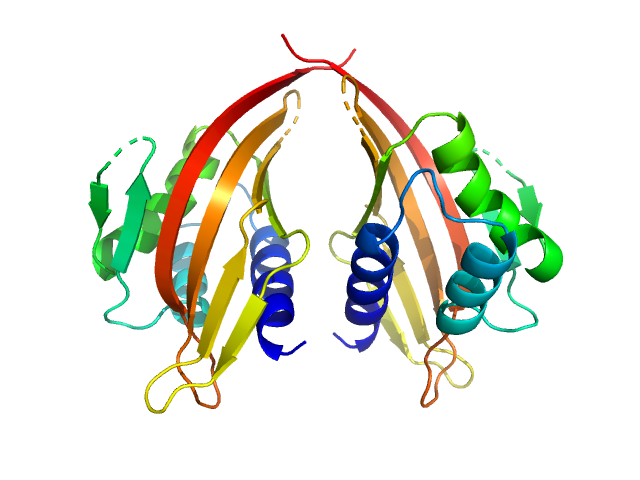
UniProt ID: P70662-3 (14-200) LIM domain-binding protein 1
|
|
|
|
| Sample: |
LIM domain-binding protein 1 dimer, 45 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP,, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 19
|
Self association of LIM domain binding protein 1, LDB1
Jacqueline Matthews
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
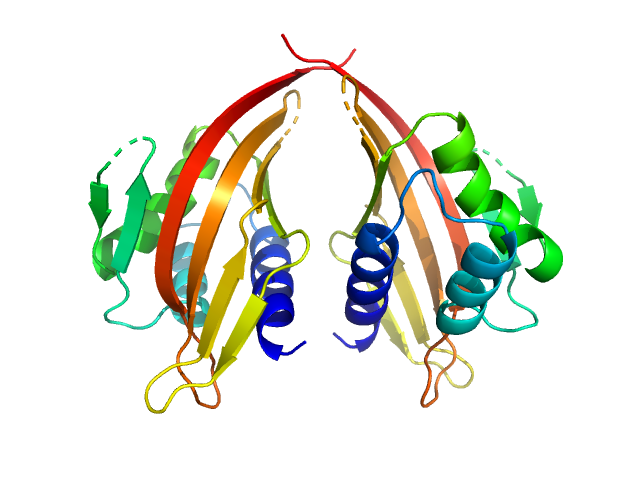
UniProt ID: P70662-3 (14-200) LIM domain-binding protein 1, L87E
|
|
|
|
| Sample: |
LIM domain-binding protein 1, L87E dimer, 45 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP,, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 19
|
Self association of LIM domain binding protein 1, LDB1
Jacqueline Matthews
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
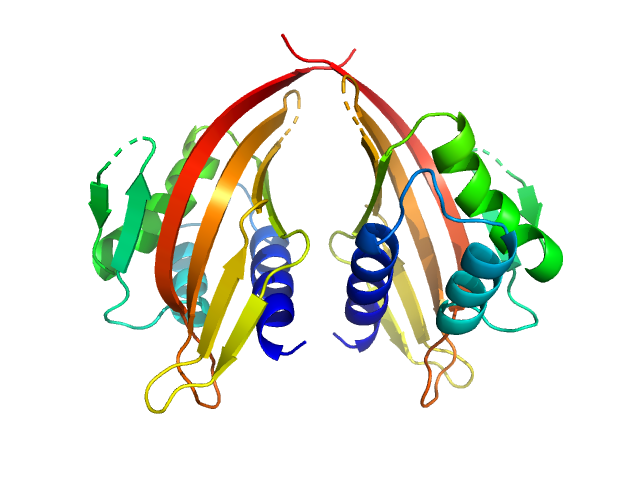
UniProt ID: P70662-3 (14-200) LIM domain-binding protein 1, L87K
|
|
|
|
| Sample: |
LIM domain-binding protein 1, L87K dimer, 45 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP,, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 19
|
Self association of LIM domain binding protein 1, LDB1
Jacqueline Matthews
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
78 |
nm3 |
|
|
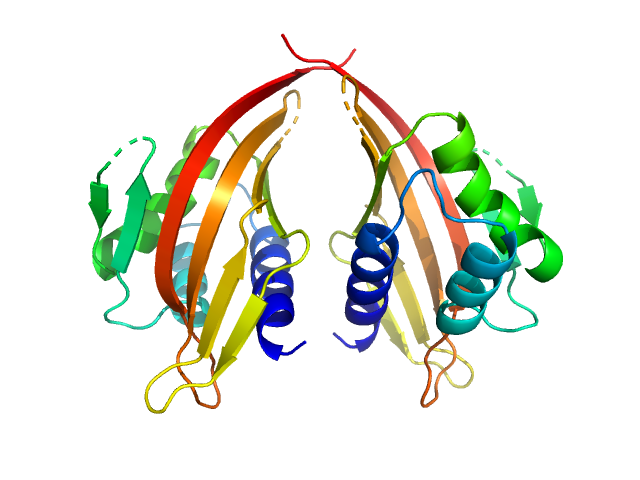
UniProt ID: P70662-3 (14-200) LIM domain-binding protein 1, L87Q
|
|
|
|
| Sample: |
LIM domain-binding protein 1, L87Q dimer, 45 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP,, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 19
|
Self association of LIM domain binding protein 1, LDB1
Jacqueline Matthews
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
UniProt ID: C6KFA3 (39-839) Adhesion G-protein coupled receptor G6 S2
|
|
|
|
| Sample: |
Adhesion G-protein coupled receptor G6 S2 monomer, 86 kDa Danio rerio protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jun 28
|
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications 11(1) (2020)
Leon K, Cunningham R, Riback J, Feldman E, Li J, Sosnick T, Zhao M, Monk K, Araç D
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
169 |
nm3 |
|
|
UniProt ID: C6KFA3 (39-839) Adhesion G-protein coupled receptor G6 S1
|
|
|
|
| Sample: |
Adhesion G-protein coupled receptor G6 S1 monomer, 89 kDa Danio rerio protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jun 28
|
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications 11(1) (2020)
Leon K, Cunningham R, Riback J, Feldman E, Li J, Sosnick T, Zhao M, Monk K, Araç D
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
191 |
nm3 |
|
|
UniProt ID: C6KFA3 (39-839) Adhesion G-protein coupled receptor G6 S2 D134A/F135A
|
|
|
|
| Sample: |
Adhesion G-protein coupled receptor G6 S2 D134A/F135A monomer, 86 kDa Danio rerio protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jun 28
|
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications 11(1) (2020)
Leon K, Cunningham R, Riback J, Feldman E, Li J, Sosnick T, Zhao M, Monk K, Araç D
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
181 |
nm3 |
|
|
UniProt ID: Q86SQ4 (41-852) Adhesion G-protein coupled receptor G6 S2
|
|
|
|
| Sample: |
Adhesion G-protein coupled receptor G6 S2 monomer, 88 kDa Homo sapiens protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jun 28
|
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications 11(1) (2020)
Leon K, Cunningham R, Riback J, Feldman E, Li J, Sosnick T, Zhao M, Monk K, Araç D
|
| RgGuinier |
4.4 |
nm |
| Dmax |
15.7 |
nm |
| VolumePorod |
199 |
nm3 |
|
|
UniProt ID: Q86SQ4 (41-852) Adhesion G-protein coupled receptor G6 S1
|
|
|
|
| Sample: |
Adhesion G-protein coupled receptor G6 S1 monomer, 91 kDa Homo sapiens protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Jun 28
|
Structural basis for adhesion G protein-coupled receptor Gpr126 function
Nature Communications 11(1) (2020)
Leon K, Cunningham R, Riback J, Feldman E, Li J, Sosnick T, Zhao M, Monk K, Araç D
|
| RgGuinier |
4.9 |
nm |
| Dmax |
17.1 |
nm |
| VolumePorod |
213 |
nm3 |
|
|
UniProt ID: Q07617 (1-926) Sperm-associated antigen 1
|
|
|
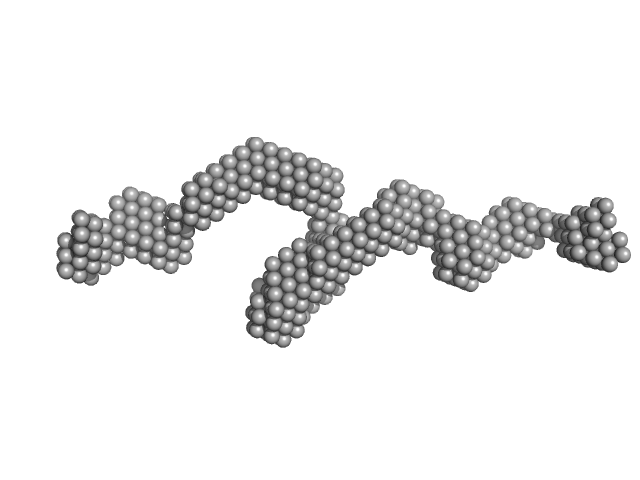
|
| Sample: |
Sperm-associated antigen 1 monomer, 104 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1% glycerol, 5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 21
|
R2SP
Raphael Dos Santos Morais
|
| RgGuinier |
7.3 |
nm |
| Dmax |
34.0 |
nm |
| VolumePorod |
374 |
nm3 |
|
|