|
|
|
|
|
| Sample: |
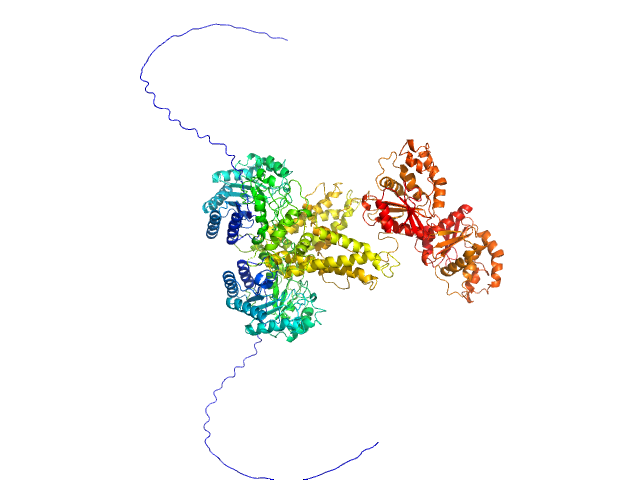
Protein O-GlcNAcase dimer, 163 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris buffer, 150 mM NaCl, and 0.5 mM TCEP, pH: 7.5
|
| Experiment: |
SAXS
data collected at Bruker Nanostar w Excillum source, Department of Chemistry, iNANO building, Aarhus Uinversity on 2023 Jun 22
|
Multi-domain O-GlcNAcase structures reveal allosteric regulatory mechanisms
Jan Skov Pedersen
|
| RgGuinier |
5.4 |
nm |
| Dmax |
18.9 |
nm |
| VolumePorod |
443 |
nm3 |
|