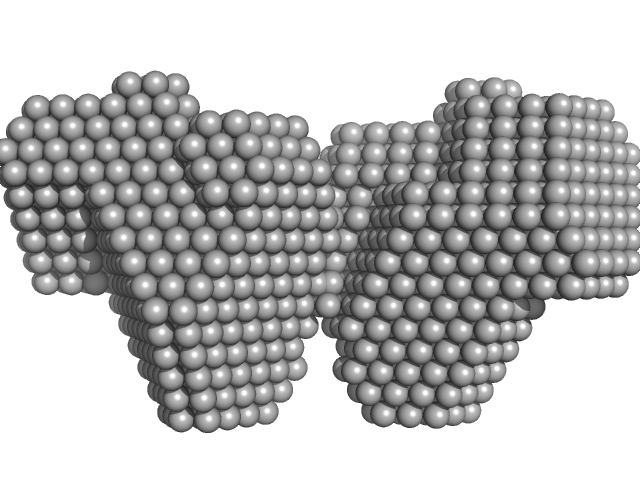
| Sample: |
Bifunctional protein PutA dimer, 238 kDa Legionella pneumophila subsp. … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 20
|
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
295 |
nm3 |
|
|
|
|
|
|

|
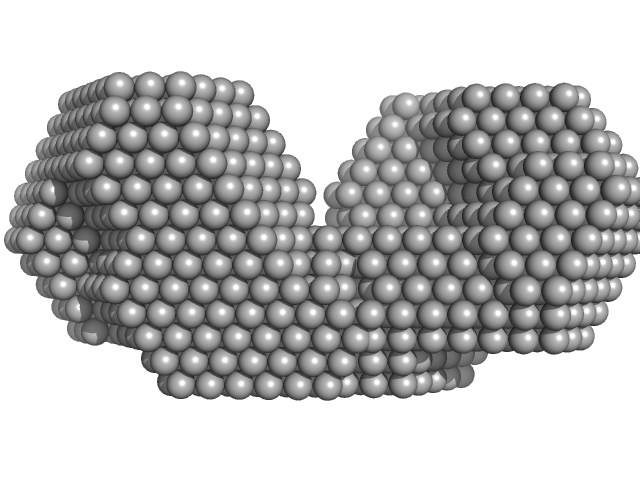
| Sample: |
Bifunctional protein PutA dimer, 229 kDa Desulfovibrio vulgaris protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
295 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bifunctional protein PutA dimer, 229 kDa Desulfovibrio vulgaris protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5., pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2012 Jun 8
|
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
294 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
281 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
|
| RgGuinier |
4.5 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
283 |
nm3 |
|
|
|
|
|
|

|
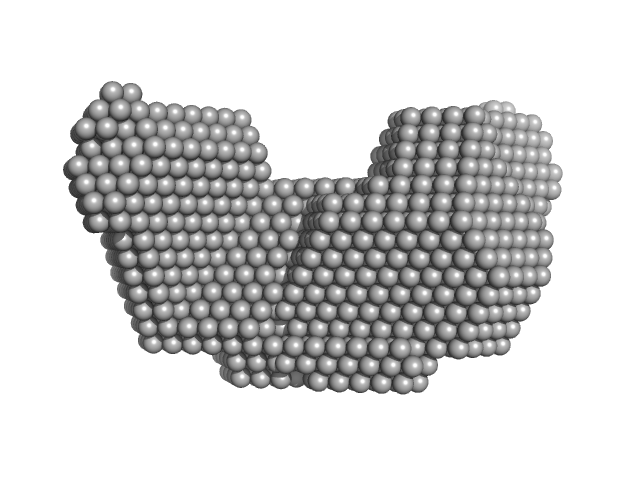
| Sample: |
Proline dehydrogenase dimer, 215 kDa Bradyrhizobium diazoefficiens protein
|
| Buffer: |
50 mM Tris (pH 7.8), 50 mM NaCl, 0.5 mM Tris(2-carboxyethyl)phosphine, and 5% (v/v) glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 16
|
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
289 |
nm3 |
|
|