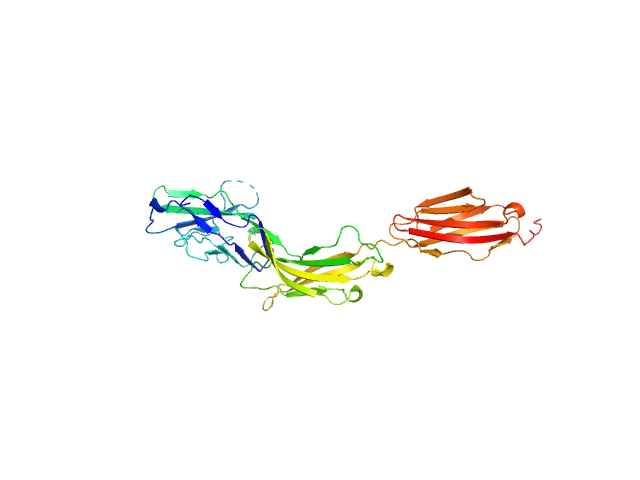
UniProt ID: P20917 (20-325) Myelin-associated glycoprotein Ig domains 1-3
|
|
|
|
| Sample: |
Myelin-associated glycoprotein Ig domains 1-3 monomer, 35 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 11
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
59 |
nm3 |
|
|
UniProt ID: Q15185 (1-131) Prostaglandin E synthase 3 (1-131)
|
|
|
|
| Sample: |
Prostaglandin E synthase 3 (1-131) monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM NaCl, 5 mM B-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS1 Beamline, Brazilian Synchrotron Light Laboratory on 2013 Jun 21
|
The C-terminal region of the human p23 chaperone modulates its structure and function.
Arch Biochem Biophys 565:57-67 (2015)
Seraphim TV, Gava LM, Mokry DZ, Cagliari TC, Barbosa LR, Ramos CH, Borges JC
|
| RgGuinier |
1.9 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
UniProt ID: P31483-2 (93-274) Nucleolysin TIA-1 isoform p40
UniProt ID: (None-None) RNA (ACUCCUUUUU)
|
|
|
|
| Sample: |
Nucleolysin TIA-1 isoform p40 monomer, 21 kDa Homo sapiens protein
RNA (ACUCCUUUUU) monomer, 1 kDa RNA
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 May 27
|
TIA-1 RRM23 binding and recognition of target oligonucleotides.
Nucleic Acids Res 45(8):4944-4957 (2017)
Waris S, García-Mauriño SM, Sivakumaran A, Beckham SA, Loughlin FE, Gorospe M, Díaz-Moreno I, Wilce MCJ, Wilce JA
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
UniProt ID: Q96FI4 (None-None) Endonuclease 8-like 1
UniProt ID: (None-None) dsDNA
UniProt ID: P12004 (None-None) Proliferating cell nuclear antigen
|
|
|

|
| Sample: |
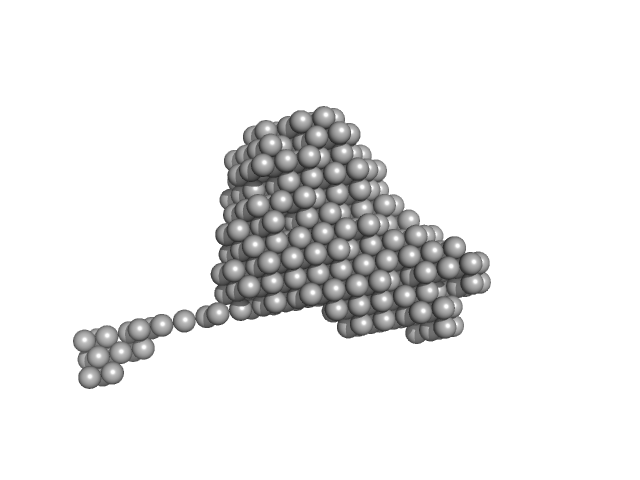
Endonuclease 8-like 1 monomer, 45 kDa Homo sapiens protein
DsDNA monomer, 2 kDa DNA
Proliferating cell nuclear antigen monomer, 30 kDa Homo sapiens protein
|
| Buffer: |
25mM HEPES 100mM NaCl 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Jan 20
|
Destabilization of the PCNA trimer mediated by its interaction with the NEIL1 DNA glycosylase.
Nucleic Acids Res 45(5):2897-2909 (2017)
Prakash A, Moharana K, Wallace SS, Doublié S
|
| RgGuinier |
3.4 |
nm |
| Dmax |
16.4 |
nm |
| VolumePorod |
113 |
nm3 |
|
|
UniProt ID: P39685 (None-None) Nucleoporin POM152
|
|
|

|
| Sample: |
Nucleoporin POM152 monomer, 26 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 10%(v/v) glycerol, 5mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2015 Apr 12
|
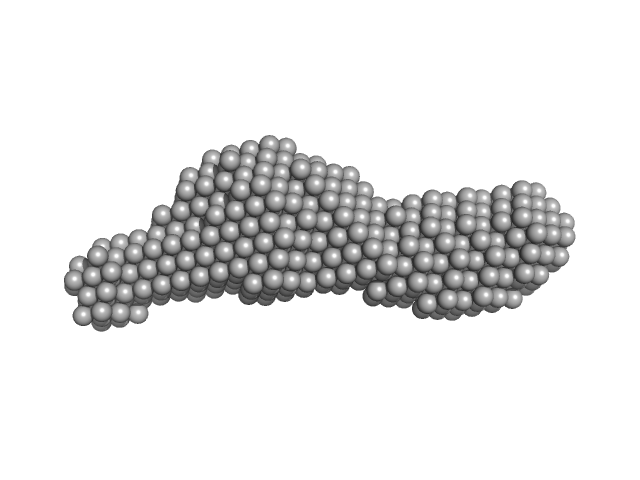
Molecular Architecture of the Major Membrane Ring Component of the Nuclear Pore Complex.
Structure 25(3):434-445 (2017)
Upla P, Kim SJ, Sampathkumar P, Dutta K, Cahill SM, Chemmama IE, Williams R, Bonanno JB, Rice WJ, Stokes DL, Cowburn D, Almo SC, Sali A, Rout MP, Fernandez-Martinez J
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
UniProt ID: Q58F21 (9-379) Bromodomain testis-specific protein
|
|
|

|
| Sample: |
Bromodomain testis-specific protein monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2% glycerol, 0.5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Jan 13
|
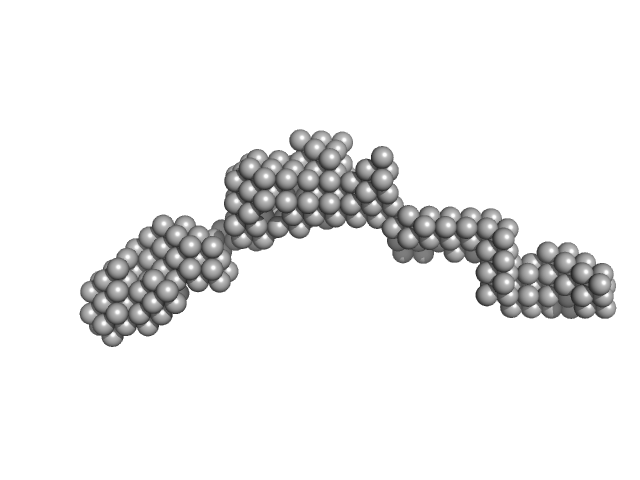
Interactome Rewiring Following Pharmacological Targeting of BET Bromodomains.
Mol Cell (2018)
Lambert JP, Picaud S, Fujisawa T, Hou H, Savitsky P, Uusküla-Reimand L, Gupta GD, Abdouni H, Lin ZY, Tucholska M, Knight JDR, Gonzalez-Badillo B, St-Denis N, Newman JA, Stucki M, Pelletier L, Bandeira N, Wilson MD, Filippakopoulos P, Gingras AC
|
| RgGuinier |
5.1 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
200 |
nm3 |
|
|
UniProt ID: Q9X721 (787-1118) Collagenase ColG segement s2s3as3b
|
|
|

|
| Sample: |
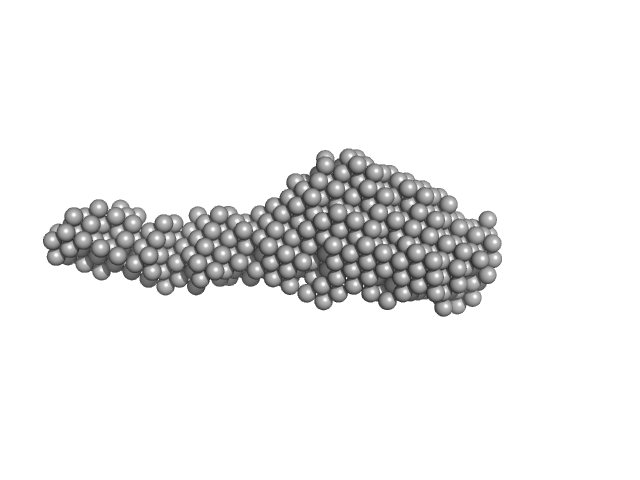
Collagenase ColG segement s2s3as3b monomer, 37 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
4.3 |
nm |
| Dmax |
19.7 |
nm |
| VolumePorod |
98 |
nm3 |
|
|
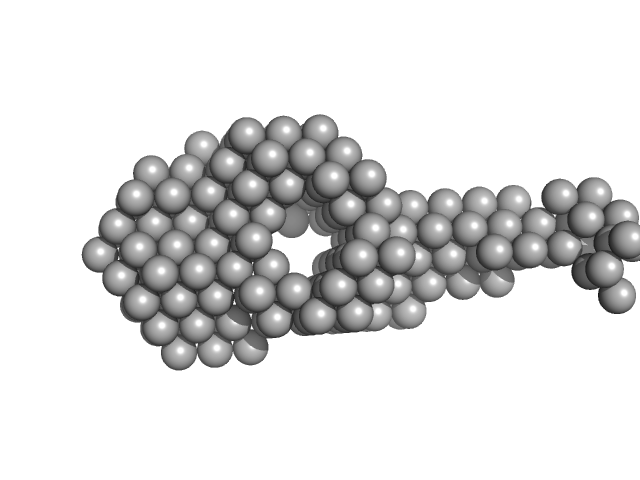
UniProt ID: P9WNV1 (None-None) DNA ligase A
UniProt ID: A0A0T9L251 (1-291) Probable exodeoxyribonuclease III protein XthA
|
|
|
|
| Sample: |
DNA ligase A monomer, 76 kDa Mycobacterium tuberculosis protein
Probable exodeoxyribonuclease III protein XthA monomer, 33 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 13
|
M. tuberculosis class II apurinic/ apyrimidinic-endonuclease/3'-5' exonuclease (XthA) engages with NAD+-dependent DNA ligase A (LigA) to counter futile cleavage and ligation cycles in base excision repair.
Nucleic Acids Res (2020)
Khanam T, Afsar M, Shukla A, Alam F, Kumar S, Soyar H, Dolma K, Pasupuleti M, Srivastava KK, Ampapathi RS, Ramachandran R
|
| RgGuinier |
6.2 |
nm |
| Dmax |
23.9 |
nm |
| VolumePorod |
662 |
nm3 |
|
|
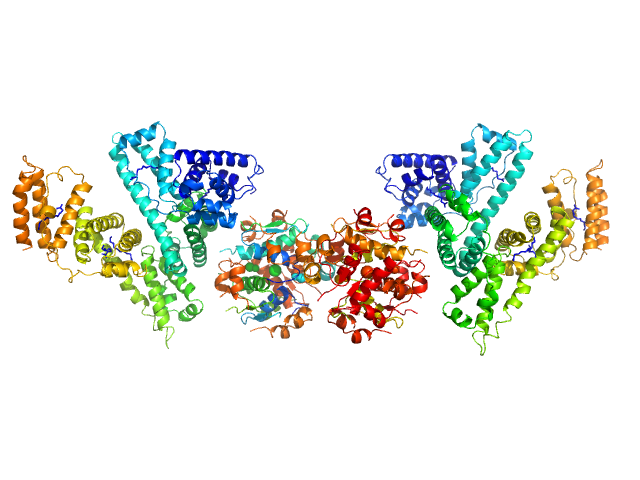
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
UniProt ID: P01308 (None-None) Insulin detemir (Levemir(R), Novo Nordisk A/S)
|
|
|
|
| Sample: |
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
Insulin detemir (Levemir(R), Novo Nordisk A/S) dodecamer, 71 kDa protein
|
| Buffer: |
8.8 mM Na2HPO4, 10.6 mM m-cresol, 12.2 mM phenol, 140.9 mM glycerol, 56.9 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
309 |
nm3 |
|
|
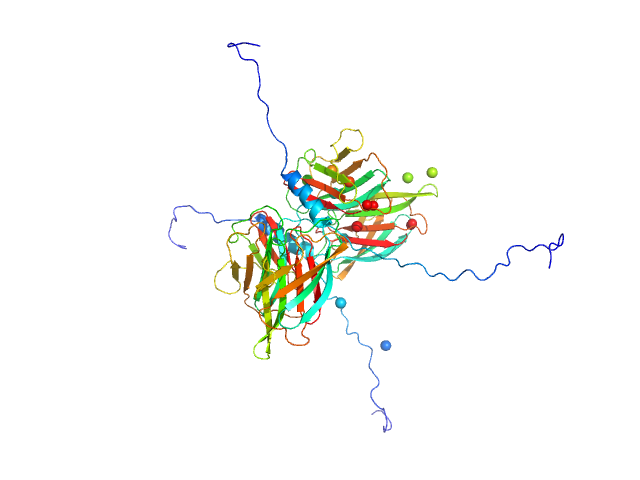
UniProt ID: Q4J9K8 (35-164) Conserved flagellar protein F
UniProt ID: Q4J9K7 (32-151) Stator protein FlaG-V118K soluble domain
|
|
|
|
| Sample: |
Conserved flagellar protein F dimer, 32 kDa Sulfolobus acidocaldarius protein
Stator protein FlaG-V118K soluble domain dimer, 30 kDa Sulfolobus acidocaldarius protein
|
| Buffer: |
25 mM citric acid/sodium citrate, 150mM NaCl, 3% Glycerol, pH: 3 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Nov 10
|
The structure of the periplasmic FlaG-FlaF complex and its essential role for archaellar swimming motility.
Nat Microbiol (2019)
Tsai CL, Tripp P, Sivabalasarma S, Zhang C, Rodriguez-Franco M, Wipfler RL, Chaudhury P, Banerjee A, Beeby M, Whitaker RJ, Tainer JA, Albers SV
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
108 |
nm3 |
|
|