|
|
|
|
|
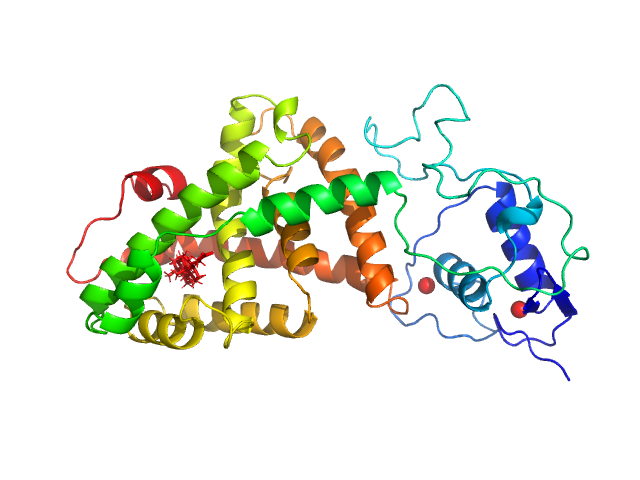
| Sample: |
Isoform 3 of Bile acid receptor monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 5 % glycerol, 1 mM DTT, 20 µM CDCA, pH: 7.4
|
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Pennsylvania State University on 2024 May 9
|
DNA induces non-canonical dimerization of the farnesoid X receptor.
Nucleic Acids Res 54(4) (2026)
...Yennawar N, Okafor CD
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
75 |
nm3 |
|