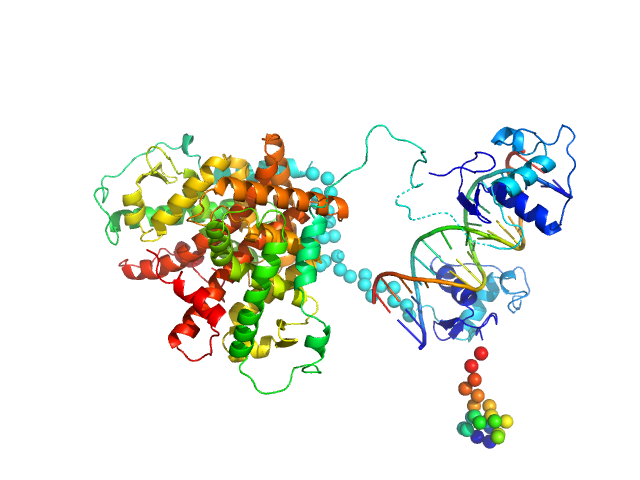
UniProt ID: P11416 (84-421) Retinoic acid receptor alpha, RAR
UniProt ID: P28700 (135-467) Retinoic acid receptor RXR-alpha
UniProt ID: None (None-None) DNA response element HoxB13 DR0
|
|
|
|
| Sample: |
Retinoic acid receptor alpha, RAR monomer, 41 kDa Mus musculus protein
Retinoic acid receptor RXR-alpha monomer, 38 kDa Mus musculus protein
DNA response element HoxB13 DR0 monomer, 10 kDa DNA
|
| Buffer: |
20 mM Tris, pH 8, 150 mM NaCl, 5% v/v glycerol, 1 mM CHAPS, 4 mM MgSO4, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2014 Jan 19
|
Structural basis for DNA recognition and allosteric control of the retinoic acid receptors RAR–RXR
Nucleic Acids Research (2020)
Osz J, McEwen A, Bourguet M, Przybilla F, Peluso-Iltis C, Poussin-Courmontagne P, Mély Y, Cianférani S, Jeffries C, Svergun D, Rochel N
|
| RgGuinier |
3.8 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
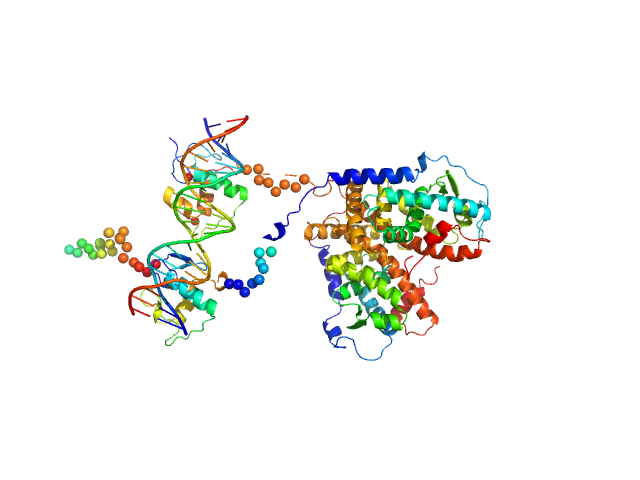
UniProt ID: P11416 (84-421) Retinoic acid receptor alpha, RAR
UniProt ID: P28700 (135-467) Retinoic acid receptor RXR-alpha
UniProt ID: None (None-None) DNA response element F11r DR5
|
|
|
|
| Sample: |
Retinoic acid receptor alpha, RAR monomer, 41 kDa Mus musculus protein
Retinoic acid receptor RXR-alpha monomer, 38 kDa Mus musculus protein
DNA response element F11r DR5 monomer, 13 kDa DNA
|
| Buffer: |
20 mM Tris, pH 8, 150 mM NaCl, 5% v/v glycerol, 1 mM CHAPS, 4 mM MgSO4, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2014 Jan 19
|
Structural basis for DNA recognition and allosteric control of the retinoic acid receptors RAR–RXR
Nucleic Acids Research (2020)
Osz J, McEwen A, Bourguet M, Przybilla F, Peluso-Iltis C, Poussin-Courmontagne P, Mély Y, Cianférani S, Jeffries C, Svergun D, Rochel N
|
| RgGuinier |
4.0 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
130 |
nm3 |
|
|
UniProt ID: P02649 (19-317) Apolipoprotein E2
UniProt ID: None (None-None) Heparin
|
|
|
|
| Sample: |
Apolipoprotein E2 tetramer, 139 kDa Homo sapiens protein
Heparin monomer, 15 kDa
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 3
|
Therapeutic approaches to ApoE
University of Sussex PhD thesis 2019 (2019)
Lucas Kraft
|
| RgGuinier |
6.2 |
nm |
| Dmax |
20.7 |
nm |
| VolumePorod |
504 |
nm3 |
|
|
UniProt ID: P02649 (19-317) Apolipoprotein E2
UniProt ID: None (None-None) Heparin
|
|
|
|
| Sample: |
Apolipoprotein E2 tetramer, 139 kDa Homo sapiens protein
Heparin monomer, 15 kDa
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 3
|
Therapeutic approaches to ApoE
University of Sussex PhD thesis 2019 (2019)
Lucas Kraft
|
| RgGuinier |
6.4 |
nm |
| Dmax |
21.7 |
nm |
| VolumePorod |
531 |
nm3 |
|
|
UniProt ID: P02649 (19-317) Apolipoprotein E2
UniProt ID: None (None-None) Heparin
|
|
|
|
| Sample: |
Apolipoprotein E2 tetramer, 139 kDa Homo sapiens protein
Heparin monomer, 15 kDa
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 3
|
Therapeutic approaches to ApoE
University of Sussex PhD thesis 2019 (2019)
Lucas Kraft
|
| RgGuinier |
6.5 |
nm |
| Dmax |
23.4 |
nm |
| VolumePorod |
584 |
nm3 |
|
|
UniProt ID: P02649 (19-317) Apolipoprotein E2
UniProt ID: None (None-None) Heparin
|
|
|
|
| Sample: |
Apolipoprotein E2 tetramer, 139 kDa Homo sapiens protein
Heparin monomer, 15 kDa
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 3
|
Therapeutic approaches to ApoE
University of Sussex PhD thesis 2019 (2019)
Lucas Kraft
|
| RgGuinier |
7.1 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
666 |
nm3 |
|
|
UniProt ID: P02649 (19-317) Apolipoprotein E2
UniProt ID: None (None-None) Heparin
|
|
|
|
| Sample: |
Apolipoprotein E2 tetramer, 139 kDa Homo sapiens protein
Heparin monomer, 15 kDa
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 3
|
Therapeutic approaches to ApoE
University of Sussex PhD thesis 2019 (2019)
Lucas Kraft
|
| RgGuinier |
7.2 |
nm |
| Dmax |
24.5 |
nm |
| VolumePorod |
696 |
nm3 |
|
|
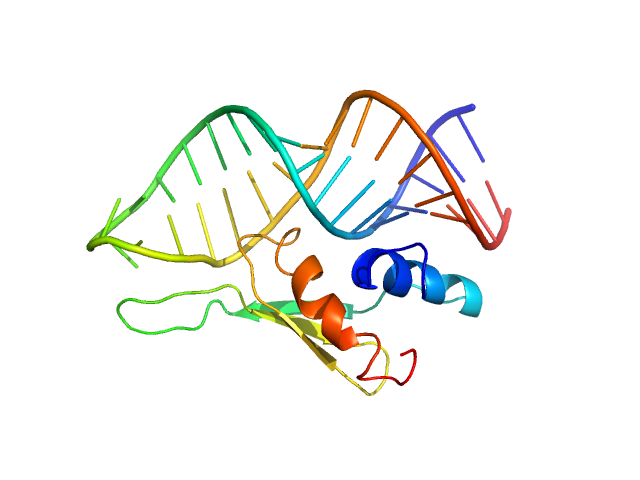
UniProt ID: O95793 (286-355) Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 4
UniProt ID: None (None-None) ADP-ribosylation factor1 - short
|
|
|
|
| Sample: |
Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 4 monomer, 8 kDa Homo sapiens protein
ADP-ribosylation factor1 - short monomer, 11 kDa Homo sapiens RNA
|
| Buffer: |
50 mM potassium phosphate, 100 mM NaCl, 3.5 mM 2-mercaptoethanol, pH: 6.8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, CEITEC on 2015 Dec 10
|
Staufen1 reads out structure and sequence features in ARF1 dsRNA for target recognition.
Nucleic Acids Res (2019)
Yadav DK, Zigáčková D, Zlobina M, Klumpler T, Beaumont C, Kubíčková M, Vaňáčová Š, Lukavsky PJ
|
| RgGuinier |
1.7 |
nm |
| Dmax |
6.0 |
nm |
| VolumePorod |
21 |
nm3 |
|
|
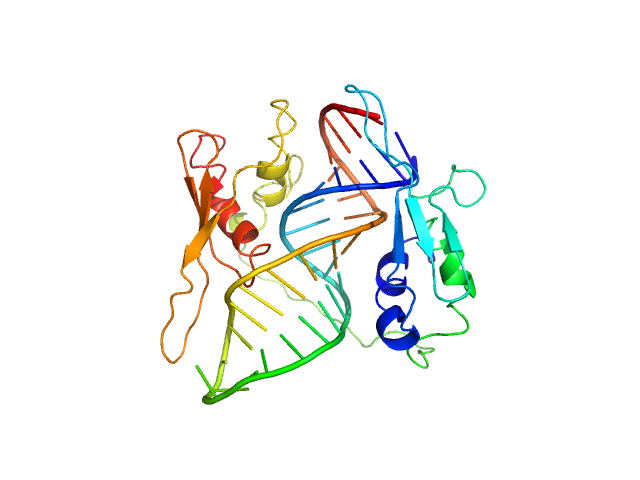
UniProt ID: None (None-None) ADP-ribosylation factor1 - short
UniProt ID: O95793 (183-355) Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 3 and 4
|
|
|
|
| Sample: |
ADP-ribosylation factor1 - short monomer, 11 kDa Homo sapiens RNA
Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 3 and 4 monomer, 20 kDa Homo sapiens protein
|
| Buffer: |
50 mM sodium phosphate buffer, 300 mM NaCl, 500 mM imidazole, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, CEITEC on 2018 May 29
|
Staufen1 reads out structure and sequence features in ARF1 dsRNA for target recognition.
Nucleic Acids Res (2019)
Yadav DK, Zigáčková D, Zlobina M, Klumpler T, Beaumont C, Kubíčková M, Vaňáčová Š, Lukavsky PJ
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.4 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
UniProt ID: O95793 (183-355) Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 3 and 4
UniProt ID: None (None-None) ADP-ribosylation factor1 - long
|
|
|
|
| Sample: |
Double-stranded RNA-binding protein Staufen homolog 1 - RNA binding domain 3 and 4 monomer, 20 kDa Homo sapiens protein
ADP-ribosylation factor1 - long monomer, 16 kDa Homo sapiens RNA
|
| Buffer: |
50 mM potassium phosphate, 100 mM NaCl, 3.5 mM 2-mercaptoethanol, pH: 6.8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 Sep 20
|
Staufen1 reads out structure and sequence features in ARF1 dsRNA for target recognition.
Nucleic Acids Res (2019)
Yadav DK, Zigáčková D, Zlobina M, Klumpler T, Beaumont C, Kubíčková M, Vaňáčová Š, Lukavsky PJ
|
| RgGuinier |
2.5 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
53 |
nm3 |
|
|