UniProt ID: Q99QS1 (32-478) Protein map
UniProt ID: P08311 (22-245) Cathepsin G
|
|
|
|
| Sample: |
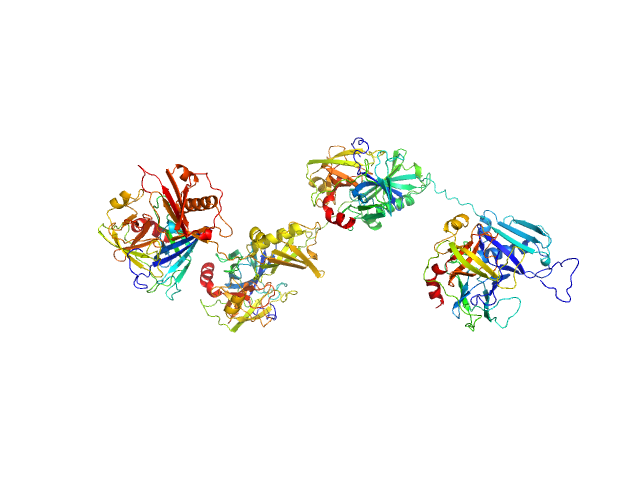
Protein map monomer, 50 kDa Staphylococcus aureus (strain … protein
Cathepsin G tetramer, 101 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Jun 3
|
S. aureus Eap is a polyvalent inhibitor of neutrophil serine proteases.
J Biol Chem 300(9):107627 (2024)
Mishra N, Gido CD, Herdendorf TJ, Hammel M, Hura GL, Fu ZQ, Geisbrecht BV
|
| RgGuinier |
5.0 |
nm |
| Dmax |
20.2 |
nm |
| VolumePorod |
166 |
nm3 |
|
|
UniProt ID: P0DN75 (2-132) Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Pigeon iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2021 Nov 1
|
A hidden property of the iron-sulfur protein in the mononuclear iron-bound state: species-dependent structural ordering induced by magnetic fields.
FEBS J (2025)
Arai S, Soga S, Hirai M, Kobayashi R, Masai H, Kimura K, Maeda K, Nagashima H
|
|
|
UniProt ID: Q8IWY9 (1005-1227) Codanin-1
|
|
|
|
| Sample: |
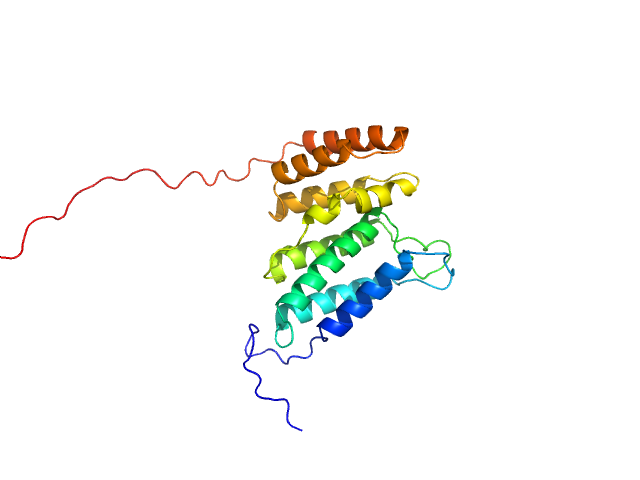
Codanin-1 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris HCl, 150 mM NaCl, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Sep 29
|
Anemia-associated mutations disrupt the CDIN1-Codanin1 complex in inherited congenital dyserythropoietic anemia I (CDA-I) disease.
FEBS J (2026)
Stojaspal M, Brom T, Nečasová I, Janovič T, Veverka P, Verma N, Uhrík L, Hernychova L, Hofr C
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
51 |
nm3 |
|
|
UniProt ID: Q8N4Q1 (1-142) Mitochondrial intermembrane space import and assembly protein 40
|
|
|
|
| Sample: |
Mitochondrial intermembrane space import and assembly protein 40 monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Oct 26
|
NADH-bound AIF activates the mitochondrial CHCHD4/MIA40 chaperone by a substrate-mimicry mechanism.
EMBO J (2025)
Brosey CA, Shen R, Tainer JA
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
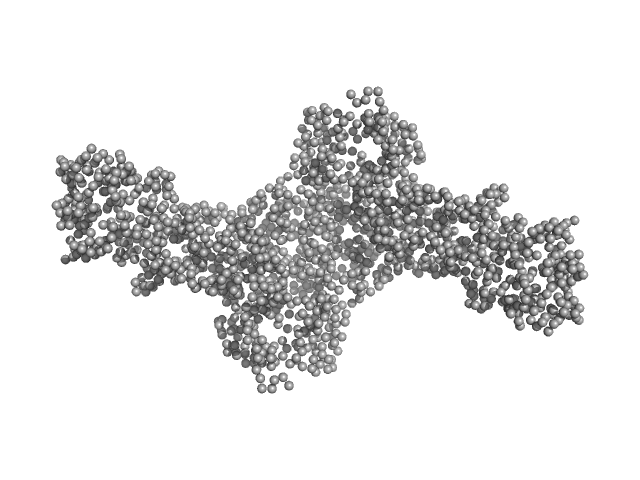
UniProt ID: Q9UBC3 (215-853) DNA methyltransferase 3 beta (215-853)
UniProt ID: Q9UJW3 (178-379) DNA methyltransferase 3-like (178-379)
|
|
|

|
| Sample: |
DNA methyltransferase 3 beta (215-853) dimer, 145 kDa Homo sapiens protein
DNA methyltransferase 3-like (178-379) dimer, 47 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM TCEP, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2021 Apr 16
|
Histone modification-driven structural remodeling unleashes DNMT3B in DNA methylation.
Sci Adv 11(13):eadu8116 (2025)
Cho CC, Huang HH, Jiang BC, Yang WZ, Chen YN, Yuan HS
|
| RgGuinier |
5.2 |
nm |
| Dmax |
21.2 |
nm |
| VolumePorod |
271263 |
nm3 |
|
|
UniProt ID: P30681 (1-166) High mobility group protein B2
|
|
|
|
| Sample: |
High mobility group protein B2 monomer, 20 kDa Mus musculus protein
|
| Buffer: |
Tris 50mM, NaCl 150mM, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2024 Apr 27
|
High mobility group protein B2 (HMGB2)
Chun I Tu
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
42 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2019 Sep 21
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
UniProt ID: P20917 (20-508) Myelin-associated glycoprotein (20-508; I473E mutant)
|
|
|
|
| Sample: |
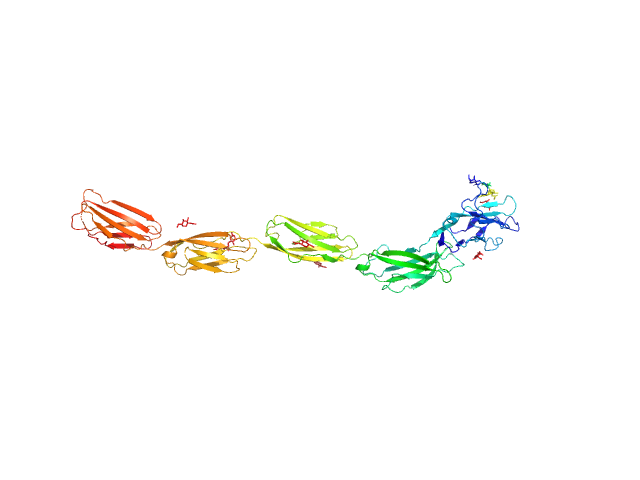
Myelin-associated glycoprotein (20-508; I473E mutant) monomer, 54 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 11
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
6.0 |
nm |
| Dmax |
21.2 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
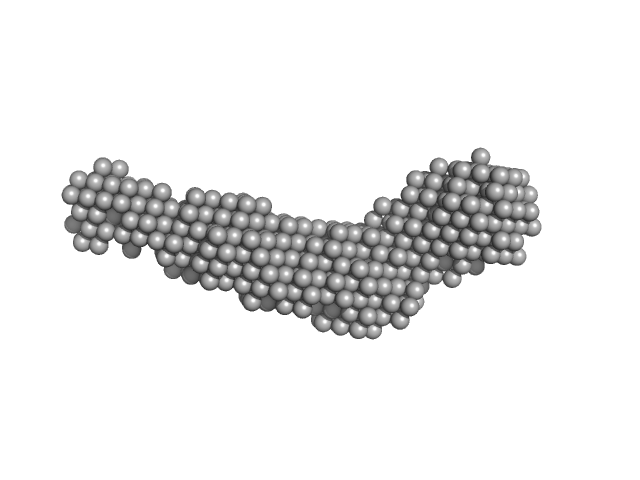
UniProt ID: Q46085 (718-1021) Collagenase ColH segement s2as2bs3
|
|
|

|
| Sample: |
Collagenase ColH segement s2as2bs3 monomer, 34 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
3.3 |
nm |
| Dmax |
13.6 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
UniProt ID: A0A0T9L251 (None-None) Probable exodeoxyribonuclease III protein XthA
UniProt ID: P9WNV1 (601-691) M. tb. LigA BRCT domain
|
|
|
|
| Sample: |
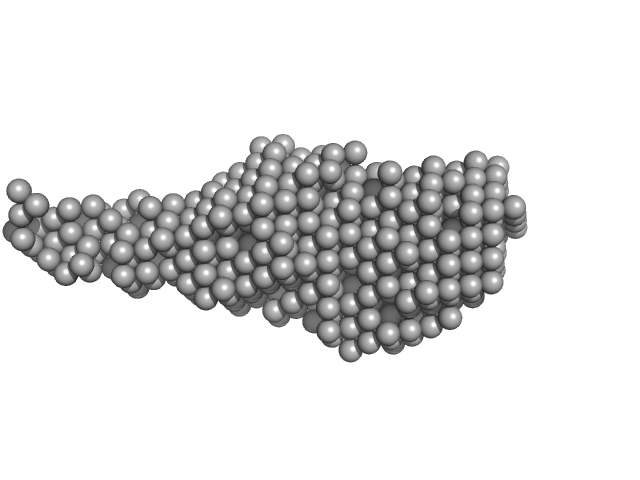
Probable exodeoxyribonuclease III protein XthA monomer, 33 kDa Mycobacterium tuberculosis protein
M. tb. LigA BRCT domain monomer, 15 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl 500 mM NaCl 5mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Mar 9
|
M. tuberculosis class II apurinic/ apyrimidinic-endonuclease/3'-5' exonuclease (XthA) engages with NAD+-dependent DNA ligase A (LigA) to counter futile cleavage and ligation cycles in base excision repair.
Nucleic Acids Res (2020)
Khanam T, Afsar M, Shukla A, Alam F, Kumar S, Soyar H, Dolma K, Pasupuleti M, Srivastava KK, Ampapathi RS, Ramachandran R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
18.5 |
nm |
| VolumePorod |
112 |
nm3 |
|
|