UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
|
|
|
|
| Sample: |
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
100 mM NaCl, 20 mM Tris, 47% v/v D₂O, pH: 8 |
| Experiment: |
SANS
data collected at D11, ILL on 2019 Sep 21
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.7 |
nm |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (None-None) DNA-binding protein HU-alpha
|
|
|
|
| Sample: |
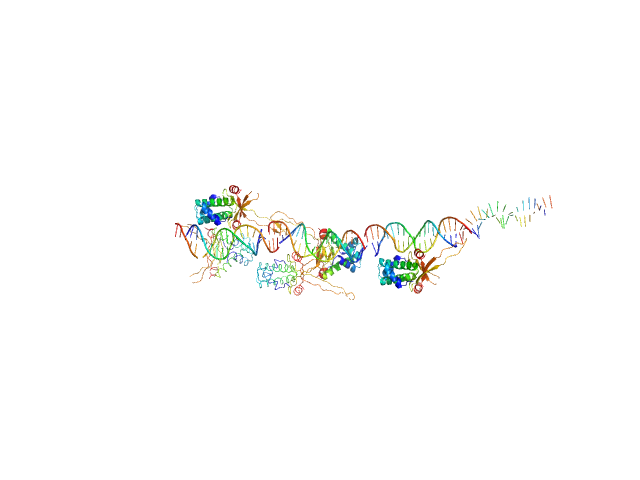
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha 14-mer, 133 kDa Escherichia coli protein
|
| Buffer: |
10 mM Bis-Tris, 150 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jun 1
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
| RgGuinier |
7.0 |
nm |
| Dmax |
26.2 |
nm |
| VolumePorod |
352 |
nm3 |
|
|
UniProt ID: P02768 (None-None) Albumin
|
|
|
|
| Sample: |
Albumin monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Dec 1
|
Albumin in patients with liver disease shows an altered conformation.
Commun Biol 4(1):731 (2021)
Paar M, Fengler VH, Rosenberg DJ, Krebs A, Stauber RE, Oettl K, Hammel M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.2 |
nm |
|
|
UniProt ID: Q9WTS6-4 (343-2708) Isoform A1B0 of Teneurin-3
|
|
|
|
| Sample: |
Isoform A1B0 of Teneurin-3 dimer, 541 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH: 7.8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Sep 10
|
Alternative splicing controls teneurin-3 compact dimer formation for neuronal recognition
Nature Communications 15(1) (2024)
Gogou C, Beugelink J, Frias C, Kresik L, Jaroszynska N, Drescher U, Janssen B, Hindges R, Meijer D
|
| RgGuinier |
7.8 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
1128 |
nm3 |
|
|
UniProt ID: P0DTC9 (176-245) Nucleoprotein (L223P, L227P, L230P)
|
|
|
|
| Sample: |
Nucleoprotein (L223P, L227P, L230P) monomer, 8 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Jan 29
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
2.8 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
14 |
nm3 |
|
|
UniProt ID: None (None-None) β-chitin nanofibers from squid pens
UniProt ID: Q9KLD5 (24-485) GlcNAc-binding protein A (perdeuterated)
|
|
|
|
| Sample: |
Β-chitin nanofibers from squid pens None, Squid
GlcNAc-binding protein A (perdeuterated) monomer, 51 kDa Vibrio cholerae serotype … protein
|
| Buffer: |
20 mM acetate, 47% v/v D₂O, pH: 5 |
| Experiment: |
SANS
data collected at BT5 USANS, NIST Center for Neutron Research on 2020 Dec 4
|
Tangled Up in Fibers: How a Multidomain Lytic Polysaccharide Monooxygenase Binds Its Chitin Substrate.
ACS Appl Mater Interfaces (2026)
Sørensen HV, Montserrat-Canals M, Coder A, Prévost S, Krueger S, Vaaje-Kolstad G, Bjerregaard-Andersen K, Lund R, Krengel U
|
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha
|
|
|
|
| Sample: |
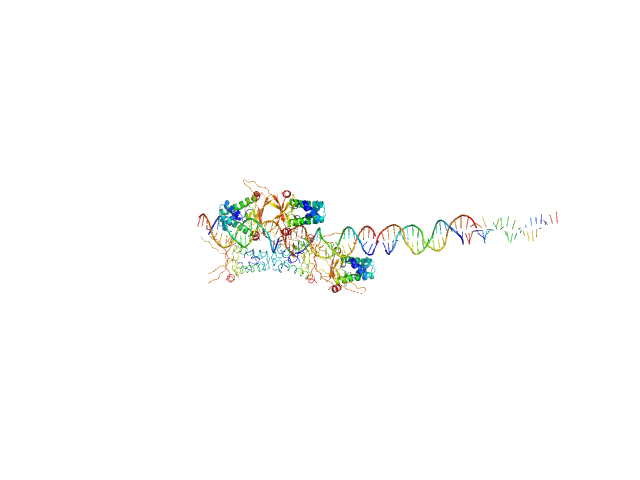
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha decamer, 95 kDa Escherichia coli protein
|
| Buffer: |
10 mM Bis-Tris, 300 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Jun 1
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
| RgGuinier |
6.0 |
nm |
| Dmax |
24.7 |
nm |
| VolumePorod |
218 |
nm3 |
|
|
UniProt ID: P02768 (None-None) Albumin
|
|
|
|
| Sample: |
Albumin monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Dec 1
|
Albumin in patients with liver disease shows an altered conformation.
Commun Biol 4(1):731 (2021)
Paar M, Fengler VH, Rosenberg DJ, Krebs A, Stauber RE, Oettl K, Hammel M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.9 |
nm |
|
|
UniProt ID: P0DTC9 (1-245) Nucleoprotein
|
|
|
|
| Sample: |
Nucleoprotein tetramer, 110 kDa Severe acute respiratory … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 May 13
|
Dynamic ensembles of SARS-CoV-2 N-protein reveal head-to-head coiled-coil-driven oligomerization and phase separation.
Nucleic Acids Res 53(11) (2025)
Hernandez G, Martins ML, Fernandes NP, Veloso T, Lopes J, Gomes T, Cordeiro TN
|
| RgGuinier |
5.2 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
UniProt ID: Q9WTS6-4 (343-2708) Isoform A1B0 of Teneurin-3
|
|
|
|
| Sample: |
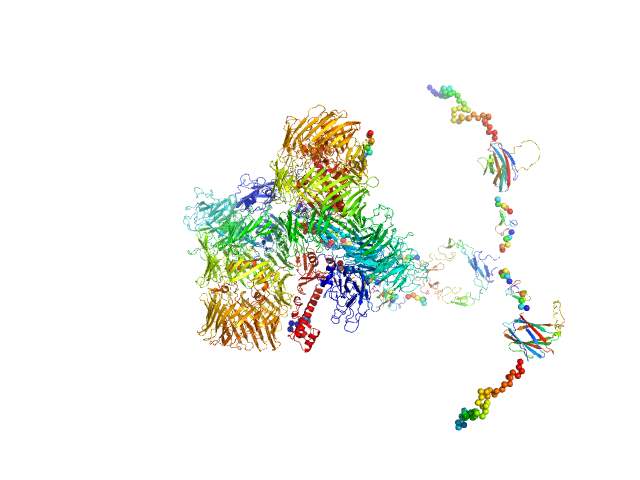
Isoform A1B0 of Teneurin-3 dimer, 541 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2 mM CaCl2, pH: 7.8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2022 Sep 10
|
Alternative splicing controls teneurin-3 compact dimer formation for neuronal recognition
Nature Communications 15(1) (2024)
Gogou C, Beugelink J, Frias C, Kresik L, Jaroszynska N, Drescher U, Janssen B, Hindges R, Meijer D
|
| RgGuinier |
7.7 |
nm |
| Dmax |
30.0 |
nm |
| VolumePorod |
1089 |
nm3 |
|
|