UniProt ID: Q8U4M2 (1-227) Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase
UniProt ID: Q8U160 (1-123) 50S ribosomal protein L7Ae
UniProt ID: Q8U4M1 (1-402) NOP5/NOP56 related protein
UniProt ID: None (None-None) pyrococcus furiosus sR26 stabilized construct
UniProt ID: None (None-None) Pyrococcus furiosus sR26 substrate D'
|
|
|
|
| Sample: |
Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase dimer, 52 kDa Pyrococcus furiosus protein
50S ribosomal protein L7Ae dimer, 27 kDa Pyrococcus furiosus protein
NOP5/NOP56 related protein dimer, 94 kDa Pyrococcus furiosus protein
Pyrococcus furiosus sR26 stabilized construct monomer, 24 kDa Pyrococcus furiosus RNA
Pyrococcus furiosus sR26 substrate D' monomer, 4 kDa Pyrococcus furiosus RNA
|
| Buffer: |
50 mM phosphate 500 mM NaCl, pH: 6.6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jun 13
|
The guide sRNA sequence determines the activity level of box C/D RNPs.
Elife 9 (2020)
Graziadei A, Gabel F, Kirkpatrick J, Carlomagno T
|
| RgGuinier |
4.7 |
nm |
| Dmax |
15.8 |
nm |
| VolumePorod |
346 |
nm3 |
|
|
UniProt ID: Q8U4M2 (1-227) Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase
UniProt ID: Q8U160 (1-123) 50S ribosomal protein L7Ae
UniProt ID: Q8U4M1 (1-402) NOP5/NOP56 related protein
UniProt ID: None (None-None) pyrococcus furiosus sR26 stabilized construct
UniProt ID: None (None-None) Pyrococcus furiosus sR26 substrate D
|
|
|
|
| Sample: |
Fibrillarin-like rRNA/tRNA 2'-O-methyltransferase dimer, 52 kDa Pyrococcus furiosus protein
50S ribosomal protein L7Ae dimer, 27 kDa Pyrococcus furiosus protein
NOP5/NOP56 related protein dimer, 94 kDa Pyrococcus furiosus protein
Pyrococcus furiosus sR26 stabilized construct monomer, 24 kDa Pyrococcus furiosus RNA
Pyrococcus furiosus sR26 substrate D monomer, 4 kDa Pyrococcus furiosus RNA
|
| Buffer: |
50 mM phosphate 500 mM NaCl, pH: 6.6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jul 13
|
The guide sRNA sequence determines the activity level of box C/D RNPs.
Elife 9 (2020)
Graziadei A, Gabel F, Kirkpatrick J, Carlomagno T
|
| RgGuinier |
5.0 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
429 |
nm3 |
|
|
UniProt ID: P04501 (130-318) Human adenovirus serotype 3 fibre knob
UniProt ID: Q14126 (157-380) Human desmoglein 2 extracellular domain 2 and 3
|
|
|
|
| Sample: |
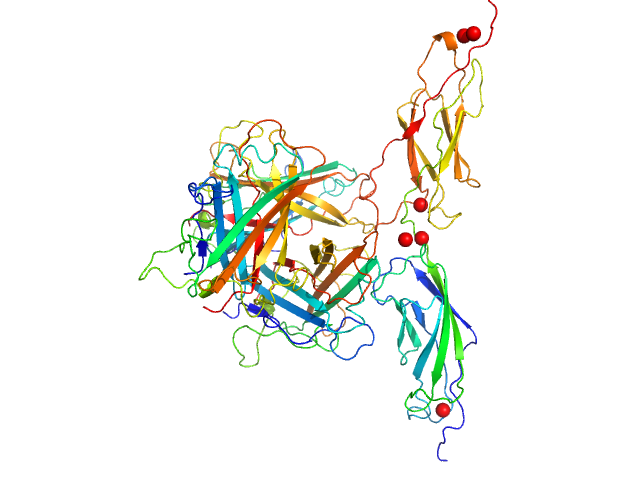
Human adenovirus serotype 3 fibre knob trimer, 69 kDa Human adenovirus serotype … protein
Human desmoglein 2 extracellular domain 2 and 3 dimer, 55 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, 3 mM CaCl2, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Jan 26
|
Intermediate-resolution crystal structure of the human adenovirus B serotype 3 fibre knob in complex with the EC2-EC3 fragment of desmoglein 2.
Acta Crystallogr F Struct Biol Commun 75(Pt 12):750-757 (2019)
Vassal-Stermann E, Hutin S, Fender P, Burmeister WP
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.2 |
nm |
| VolumePorod |
188 |
nm3 |
|
|
UniProt ID: P22897 (19-1388) Macrophage mannose receptor 1
|
|
|
|
| Sample: |
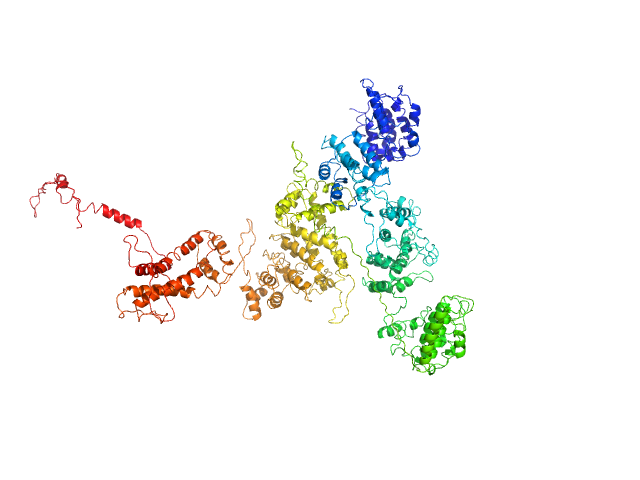
Macrophage mannose receptor 1 dimer, 315 kDa Mouse myeloma cell … protein
|
| Buffer: |
50mM Hepes, 100mM NaCl, 1mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Apr 15
|
Mannose receptor (CD206) activation in tumor-associated macrophages enhances adaptive and innate antitumor immune responses.
Sci Transl Med 12(530) (2020)
Jaynes JM, Sable R, Ronzetti M, Bautista W, Knotts Z, Abisoye-Ogunniyan A, Li D, Calvo R, Dashnyam M, Singh A, Guerin T, White J, Ravichandran S, Kumar P, Talsania K, Chen V, Ghebremedhin A, Karanam B, Bin Salam A, Amin R, Odzorig T, Aiken T, Nguyen V, Bian Y, Zarif JC, de Groot AE, Mehta M, Fan L, Hu X, Simeonov A, Pate N, Abu-Asab M, Ferrer M, Southall N, Ock CY, Zhao Y, Lopez H, Kozlov S, de Val N, Yates CC, Baljinnyam B, Marugan J, Rudloff U
|
| RgGuinier |
7.9 |
nm |
| Dmax |
30.1 |
nm |
| VolumePorod |
584 |
nm3 |
|
|
UniProt ID: Q15291 (2-381) Retinoblastoma-binding protein 5
UniProt ID: Q03164 (3754-3969) Histone-lysine N-methyltransferase 2A
UniProt ID: P61964 (23-334) WD repeat-containing protein 5
UniProt ID: Q9UBL3-3 (1-534) Set1/Ash2 histone methyltransferase complex subunit ASH2
|
|
|
|
| Sample: |
Retinoblastoma-binding protein 5 monomer, 42 kDa Homo sapiens protein
Histone-lysine N-methyltransferase 2A monomer, 25 kDa Homo sapiens protein
WD repeat-containing protein 5 monomer, 34 kDa Homo sapiens protein
Set1/Ash2 histone methyltransferase complex subunit ASH2 monomer, 60 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 25mM Tris-HCl, 4% glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2019 Jun 22
|
The internal interaction in RBBP5 regulates assembly and activity of MLL1 methyltransferase complex.
Nucleic Acids Res (2019)
Han J, Li T, Li Y, Li M, Wang X, Peng C, Su C, Li N, Li Y, Xu Y, Chen Y
|
| RgGuinier |
5.7 |
nm |
| Dmax |
18.6 |
nm |
| VolumePorod |
360 |
nm3 |
|
|
UniProt ID: Q03164 (3754-3969) Histone-lysine N-methyltransferase 2A
UniProt ID: P61964 (23-334) WD repeat-containing protein 5
UniProt ID: Q9UBL3-3 (1-534) Set1/Ash2 histone methyltransferase complex subunit ASH2
UniProt ID: Q15291 (2-480) Retinoblastoma-binding protein 5
|
|
|
|
| Sample: |
Histone-lysine N-methyltransferase 2A monomer, 25 kDa Homo sapiens protein
WD repeat-containing protein 5 monomer, 34 kDa Homo sapiens protein
Set1/Ash2 histone methyltransferase complex subunit ASH2 monomer, 60 kDa Homo sapiens protein
Retinoblastoma-binding protein 5 monomer, 53 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 25mM Tris-HCl, 4% glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2019 Jun 22
|
The internal interaction in RBBP5 regulates assembly and activity of MLL1 methyltransferase complex.
Nucleic Acids Res (2019)
Han J, Li T, Li Y, Li M, Wang X, Peng C, Su C, Li N, Li Y, Xu Y, Chen Y
|
| RgGuinier |
5.0 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
256 |
nm3 |
|
|
UniProt ID: Q03164 (3754-3969) Histone-lysine N-methyltransferase 2A
UniProt ID: P61964 (23-334) WD repeat-containing protein 5
UniProt ID: Q9UBL3-3 (1-534) Set1/Ash2 histone methyltransferase complex subunit ASH2
UniProt ID: Q15291 (2-480) Retinoblastoma-binding protein 5
|
|
|
|
| Sample: |
Histone-lysine N-methyltransferase 2A monomer, 25 kDa Homo sapiens protein
WD repeat-containing protein 5 monomer, 34 kDa Homo sapiens protein
Set1/Ash2 histone methyltransferase complex subunit ASH2 monomer, 60 kDa Homo sapiens protein
Retinoblastoma-binding protein 5 monomer, 53 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 25mM Tris-HCl, 4% glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2019 Jun 22
|
The internal interaction in RBBP5 regulates assembly and activity of MLL1 methyltransferase complex.
Nucleic Acids Res (2019)
Han J, Li T, Li Y, Li M, Wang X, Peng C, Su C, Li N, Li Y, Xu Y, Chen Y
|
| RgGuinier |
5.3 |
nm |
| Dmax |
17.2 |
nm |
| VolumePorod |
313 |
nm3 |
|
|
UniProt ID: Q03164 (3754-3969) Histone-lysine N-methyltransferase 2A
UniProt ID: P61964 (23-334) WD repeat-containing protein 5
UniProt ID: Q9UBL3-3 (1-534) Set1/Ash2 histone methyltransferase complex subunit ASH2
UniProt ID: Q15291 (2-538) Retinoblastoma-binding protein 5
|
|
|
|
| Sample: |
Histone-lysine N-methyltransferase 2A monomer, 25 kDa Homo sapiens protein
WD repeat-containing protein 5 monomer, 34 kDa Homo sapiens protein
Set1/Ash2 histone methyltransferase complex subunit ASH2 monomer, 60 kDa Homo sapiens protein
Retinoblastoma-binding protein 5 monomer, 59 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 25mM Tris-HCl, 4% glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2019 Jun 22
|
The internal interaction in RBBP5 regulates assembly and activity of MLL1 methyltransferase complex.
Nucleic Acids Res (2019)
Han J, Li T, Li Y, Li M, Wang X, Peng C, Su C, Li N, Li Y, Xu Y, Chen Y
|
| RgGuinier |
5.0 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
282 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
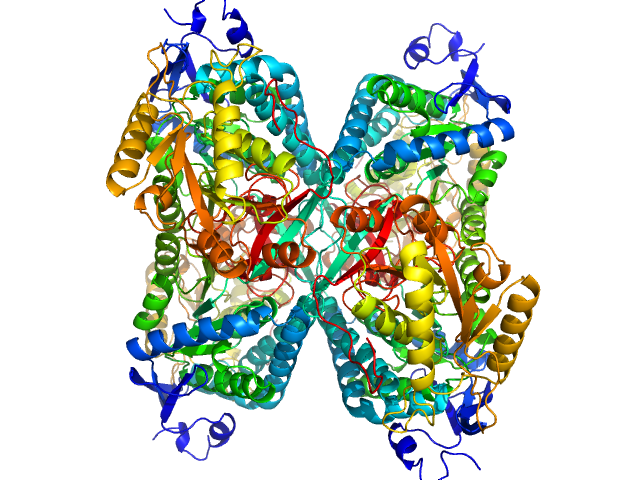
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 1 mM DTT, 10 mM NAD, 2% (v/v) glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 1
|
Structural Analysis of Pathogenic Mutations Targeting Glu427 of ALDH7A1, the Hot Spot Residue of Pyridoxine-Dependent Epilepsy.
J Inherit Metab Dis (2019)
Laciak AR, Korasick DA, Gates KS, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
350 |
nm3 |
|
|
UniProt ID: P49419 (None-None) Alpha-aminoadipic semialdehyde dehydrogenase
|
|
|
|
| Sample: |
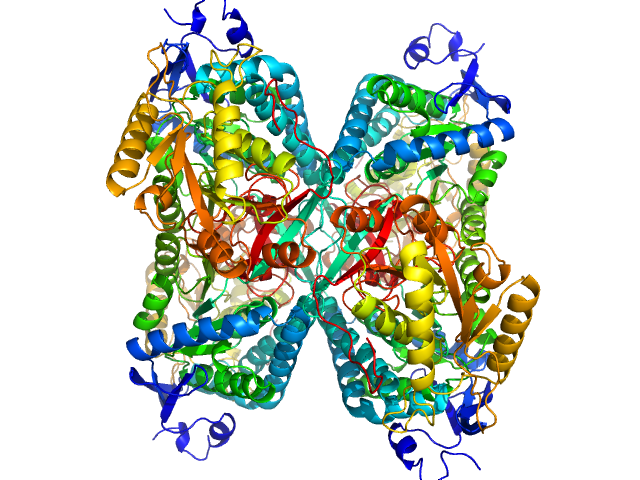
Alpha-aminoadipic semialdehyde dehydrogenase tetramer, 222 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 1 mM DTT, 10 mM NAD, 2% (v/v) glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 1
|
Structural Analysis of Pathogenic Mutations Targeting Glu427 of ALDH7A1, the Hot Spot Residue of Pyridoxine-Dependent Epilepsy.
J Inherit Metab Dis (2019)
Laciak AR, Korasick DA, Gates KS, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
326 |
nm3 |
|
|