UniProt ID: P73129 (1-96) Ssr1698 protein
|
|
|
|
| Sample: |
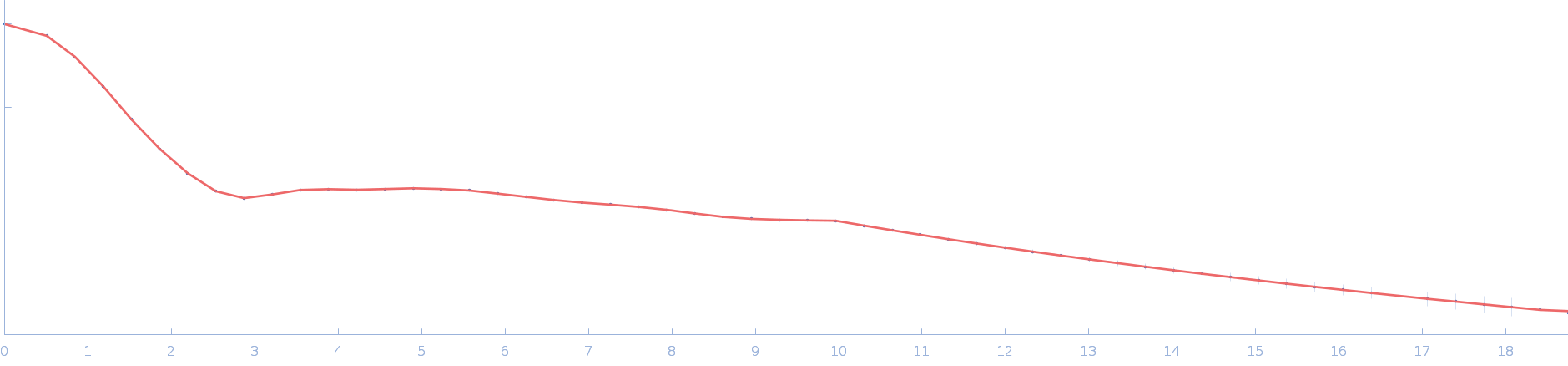
Ssr1698 protein dimer, 23 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Hepes, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2022 Sep 27
|
A hemoprotein with a zinc-mirror heme site ties heme availability to carbon metabolism in cyanobacteria.
Nat Commun 15(1):3167 (2024)
Grosjean N, Yee EF, Kumaran D, Chopra K, Abernathy M, Biswas S, Byrnes J, Kreitler DF, Cheng JF, Ghosh A, Almo SC, Iwai M, Niyogi KK, Pakrasi HB, Sarangi R, van Dam H, Yang L, Blaby IK, Blaby-Haas CE
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.2 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
UniProt ID: P54760 (586-602) Ephrin type-B receptor 4
UniProt ID: P20936 (174-1047) Ras GTPase-activating protein 1
|
|
|
|
| Sample: |
Ephrin type-B receptor 4 monomer, 2 kDa Homo sapiens protein
Ras GTPase-activating protein 1 monomer, 101 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 150 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 4
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
176 |
nm3 |
|
|
UniProt ID: Q9NZ71-6 (756-1300) Regulator of telomere elongation helicase 1 (Isoform 6, 756-1300)
|
|
|
|
| Sample: |
Regulator of telomere elongation helicase 1 (Isoform 6, 756-1300) monomer, 59 kDa Homo sapiens protein
|
| Buffer: |
50 mM Hepes, 150 mM NaCl,10 (v/v)% glycerol, 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 22
|
Structural and biochemical characterization of the C‐terminal region of the human RTEL
1 helicase
Protein Science 33(9) (2024)
Cortone G, Graewert M, Kanade M, Longo A, Hegde R, González‐Magaña A, Chaves‐Arquero B, Blanco F, Napolitano L, Onesti S
|
| RgGuinier |
4.9 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
UniProt ID: A0A804LC01 (30-200) Protein PAIR1
|
|
|

|
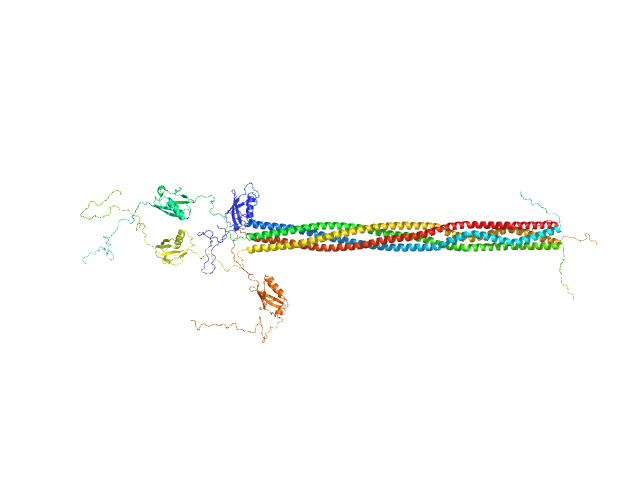
| Sample: |
Protein PAIR1 tetramer, 132 kDa Zea mays protein
|
| Buffer: |
25 mM HEPES-NaOH, 500 mM NaCl, 5 mM EDTA, 5% glycerol, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2023 Apr 14
|
Evolutionary conservation of the structure and function of meiotic Rec114-Mei4 and Mer2 complexes.
Genes Dev 37(11-12):535-553 (2023)
Daccache D, De Jonge E, Liloku P, Mechleb K, Haddad M, Corthaut S, Sterckx YG, Volkov AN, Claeys Bouuaert C
|
| RgGuinier |
8.0 |
nm |
| Dmax |
31.7 |
nm |
|
|
UniProt ID: Q96RR4 (93-517) Calcium/calmodulin-dependent protein kinase kinase 2
|
|
|
|
| Sample: |
Calcium/calmodulin-dependent protein kinase kinase 2 monomer, 48 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM TCEP, 3% (w/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
14-3-3 protein inhibits CaMKK1 by blocking the kinase active site with its last two C-terminal helices.
Protein Sci :e4805 (2023)
Petrvalska O, Honzejkova K, Koupilova N, Herman P, Obsilova V, Obsil T
|
| RgGuinier |
3.0 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
UniProt ID: P09651-2 (186-320) Isoform A1-A of Heterogeneous nuclear ribonucleoprotein A1
|
|
|
|
| Sample: |
Isoform A1-A of Heterogeneous nuclear ribonucleoprotein A1 monomer, 13 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 19
|
Design of intrinsically disordered protein variants with diverse structural properties
Science Advances 10(35) (2024)
Pesce F, Bremer A, Tesei G, Hopkins J, Grace C, Mittag T, Lindorff-Larsen K
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
19 |
nm3 |
|
|
UniProt ID: Q9WTS6-1 (343-2715) Isoform A1B1 of Teneurin-3
|
|
|
|
| Sample: |
Isoform A1B1 of Teneurin-3 dimer, 545 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH: 7.8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Sep 10
|
Alternative splicing controls teneurin-3 compact dimer formation for neuronal recognition
Nature Communications 15(1) (2024)
Gogou C, Beugelink J, Frias C, Kresik L, Jaroszynska N, Drescher U, Janssen B, Hindges R, Meijer D
|
| RgGuinier |
7.9 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
1129 |
nm3 |
|
|
UniProt ID: Q1WK95 (29-462) ISG75
|
|
|
|
| Sample: |
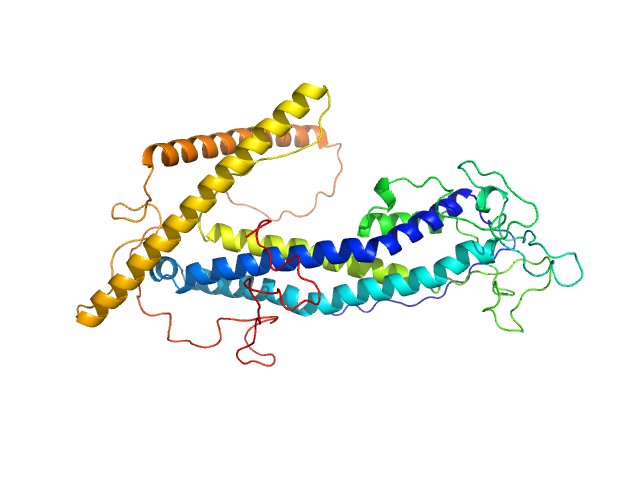
ISG75 monomer, 50 kDa Trypanosoma brucei gambiense protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 3% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Nov 25
|
Beyond the VSG Layer: Exploring the Role of Intrinsic Disorder in the Invariant Surface Glycoproteins of African Trypanosomes.
PLoS Pathog 20(4):e1012186 (2024)
Sülzen H, Volkov AN, Geens R, Zahedifard F, Stijlemans B, Zoltner M, Magez S, Sterckx YG, Zoll S
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
UniProt ID: P0DTC9 (1-365) Nucleoprotein
|
|
|
|
| Sample: |
Nucleoprotein hexamer, 245 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 13
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
7.1 |
nm |
| Dmax |
29.0 |
nm |
| VolumePorod |
708 |
nm3 |
|
|
UniProt ID: J3QLL0 (2-73) Importin subunit alpha-1
|
|
|
|
| Sample: |
Importin subunit alpha-1 monomer, 9 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline containing 6 M urea, 0.3 M KCl, 5 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Jun 29
|
A genetically encoded anomalous SAXS ruler to probe the dimensions of intrinsically disordered proteins.
Proc Natl Acad Sci U S A 121(50):e2415220121 (2024)
Yu M, Gruzinov AY, Ruan H, Scheidt T, Chowdhury A, Giofrè S, Mohammed ASA, Caria J, Sauter PF, Svergun DI, Lemke EA
|
| RgGuinier |
2.7 |
nm |
| Dmax |
11.9 |
nm |
| VolumePorod |
23 |
nm3 |
|
|