UniProt ID: P0DTC9 (1-419) Nucleoprotein
|
|
|
|
| Sample: |
Nucleoprotein hexamer, 281 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 13
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
7.6 |
nm |
| Dmax |
32.0 |
nm |
| VolumePorod |
764 |
nm3 |
|
|
UniProt ID: P08311 (22-245) Cathepsin G
UniProt ID: P08246 (31-249) Neutrophil elastase
UniProt ID: Q99QS1 (268-438) Eap34
|
|
|
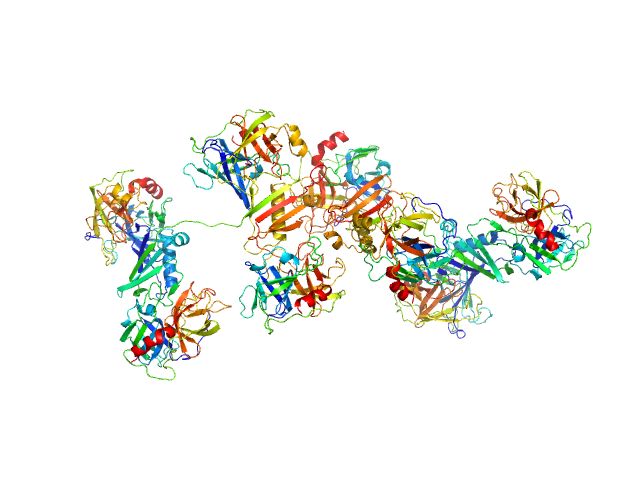
|
| Sample: |
Cathepsin G tetramer, 101 kDa Homo sapiens protein
Neutrophil elastase tetramer, 93 kDa Homo sapiens protein
Eap34 dimer, 48 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Jun 3
|
S. aureus Eap is a polyvalent inhibitor of neutrophil serine proteases.
J Biol Chem 300(9):107627 (2024)
Mishra N, Gido CD, Herdendorf TJ, Hammel M, Hura GL, Fu ZQ, Geisbrecht BV
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.4 |
nm |
| VolumePorod |
500 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
GlcNAc-binding protein A monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at D11, ILL on 2020 Aug 18
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
UniProt ID: Q46085 (718-1021) Collagenase ColH segement s2as2bs3
|
|
|
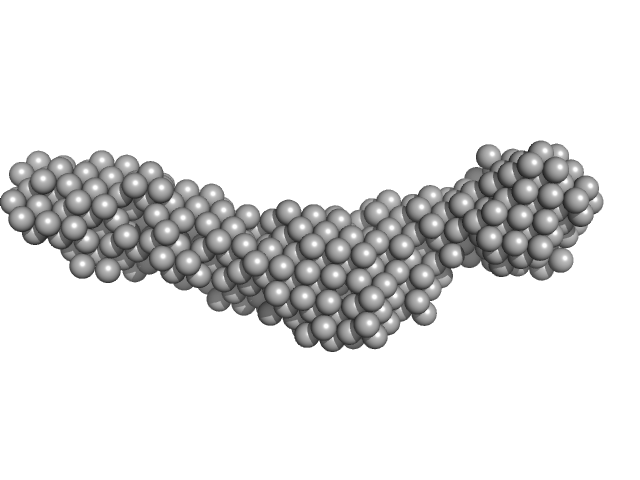
|
| Sample: |
Collagenase ColH segement s2as2bs3 monomer, 34 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
3.2 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha
|
|
|
|
| Sample: |
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, 10 kDa Escherichia coli protein
|
| Buffer: |
10 mM sodium acetate, 150 mM NaCl, pH: 4.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jun 1
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
|
|
UniProt ID: Q9GZZ9 (57-346) Ubiquitin-like modifier-activating enzyme 5
|
|
|
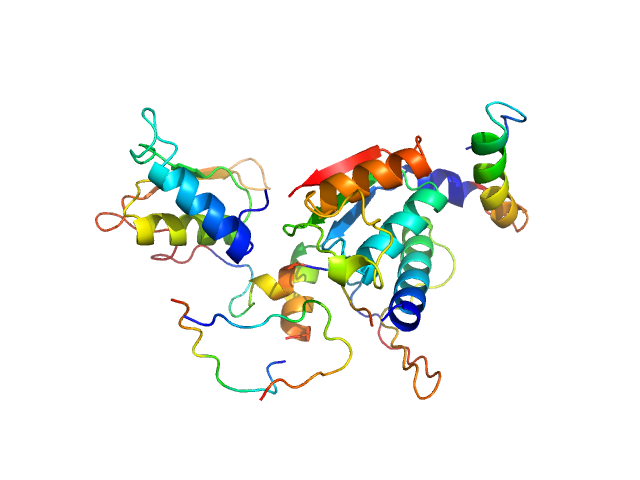
|
| Sample: |
Ubiquitin-like modifier-activating enzyme 5 dimer, 68 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Jun 13
|
Structure and dynamics of UBA5-UFM1 complex formation showing new insights in the UBA5 activation mechanism
Journal of Structural Biology :107796 (2021)
Fuchs S, Kikhney A, Schubert R, Kaiser C, Liebau E, Svergun D, Betzel C, Perbandt M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
UniProt ID: O60885 (38-460) Bromodomain-containing protein 4
|
|
|
|
| Sample: |
Bromodomain-containing protein 4 monomer, 48 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 25
|
Multivalent nucleosome scaffolding by bromodomain and extraterminal domain tandem bromodomains.
J Biol Chem :108289 (2025)
Olp MD, Bursch KL, Wynia-Smith SL, Nuñez R, Goetz CJ, Jackson V, Smith BC
|
| RgGuinier |
5.5 |
nm |
| Dmax |
20.2 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
UniProt ID: P02768 (None-None) Albumin
|
|
|
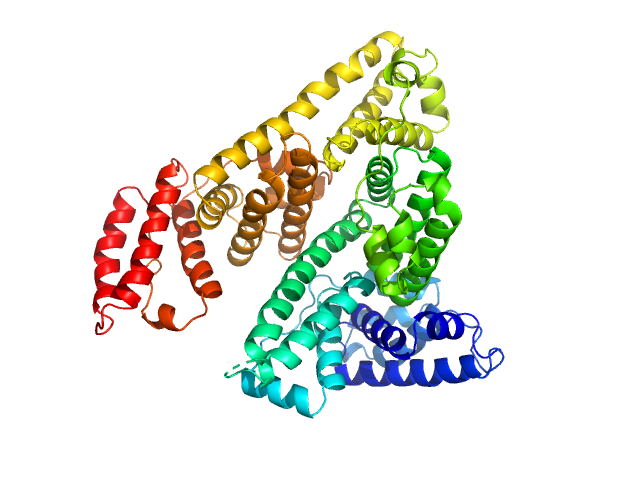
|
| Sample: |
Albumin monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Dec 1
|
Albumin in patients with liver disease shows an altered conformation.
Commun Biol 4(1):731 (2021)
Paar M, Fengler VH, Rosenberg DJ, Krebs A, Stauber RE, Oettl K, Hammel M
|
|
|
UniProt ID: P0DN75 (2-132) Iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Nov 30
|
Magnetic field effects on the structure and molecular behavior of pigeon iron–sulfur protein
Protein Science 31(6) (2022)
Arai S, Shimizu R, Adachi M, Hirai M
|
| RgGuinier |
5.0 |
nm |
| Dmax |
16.2 |
nm |
|
|
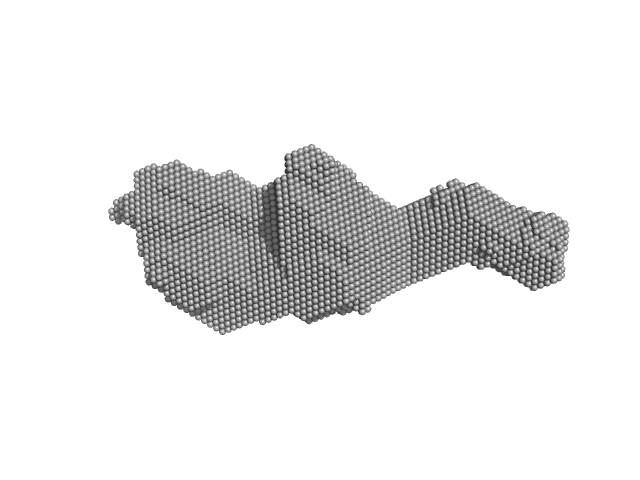
UniProt ID: P74102 (2-317) Orange carotenoid-binding protein
|
|
|

|
| Sample: |
Orange carotenoid-binding protein dimer, 70 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM NaCL, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 23
|
Oligomerization processes limit photoactivation and recovery of the orange carotenoid protein.
Biophys J 121(15):2849-2872 (2022)
Andreeva EA, Niziński S, Wilson A, Levantino M, De Zitter E, Munro R, Muzzopappa F, Thureau A, Zala N, Burdzinski G, Sliwa M, Kirilovsky D, Schirò G, Colletier JP
|
| RgGuinier |
2.9 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
63 |
nm3 |
|
|