Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Furlong EJ,
Kurth F,
Premkumar L,
Whitten AE,
Martin JL
Acta Crystallogr D Struct Biol
75(Pt 3):296-307
(2019 Mar 1)
|
|
|
|
|
| Sample: |
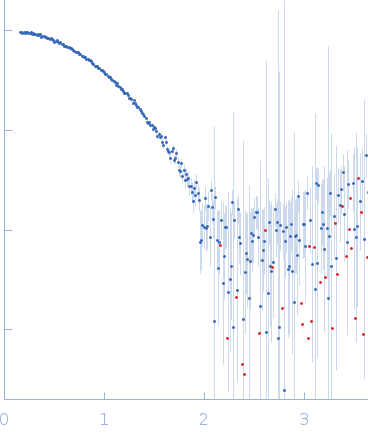
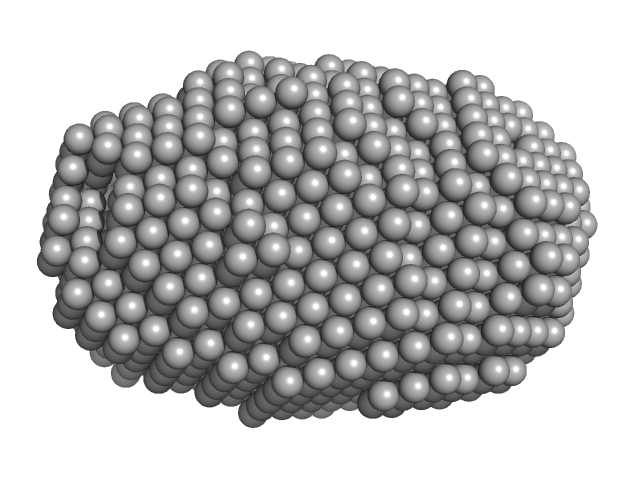
C-terminal catalytic domain of Suppressor of Copper Sensitivity C protein monomer, 20 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
|
| RgGuinier |
1.7 |
nm |
| Dmax |
5.4 |
nm |
| VolumePorod |
22200 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
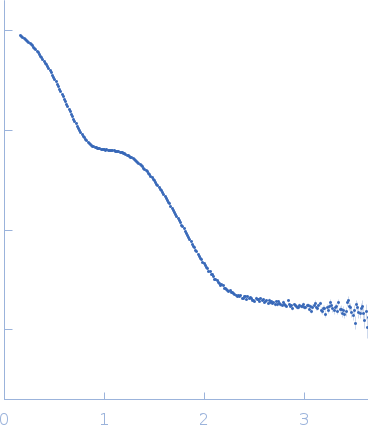
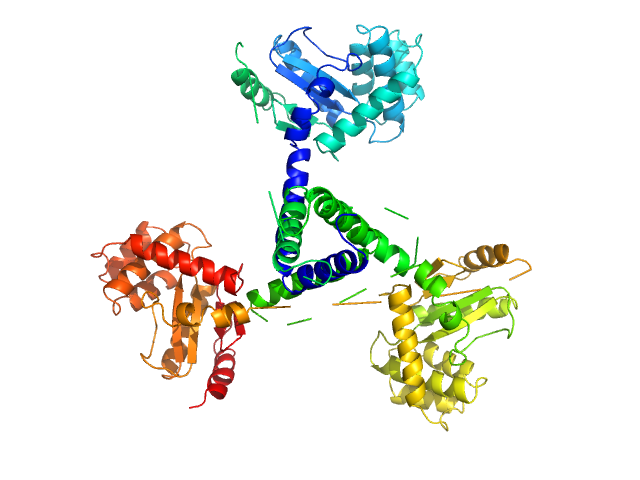
Suppressor of Copper Sensitivity C protein trimer, 74 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
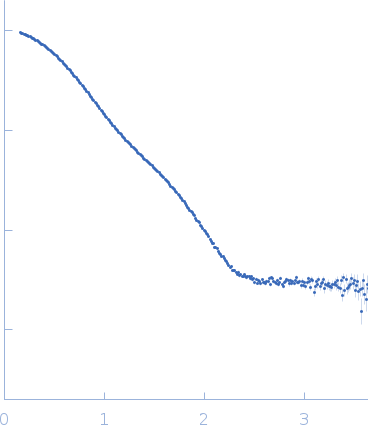
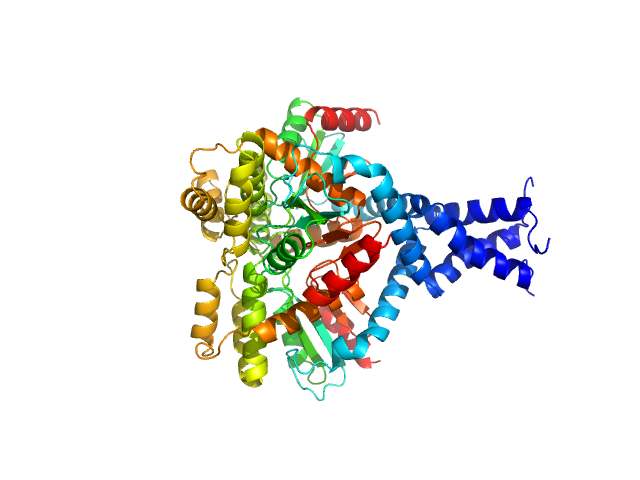
Deletion mutant of PmScsC, 23 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
54 |
nm3 |
|
|