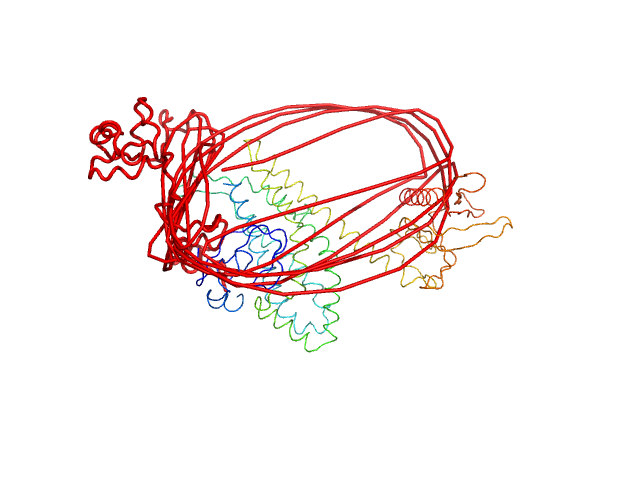
UniProt ID: Q92974-1 (28-582) Isoform 1 of Rho guanine nucleotide exchange factor 2
|
|
|
|
| Sample: |
Isoform 1 of Rho guanine nucleotide exchange factor 2 monomer, 64 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris-Cl, 150mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Feb 19
|
Structural basis of microtubule-mediated signal transduction.
Cell (2025)
Choi SR, Blum TB, Giono M, Roy B, Vakonakis I, Schmid D, Oelgarth N, Ranganathan A, Gossert AD, Shivashankar GV, Zippelius A, Steinmetz MO
|
|
|
UniProt ID: Q8N4Q1 (1-142) Mitochondrial intermembrane space import and assembly protein 40
|
|
|
|
| Sample: |
Mitochondrial intermembrane space import and assembly protein 40 monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Oct 26
|
NADH-bound AIF activates the mitochondrial CHCHD4/MIA40 chaperone by a substrate-mimicry mechanism.
EMBO J (2025)
Brosey CA, Shen R, Tainer JA
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
62 |
nm3 |
|
|
UniProt ID: Q86YC2 (1-194) Partner and localizer of BRCA2 (L24A)
|
|
|
|
| Sample: |
Partner and localizer of BRCA2 (L24A) monomer, 23 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 160 mM NaCl, 0.5 mM TCEP pH 7.5, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2023 Dec 9
|
The strand exchange domain of tumor suppressor PALB2 is intrinsically disordered and promotes oligomerization-dependent DNA compaction.
iScience 27(12):111259 (2024)
Kyriukha Y, Watkins MB, Redington JM, Chintalapati N, Ganti A, Dastvan R, Uversky VN, Hopkins JB, Pozzi N, Korolev S
|
| RgGuinier |
5.2 |
nm |
| Dmax |
21.6 |
nm |
| VolumePorod |
127 |
nm3 |
|
|
UniProt ID: Q4WF55 (1-200) N(5)-hydroxyornithine:cis-anhydromevalonyl coenzyme A-N(5)-transacylase sidF
|
|
|
|
| Sample: |
N(5)-hydroxyornithine:cis-anhydromevalonyl coenzyme A-N(5)-transacylase sidF monomer, 25 kDa Aspergillus fumigatus (strain … protein
|
| Buffer: |
50 mM Tris, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2023 Jan 18
|
SidF, a dual substrate N5-acetyl-N5-hydroxy-L-ornithine transacetylase involved in Aspergillus fumigatus siderophore biosynthesis
Journal of Structural Biology: X 11:100119 (2025)
Poonsiri T, Stransky J, Demitri N, Haas H, Cianci M, Benini S
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
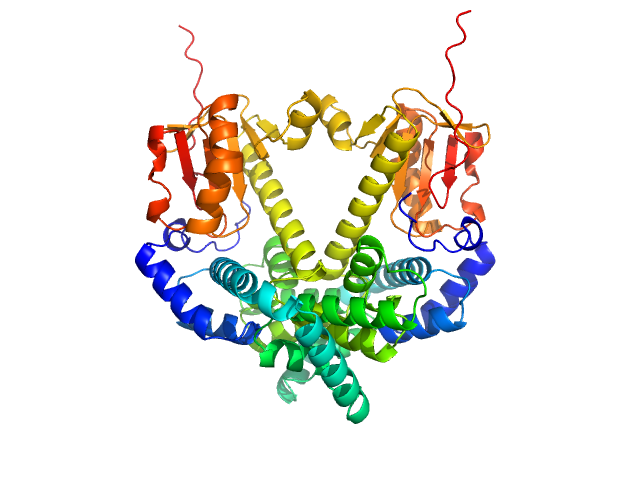
UniProt ID: Q9Y2V0 (1-281) CDAN1-interacting nuclease 1
|
|
|
|
| Sample: |
CDAN1-interacting nuclease 1 monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Sep 29
|
Anemia-associated mutations disrupt the CDIN1-Codanin1 complex in inherited congenital dyserythropoietic anemia I (CDA-I) disease.
FEBS J (2026)
Stojaspal M, Brom T, Nečasová I, Janovič T, Veverka P, Verma N, Uhrík L, Hernychova L, Hofr C
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
104 |
nm3 |
|
|
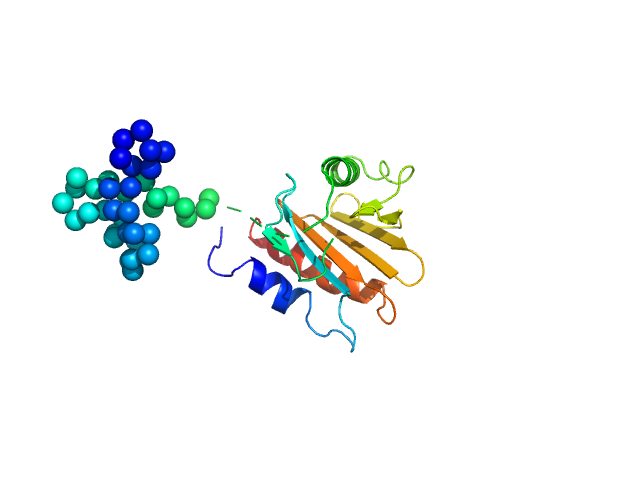
UniProt ID: Q9STB6 (1-131) Profilin-2
|
|
|
|
| Sample: |
Profilin-2 monomer, 18 kDa Hevea brasiliensis protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 3
|
Allergen-induced structural rearrangements in IgE: insights from SAXS and molecular dynamics.
Int J Biol Macromol :147658 (2025)
Gómez-Velasco H, García-Ramírez B, Siliqi D, Graewert MA, Quintero-Martinez A, Ortega E, Rodríguez-Romero A
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
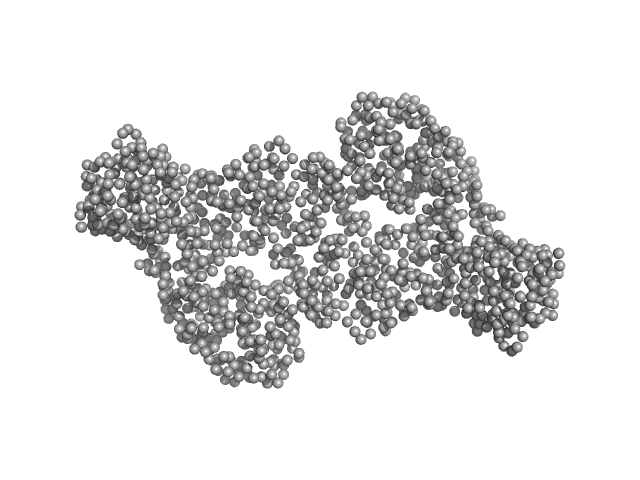
UniProt ID: Q9UJW3 (178-379) DNA methyltransferase 3-like (178-379)
UniProt ID: Q9UBC3 (413-853) DNA methyltransferase 3 beta (413-853)
|
|
|
|
| Sample: |
DNA methyltransferase 3-like (178-379) dimer, 47 kDa Homo sapiens protein
DNA methyltransferase 3 beta (413-853) dimer, 100 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM TCEP, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2022 Nov 2
|
Histone modification-driven structural remodeling unleashes DNMT3B in DNA methylation.
Sci Adv 11(13):eadu8116 (2025)
Cho CC, Huang HH, Jiang BC, Yang WZ, Chen YN, Yuan HS
|
| RgGuinier |
4.5 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
191 |
nm3 |
|
|
UniProt ID: P30681 (1-210) High mobility group protein B2
UniProt ID: None (None-None) Double stranded DNA 20mer
|
|
|
|
| Sample: |
High mobility group protein B2 monomer, 26 kDa Mus musculus protein
Double stranded DNA 20mer monomer, 12 kDa DNA
|
| Buffer: |
Tris 50mM, NaCl 150mM, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Apr 27
|
High mobility group protein B2 (HMGB2)
Chun I Tu
|
| RgGuinier |
3.3 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
49 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2019 Sep 21
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
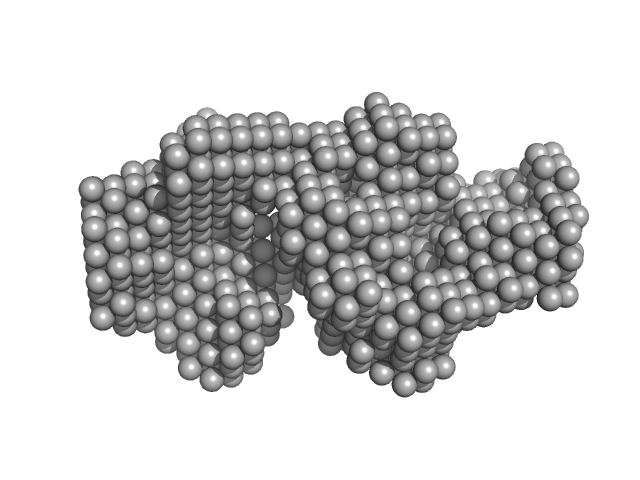
UniProt ID: P01267 (None-None) Thyroglobulin
|
|
|

|
| Sample: |
Thyroglobulin dimer, 606 kDa Bos taurus protein
|
| Buffer: |
100 mM Tris/HCl 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 May 19
|
Accuracy of molecular mass determination of proteins in solution by small-angle X-ray scattering
Journal of Applied Crystallography 40(s1):s245-s249 (2007)
Mylonas E, Svergun D
|
| RgGuinier |
7.9 |
nm |
| Dmax |
24.0 |
nm |
|
|