|
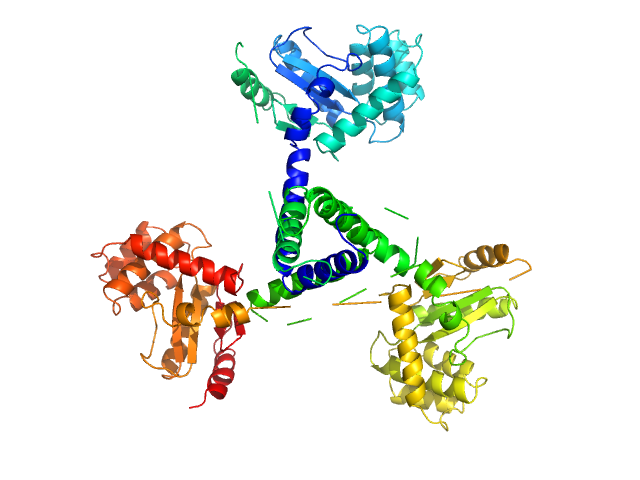
X-ray synchrotron radiation scattering data from solutions of Suppressor of Copper Sensitivity C protein (ScsC) protein from Proteus mirabilis in 10 mM HEPES 150mM NaCl, pH 7.4 were collected on the SAXS/WAXS beam line of the Australian Synchrotron (Melbourne, Australia) using a 2D Photon counting Pilatus 1M-W pixel detector (s = 4π sin θ/λ, where 2θ is the scattering angle). Thirty eight successive 1 second frames were collected across solute concentrations of 0.1-10.0 mg/ml at a sample temperature of 10°C. The data were normalized to the intensity of the transmitted beam and radially averaged and the scattering of the solvent-blank was subtracted. The SAXS data displayed this entry was derived from a 10.0 mg/ml sample and show that essentially only trimer is present in solution. A linear combination of monomer and trimer are fit to the scattering curve, and the final concentration of each species in solution was determined to be: 0.08 µM (monomer); 95.05 µM (trimer). Together, with data collected at other concentration (available to download as a zip archive), it was estimated that K1 (3monomer <-> trimer) was 6.3 nM^2.
|
|
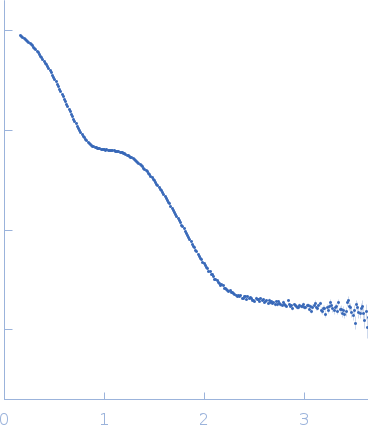 s, nm-1
s, nm-1