UniProt ID: Q14457 (1-150) Beclin-1
|
|
|
|
| Sample: |
Beclin-1 monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 300 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Feb 12
|
Invariant BECN1 CXXC motifs bind Zn(2+) and regulate structure and function of the BECN1 intrinsically disordered region.
Autophagy :1-17 (2023)
Mukhopadhyay S, Subedi S, Hopkins JB, Ugrinov A, Chakravarthy S, Colbert CL, Sinha SC
|
| RgGuinier |
4.3 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
UniProt ID: Q14457 (1-150) Beclin-1
|
|
|
|
| Sample: |
Beclin-1 monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris, 300 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Feb 12
|
Invariant BECN1 CXXC motifs bind Zn(2+) and regulate structure and function of the BECN1 intrinsically disordered region.
Autophagy :1-17 (2023)
Mukhopadhyay S, Subedi S, Hopkins JB, Ugrinov A, Chakravarthy S, Colbert CL, Sinha SC
|
| RgGuinier |
2.8 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
49 |
nm3 |
|
|
UniProt ID: Q05586 (1-847) Glutamate receptor ionotropic, NMDA 1
UniProt ID: Q12879 (1-842) Glutamate receptor ionotropic, NMDA 2A
|
|
|
|
| Sample: |
Glutamate receptor ionotropic, NMDA 1 dimer, 193 kDa Homo sapiens protein
Glutamate receptor ionotropic, NMDA 2A dimer, 191 kDa Homo sapiens protein
|
| Buffer: |
150 mM NaCl, 0.1% digitonin, 5 µM Cholesteryl Hemisuccinate TRIS Salt, 0.1 mM CHAPSO, 50 µM EDTA,1 mM Gly/Glu, 20 mM HEPES, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2022 Dec 8
|
Structural basis for antibody-mediated NMDA receptor clustering and endocytosis in autoimmune encephalitis.
Nat Struct Mol Biol (2024)
Wang H, Xie C, Deng B, Ding J, Li N, Kou Z, Jin M, He J, Wang Q, Wen H, Zhang J, Zhou Q, Chen S, Chen X, Yuan TF, Zhu S
|
| RgGuinier |
6.6 |
nm |
| Dmax |
20.4 |
nm |
| VolumePorod |
1180 |
nm3 |
|
|
UniProt ID: Q05586 (1-847) Glutamate receptor ionotropic, NMDA 1
UniProt ID: Q12879 (1-842) Glutamate receptor ionotropic, NMDA 2A
UniProt ID: None (None-None) Human derived autoantibody mAb2G7 heavy chain, mAb2G7 VH
UniProt ID: None (None-None) Human derived autoantibody mAb2G7 light chain, mAb2G7 VL
|
|
|
|
| Sample: |
Glutamate receptor ionotropic, NMDA 1 dimer, 193 kDa Homo sapiens protein
Glutamate receptor ionotropic, NMDA 2A dimer, 191 kDa Homo sapiens protein
Human derived autoantibody mAb2G7 heavy chain, mAb2G7 VH dimer, 103 kDa protein
Human derived autoantibody mAb2G7 light chain, mAb2G7 VL dimer, 51 kDa protein
|
| Buffer: |
150 mM NaCl, 0.1% digitonin, 5 µM Cholesteryl Hemisuccinate TRIS Salt, 0.1 mM CHAPSO, 50 µM EDTA,1 mM Gly/Glu, 20 mM HEPES, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2022 Dec 8
|
Structural basis for antibody-mediated NMDA receptor clustering and endocytosis in autoimmune encephalitis.
Nat Struct Mol Biol (2024)
Wang H, Xie C, Deng B, Ding J, Li N, Kou Z, Jin M, He J, Wang Q, Wen H, Zhang J, Zhou Q, Chen S, Chen X, Yuan TF, Zhu S
|
| RgGuinier |
7.7 |
nm |
| Dmax |
25.4 |
nm |
| VolumePorod |
1260 |
nm3 |
|
|
UniProt ID: Q05586 (1-847) Glutamate receptor ionotropic, NMDA 1
UniProt ID: Q12879 (1-842) Glutamate receptor ionotropic, NMDA 2A
UniProt ID: None (None-None) Human derived autoantibody mAb5F6 heavy chain, mAb5F6 VH
UniProt ID: None (None-None) Human derived autoantibody mAb5F6 light chain, mAb5F6 VL
|
|
|
|
| Sample: |
Glutamate receptor ionotropic, NMDA 1 dimer, 193 kDa Homo sapiens protein
Glutamate receptor ionotropic, NMDA 2A dimer, 191 kDa Homo sapiens protein
Human derived autoantibody mAb5F6 heavy chain, mAb5F6 VH dimer, 104 kDa Homo sapiens protein
Human derived autoantibody mAb5F6 light chain, mAb5F6 VL dimer, 52 kDa protein
|
| Buffer: |
150 mM NaCl, 0.1% digitonin, 5 µM Cholesteryl Hemisuccinate TRIS Salt, 0.1 mM CHAPSO, 50 µM EDTA,1 mM Gly/Glu, 20 mM HEPES, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2022 Dec 8
|
Structural basis for antibody-mediated NMDA receptor clustering and endocytosis in autoimmune encephalitis.
Nat Struct Mol Biol (2024)
Wang H, Xie C, Deng B, Ding J, Li N, Kou Z, Jin M, He J, Wang Q, Wen H, Zhang J, Zhou Q, Chen S, Chen X, Yuan TF, Zhu S
|
| RgGuinier |
9.9 |
nm |
| Dmax |
31.7 |
nm |
| VolumePorod |
2500 |
nm3 |
|
|
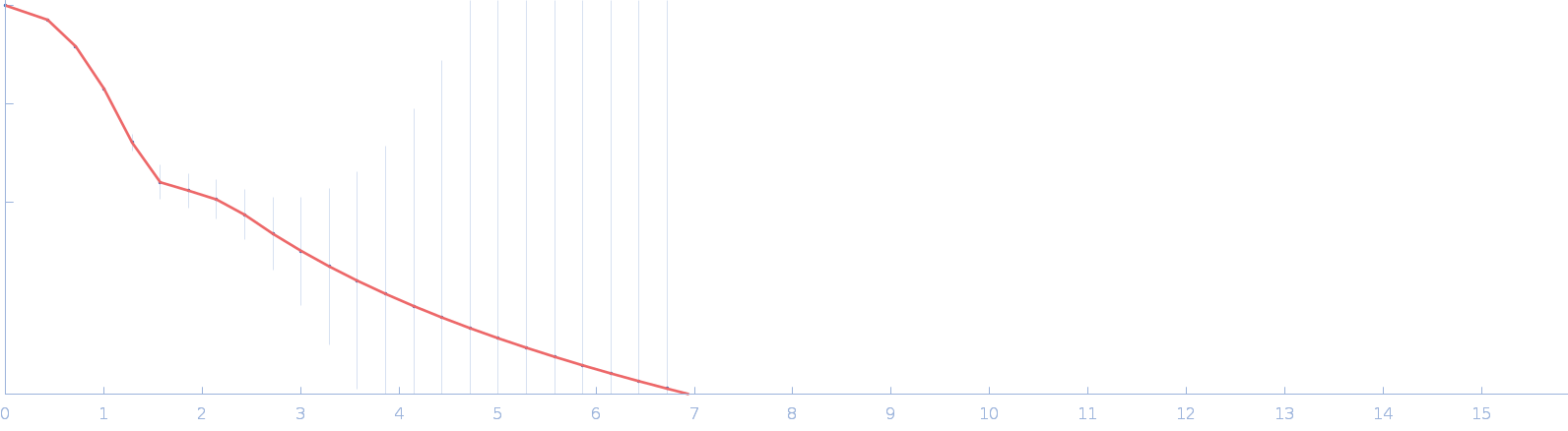
UniProt ID: Q8A622 (41-464) DUF4374 domain-containing protein
|
|
|
|
| Sample: |
DUF4374 domain-containing protein monomer, 46 kDa Bacteroides thetaiotaomicron (strain … protein
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 May 9
|
Iron acquisition by a commensal bacterium modifies host nutritional immunity during Salmonella infection.
Cell Host Microbe 31(10):1639-1654.e10 (2023)
Spiga L, Fansler RT, Perera YR, Shealy NG, Munneke MJ, David HE, Torres TP, Lemoff A, Ran X, Richardson KL, Pudlo N, Martens EC, Folta-Stogniew E, Yang ZJ, Skaar EP, Byndloss MX, Chazin WJ, Zhu W
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
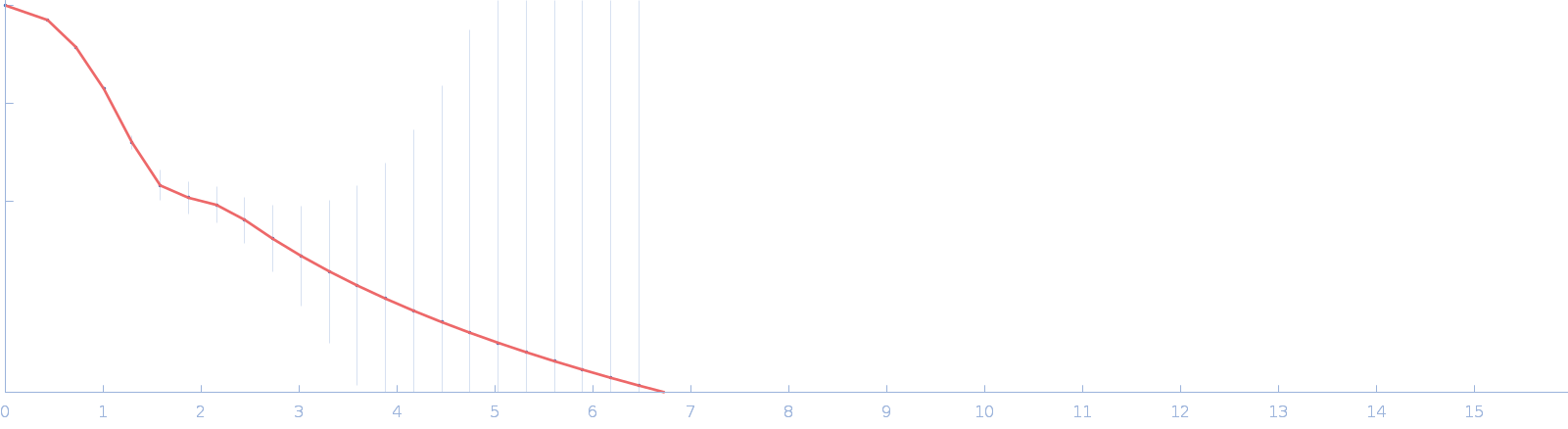
UniProt ID: Q8A622 (41-464) DUF4374 domain-containing protein
|
|
|
|
| Sample: |
DUF4374 domain-containing protein monomer, 46 kDa Bacteroides thetaiotaomicron (strain … protein
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 12-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 May 9
|
Iron acquisition by a commensal bacterium modifies host nutritional immunity during Salmonella infection.
Cell Host Microbe 31(10):1639-1654.e10 (2023)
Spiga L, Fansler RT, Perera YR, Shealy NG, Munneke MJ, David HE, Torres TP, Lemoff A, Ran X, Richardson KL, Pudlo N, Martens EC, Folta-Stogniew E, Yang ZJ, Skaar EP, Byndloss MX, Chazin WJ, Zhu W
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
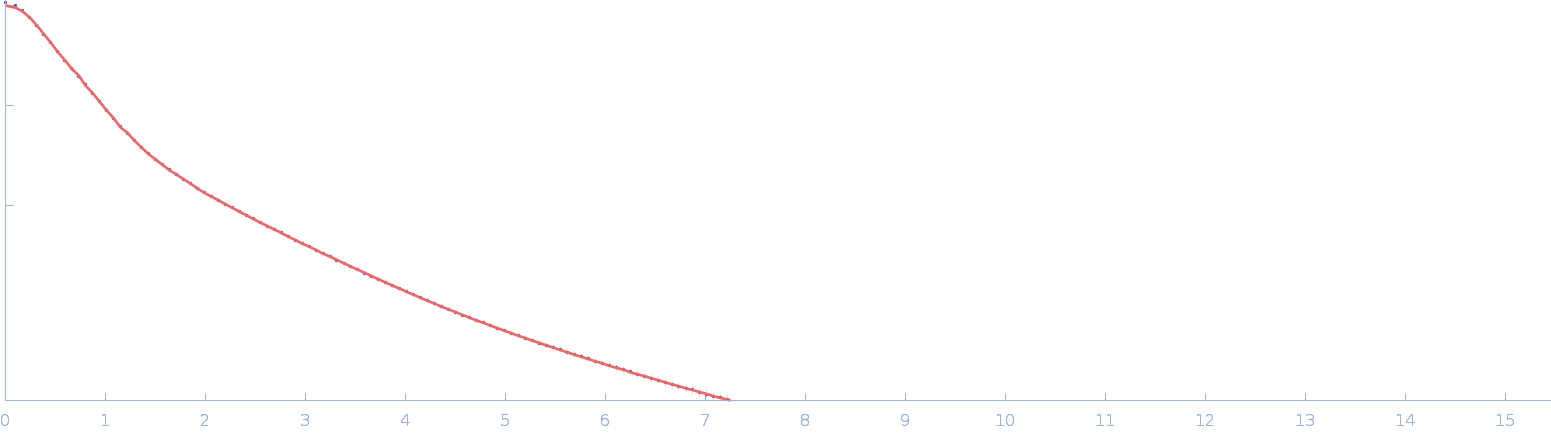
UniProt ID: Q90922 (26-458) Netrin-1
|
|
|
|
| Sample: |
Netrin-1 monomer, 50 kDa Gallus gallus protein
|
| Buffer: |
20 mM Tris-HCl, 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Oct 24
|
The dynamic nature of netrin-1 and the structural basis for glycosaminoglycan fragment-induced filament formation
Nature Communications 14(1) (2023)
Meier M, Gupta M, Akgül S, McDougall M, Imhof T, Nikodemus D, Reuten R, Moya-Torres A, To V, Ferens F, Heide F, Padilla-Meier G, Kukura P, Huang W, Gerisch B, Mörgelin M, Poole K, Antebi A, Koch M, Stetefeld J
|
| RgGuinier |
4.0 |
nm |
| Dmax |
18.0 |
nm |
| VolumePorod |
103 |
nm3 |
|
|
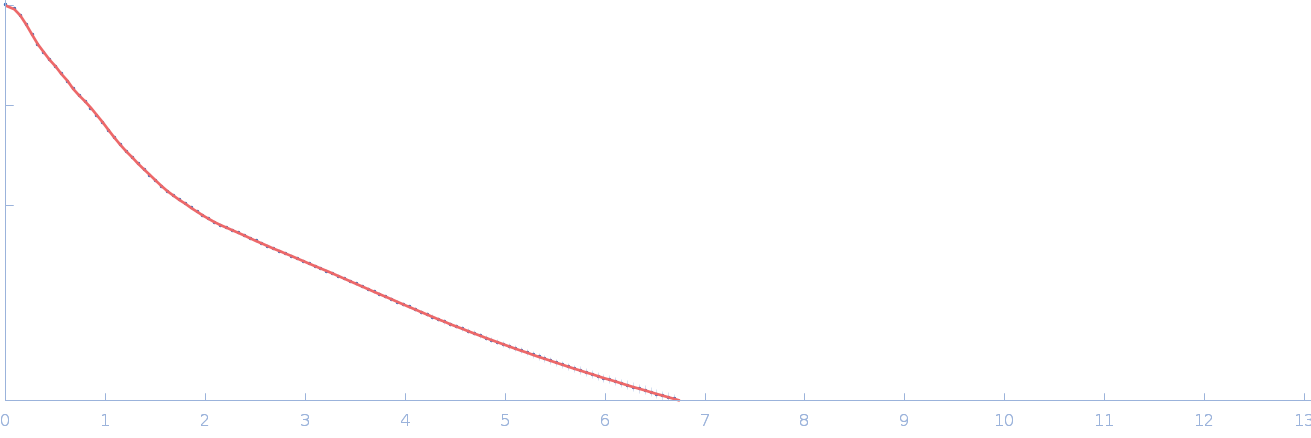
UniProt ID: Q90922 (26-458) Netrin-1
|
|
|
|
| Sample: |
Netrin-1 dimer, 99 kDa Gallus gallus protein
|
| Buffer: |
20 mM Tris-HCl, 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Oct 24
|
The dynamic nature of netrin-1 and the structural basis for glycosaminoglycan fragment-induced filament formation
Nature Communications 14(1) (2023)
Meier M, Gupta M, Akgül S, McDougall M, Imhof T, Nikodemus D, Reuten R, Moya-Torres A, To V, Ferens F, Heide F, Padilla-Meier G, Kukura P, Huang W, Gerisch B, Mörgelin M, Poole K, Antebi A, Koch M, Stetefeld J
|
| RgGuinier |
5.9 |
nm |
| Dmax |
21.3 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
UniProt ID: Q90922 (26-458) Netrin-1
|
|
|
|
| Sample: |
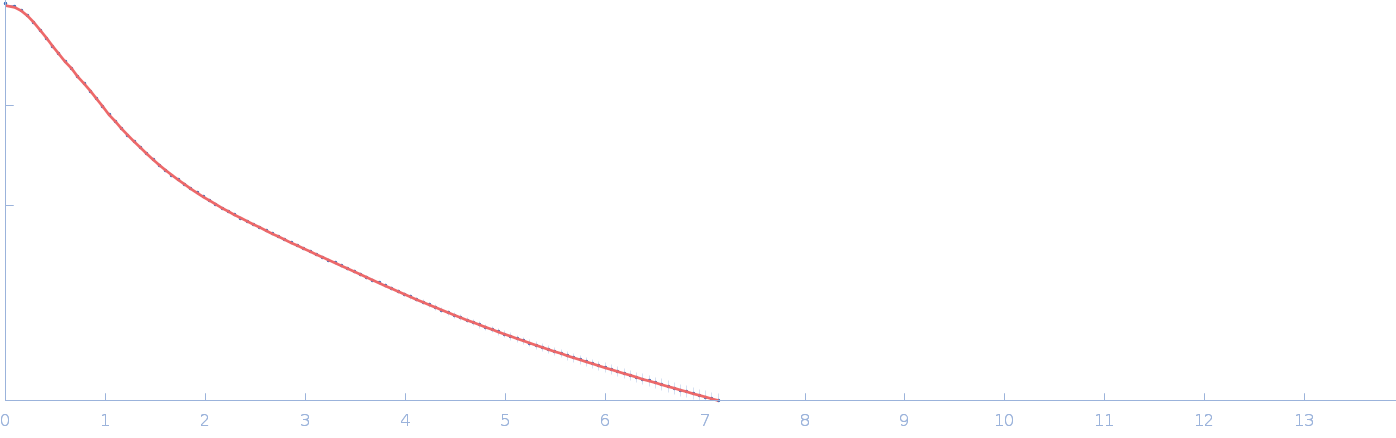
Netrin-1 monomer, 50 kDa Gallus gallus protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Oct 24
|
The dynamic nature of netrin-1 and the structural basis for glycosaminoglycan fragment-induced filament formation
Nature Communications 14(1) (2023)
Meier M, Gupta M, Akgül S, McDougall M, Imhof T, Nikodemus D, Reuten R, Moya-Torres A, To V, Ferens F, Heide F, Padilla-Meier G, Kukura P, Huang W, Gerisch B, Mörgelin M, Poole K, Antebi A, Koch M, Stetefeld J
|
| RgGuinier |
4.1 |
nm |
| Dmax |
21.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|