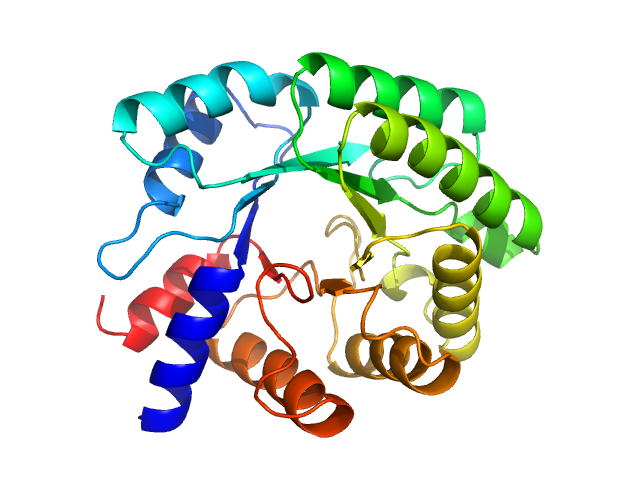
UniProt ID: P0A6L0 (1-259) Deoxyribose-phosphate aldolase
|
|
|
|
| Sample: |
Deoxyribose-phosphate aldolase, 28 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 1
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
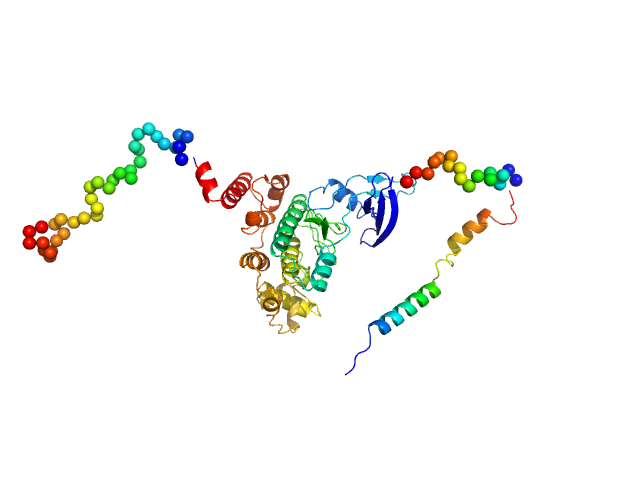
UniProt ID: O77344 (1-440) Glycogen synthase kinase 3
|
|
|
|
| Sample: |
Glycogen synthase kinase 3 monomer, 52 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
20 mM Tris pH 8.0, 100 mM NaCl, 0.5 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Nov 19
|
N-terminal phosphorylation regulates the activity of glycogen synthase kinase 3 from Plasmodium falciparum.
Biochem J 479(3):337-356 (2022)
Pazicky S, Alder A, Mertens H, Svergun D, Gilberger T, Löw C
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
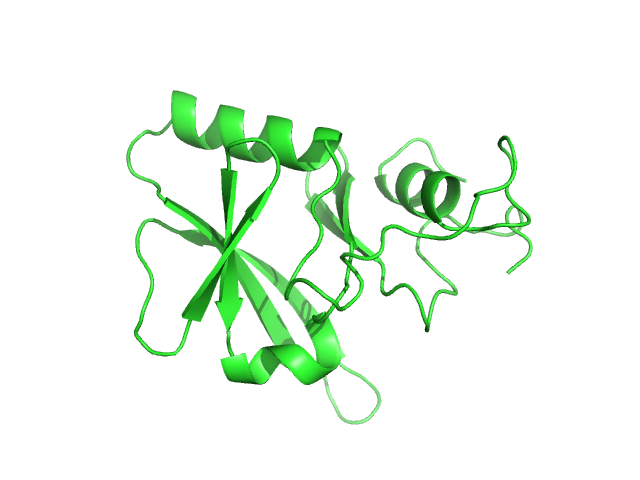
UniProt ID: P0DN75 (1-132) Iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Feb 25
|
Magnetic field effects on the structure and molecular behavior of pigeon iron–sulfur protein
Protein Science 31(6) (2022)
Arai S, Shimizu R, Adachi M, Hirai M
|
|
|
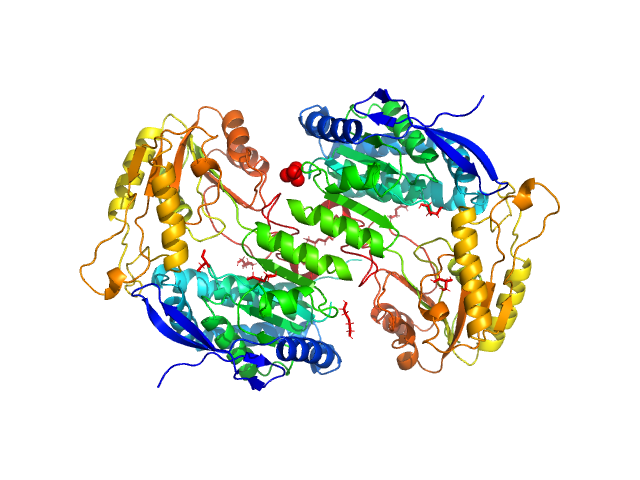
UniProt ID: Q9KWS5 (1-476) 2-aminomuconic 6-semialdehyde dehydrogenase
|
|
|
|
| Sample: |
2-aminomuconic 6-semialdehyde dehydrogenase dimer, 101 kDa Pseudomonas sp. protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2021 Feb 1
|
The tetrameric assembly of 2-aminomuconic 6-semialdehyde dehydrogenase is a functional requirement of cofactor NAD(+) binding.
Environ Microbiol 24(7):2994-3012 (2022)
Shi Q, Chen Y, Li X, Dong H, Chen C, Zhong Z, Yang C, Liu G, Su D
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
UniProt ID: Q45589 (97-273) Cyclic di-AMP synthase CdaA
UniProt ID: O34824 (1-369) Phosphoglucosamine mutase
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
Phosphoglucosamine mutase dimer, 83 kDa Bacillus subtilis protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Aug 2
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.1 |
nm |
| VolumePorod |
156 |
nm3 |
|
|
UniProt ID: J7M2L4 (41-454) NAD glycohydrolase
UniProt ID: P0C0I3 (32-571) Streptolysin O (T66M)
|
|
|
|
| Sample: |
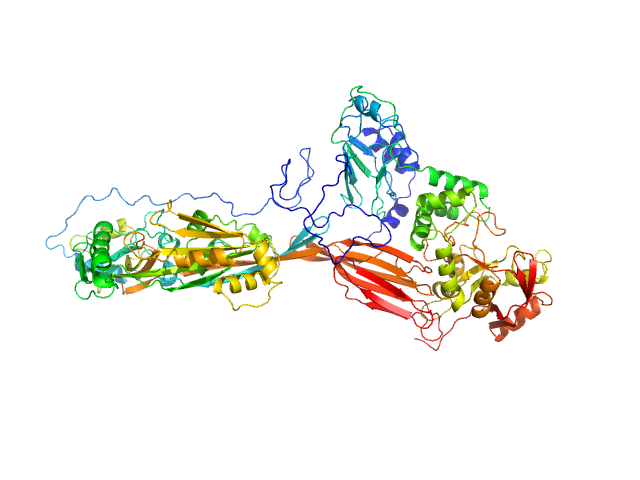
NAD glycohydrolase monomer, 47 kDa Streptococcus pyogenes M1 … protein
Streptolysin O (T66M) monomer, 63 kDa Streptococcus pyogenes serotype … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 16
|
Structural basis underlying the synergism of NADase and SLO during group A Streptococcus infection.
Commun Biol 6(1):124 (2023)
Tsai WJ, Lai YH, Shi YA, Hammel M, Duff AP, Whitten AE, Wilde KL, Wu CM, Knott R, Jeng US, Kang CY, Hsu CY, Wu JL, Tsai PJ, Chiang-Ni C, Wu JJ, Lin YS, Liu CC, Senda T, Wang S
|
| RgGuinier |
4.8 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
125 |
nm3 |
|
|
UniProt ID: P9WJD9 (2-348) ESX-1 secretion-associated protein EspB
|
|
|
|
| Sample: |
ESX-1 secretion-associated protein EspB monomer, 37 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM Tris-HCl, 300 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Sep 21
|
The crystal structure of the EspB-EspK virulence factor-chaperone complex suggests an additional type VII secretion mechanism in M. tuberculosis.
J Biol Chem :102761 (2022)
Gijsbers A, Eymery M, Gao Y, Menart I, Vinciauskaite V, Siliqi D, Peters PJ, McCarthy A, Ravelli RBG
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.3 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
UniProt ID: P02787 (1-698) Serotransferrin
|
|
|
|
| Sample: |
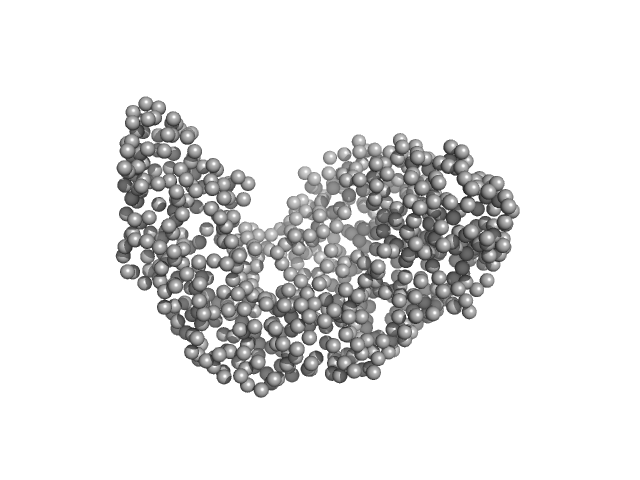
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (APO Buffer), pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.6 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
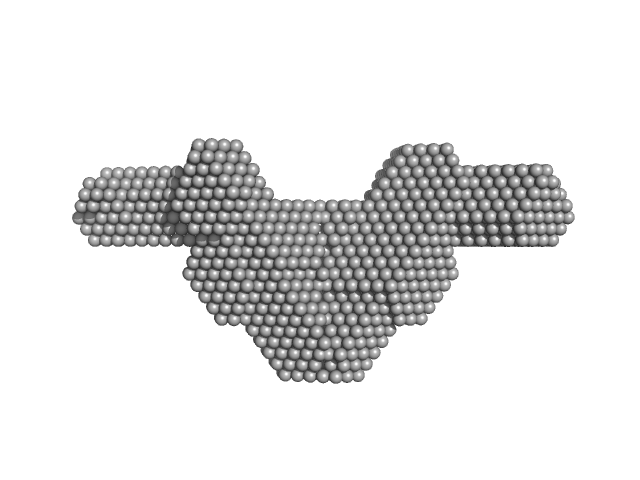
UniProt ID: Q15233 (53-312) Non-POU domain-containing octamer-binding protein
UniProt ID: P23246 (276-598) Splicing factor, proline- and glutamine-rich SFPQ276-598(R542C)/NONO53-312 dimer)
|
|
|

|
| Sample: |
Non-POU domain-containing octamer-binding protein dimer, 60 kDa Homo sapiens protein
Splicing factor, proline- and glutamine-rich SFPQ276-598(R542C)/NONO53-312 dimer) dimer, 76 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 250 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Apr 28
|
Structural plasticity of the coiled-coil interactions in human SFPQ.
Nucleic Acids Res (2024)
Koning HJ, Lai JY, Marshall AC, Stroeher E, Monahan G, Pullakhandam A, Knott GJ, Ryan TM, Fox AH, Whitten A, Lee M, Bond CS
|
| RgGuinier |
5.2 |
nm |
| Dmax |
21.0 |
nm |
| VolumePorod |
257 |
nm3 |
|
|
UniProt ID: None (44-60) Protein Tat
UniProt ID: None (None-None) Homo sapiens RNA component of 7SK nuclear ribonucleoprotein (RN7SK), small nuclear RNA
|
|
|
|
| Sample: |
Protein Tat monomer, 2 kDa Human immunodeficiency virus … protein
Homo sapiens RNA component of 7SK nuclear ribonucleoprotein (RN7SK), small nuclear RNA monomer, 18 kDa Homo sapiens RNA
|
| Buffer: |
10 mM phosphate, 70 mM NaCl, 0.1 mM EDTA, pH: 5.6 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 9
|
A structure-based mechanism for displacement of the HEXIM adapter from 7SK small nuclear RNA.
Commun Biol 5(1):819 (2022)
Pham VV, Gao M, Meagher JL, Smith JL, D'Souza VM
|
| RgGuinier |
2.3 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
28 |
nm3 |
|
|