UniProt ID: P53893 (None-None) Ribosome assembly protein 1
UniProt ID: Q07953 (None-None) Ribosome maturation protein SDO1
|
|
|

|
| Sample: |
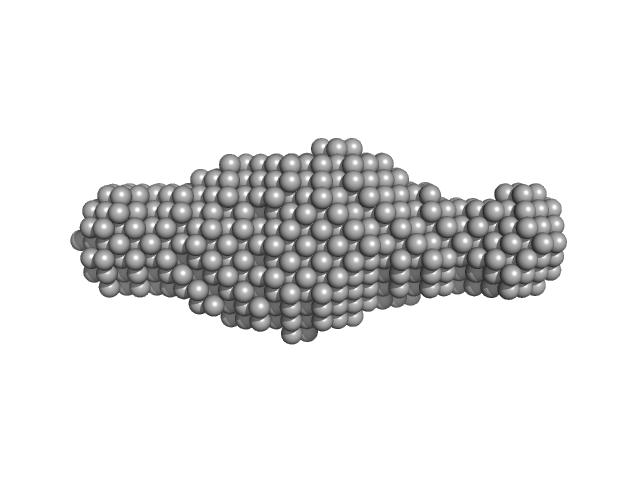
Ribosome assembly protein 1 monomer, 124 kDa Saccharomyces cerevisiae protein
Ribosome maturation protein SDO1 monomer, 28 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Sep 21
|
Interaction of the GTPase Elongation Factor Like-1 with the Shwachman-Diamond Syndrome Protein and Its Missense Mutations.
Int J Mol Sci 19(12) (2018)
Gijsbers A, Montagut DC, Méndez-Godoy A, Altamura D, Saviano M, Siliqi D, Sánchez-Puig N
|
| RgGuinier |
5.0 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
333 |
nm3 |
|
|
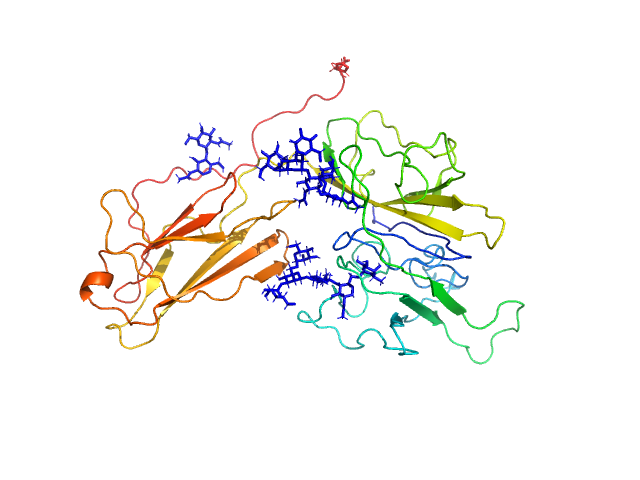
UniProt ID: O95256 (19-354) Interleukin-18 receptor accessory protein
|
|
|
|
| Sample: |
Interleukin-18 receptor accessory protein monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 3% glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2017 Jul 24
|
Functional Relevance of Interleukin-1 Receptor Inter-domain Flexibility for Cytokine Binding and Signaling.
Structure 27(8):1296-1307.e5 (2019)
Ge J, Remesh SG, Hammel M, Pan S, Mahan AD, Wang S, Wang X
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
UniProt ID: P11532 (1461-1973) Dystrophin (R11-15 human dystrophin fragment)
|
|
|

|
| Sample: |
Dystrophin (R11-15 human dystrophin fragment) monomer, 60 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-d11, 150 mM NaCl, 0.1 mM EDTA-d16, in 100% v/v D2O, pH: 7.1 |
| Experiment: |
SANS
data collected at D22, Institut Laue-Langevin (ILL) on 2016 Nov 1
|
How the central domain of dystrophin acts to bridge F-actin to sarcolemmal lipids.
J Struct Biol :107411 (2019)
Mias-Lucquin D, Dos Santos Morais R, Chéron A, Lagarrigue M, Winder SJ, Chenuel T, Pérez J, Appavou MS, Martel A, Alviset G, Le Rumeur E, Combet S, Hubert JF, Delalande O
|
| RgGuinier |
6.2 |
nm |
| Dmax |
28.1 |
nm |
| VolumePorod |
144 |
nm3 |
|
|
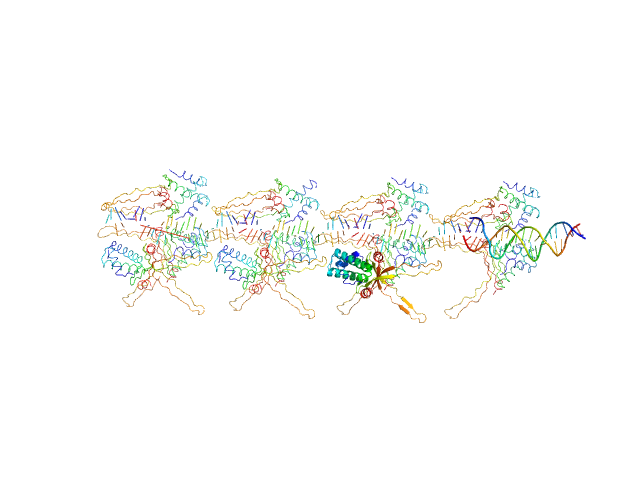
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha, E34K
|
|
|
|
| Sample: |
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, E34K, 305 kDa Escherichia coli protein
|
| Buffer: |
20mM HEPES, 100mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Jul 8
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
| RgGuinier |
8.0 |
nm |
| Dmax |
24.8 |
nm |
| VolumePorod |
448 |
nm3 |
|
|
UniProt ID: P00330 (1-348) Alcohol dehydrogenase 1
|
|
|

|
| Sample: |
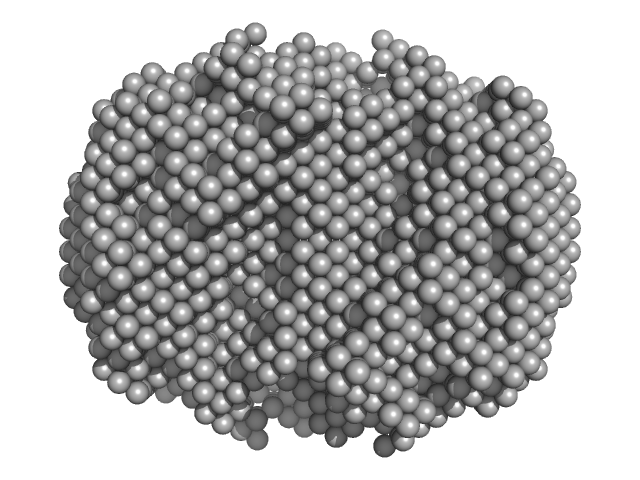
Alcohol dehydrogenase 1 tetramer, 147 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 2% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Apr 5
|
Adding Size Exclusion Chromatography (SEC) and Light Scattering (LS) Devices to Obtain High-Quality Small Angle X-Ray Scattering (SAXS) Data
Crystals 10(11):975 (2020)
Graewert M, Da Vela S, Gräwert T, Molodenskiy D, Blanchet C, Svergun D, Jeffries C
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
201 |
nm3 |
|
|
UniProt ID: O08967 (14-390) Cytohesin-3
|
|
|

|
| Sample: |
Cytohesin-3 dimer, 90 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2 mM MgCl2, 0.1% 2-mercaptoethanol, 5% glycerol, 0.001 mM insitol 1,3,4,5-tetrakis phosphate, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2013 Nov 15
|
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Structure (2019)
Das S, Malaby AW, Nawrotek A, Zhang W, Zeghouf M, Maslen S, Skehel M, Chakravarthy S, Irving TC, Bilsel O, Cherfils J, Lambright DG
|
| RgGuinier |
5.3 |
nm |
| Dmax |
25.7 |
nm |
| VolumePorod |
168 |
nm3 |
|
|
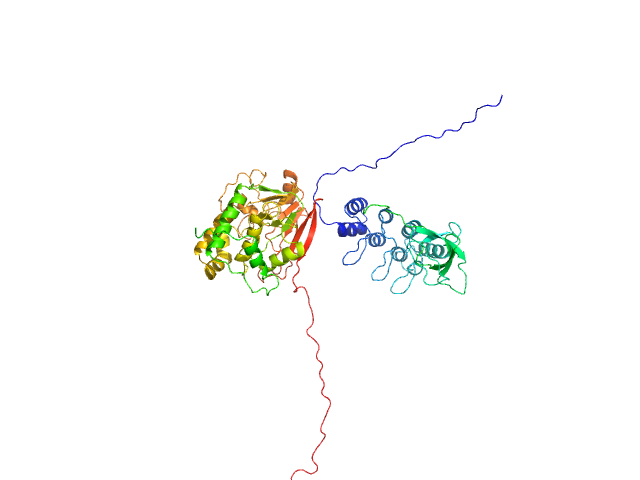
UniProt ID: P62136 (1-330) Serine/threonine-protein phosphatase PP1-alpha catalytic subunit
UniProt ID: Q8WUF5 (621-828) Inhibitor of apoptosis-stimulating protein of p53 (RelA-associated inhibitor)
|
|
|
|
| Sample: |
Serine/threonine-protein phosphatase PP1-alpha catalytic subunit monomer, 38 kDa Homo sapiens protein
Inhibitor of apoptosis-stimulating protein of p53 (RelA-associated inhibitor) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Apr 25
|
Flexible Tethering of ASPP Proteins Facilitates PP-1c Catalysis.
Structure 27(10):1485-1496.e4 (2019)
Zhou Y, Millott R, Kim HJ, Peng S, Edwards RA, Skene-Arnold T, Hammel M, Lees-Miller SP, Tainer JA, Holmes CFB, Glover JNM
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
UniProt ID: P52129 (2-357) mRNA endoribonuclease toxin LS - R255A mutant; N-terminal His-tagged
|
|
|
|
| Sample: |
MRNA endoribonuclease toxin LS - R255A mutant; N-terminal His-tagged dimer, 84 kDa Escherichia coli protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 25
|
Alternative dimerization is required for activity and inhibition of the HEPN ribonuclease RnlA
Nucleic Acids Research 49(12):7164-7178 (2021)
Garcia-Rodriguez G, Charlier D, Wilmaerts D, Michiels J, Loris R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
20.1 |
nm |
| VolumePorod |
133 |
nm3 |
|
|
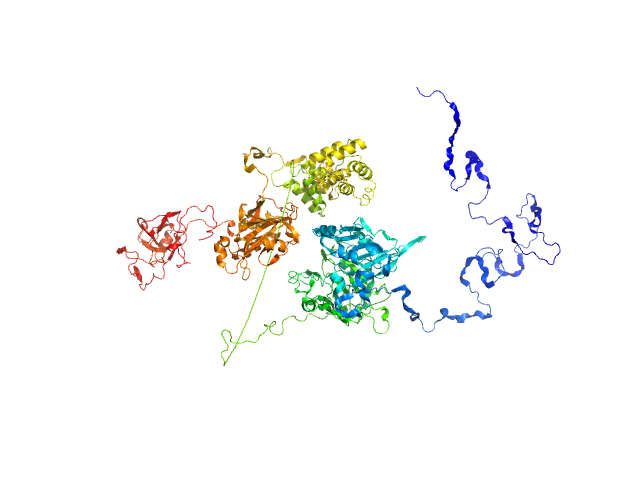
UniProt ID: Q9NUW8 (1-608) Tyrosyl-DNA phosphodiesterase 1
UniProt ID: P49916-3 (170-755) Isoform 3 of DNA ligase 3 (DNA ligase III alpha)
|
|
|
|
| Sample: |
Tyrosyl-DNA phosphodiesterase 1 monomer, 71 kDa Homo sapiens protein
Isoform 3 of DNA ligase 3 (DNA ligase III alpha) monomer, 68 kDa Homo sapiens protein
|
| Buffer: |
200 mM NaCl, 40 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jul 5
|
Direct interaction of DNA repair protein tyrosyl DNA phosphodiesterase 1 and the DNA ligase III catalytic domain is regulated by phosphorylation of its flexible N-terminus.
J Biol Chem :100921 (2021)
Rashid I, Hammel M, Sverzhinsky A, Tsai MS, Pascal JM, Tainer JA, Tomkinson AE
|
| RgGuinier |
4.5 |
nm |
| Dmax |
18.0 |
nm |
| VolumePorod |
211 |
nm3 |
|
|
UniProt ID: P11717 (1222-1510) Cation-independent mannose-6-phosphate receptor
|
|
|
|
| Sample: |
Cation-independent mannose-6-phosphate receptor monomer, 34 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Mar 3
|
Structure of the Human Cation-Independent Mannose 6-Phosphate/IGF2 Receptor Domains 7–11 Uncovers the Mannose 6-Phosphate Binding Site of Domain 9
Structure (2020)
Bochel A, Williams C, McCoy A, Hoppe H, Winter A, Nicholls R, Harlos K, Jones E, Berger I, Hassan A, Crump M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
69 |
nm3 |
|
|