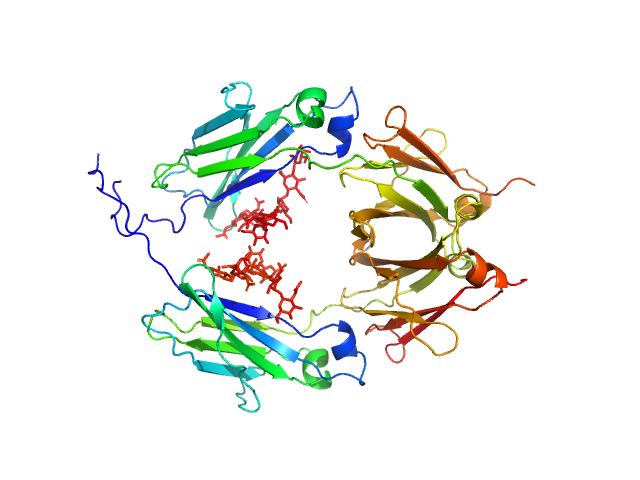
UniProt ID: P01857 (108-329) Glycosylated human immunoglobulin G Fc region
|
|
|
|
| Sample: |
Glycosylated human immunoglobulin G Fc region dimer, 53 kDa Homo sapiens protein
|
| Buffer: |
20 mM Citrate-Phosphate, pH: 7 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2017 Mar 5
|
CH2 domain orientation of human immunoglobulin G in solution: Structural comparison of glycosylated and aglycosylated Fc regions using small-angle X-ray scattering.
MAbs (2018)
Yageta S, Imamura H, Shibuya R, Honda S
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
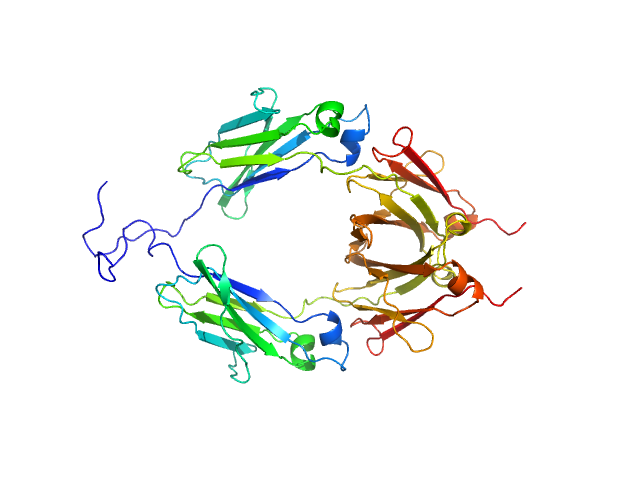
UniProt ID: P01857 (104-330) Aglycosylated human immunoglobulin G Fc region
|
|
|
|
| Sample: |
Aglycosylated human immunoglobulin G Fc region dimer, 51 kDa Homo sapiens protein
|
| Buffer: |
20 mM Citrate-Phosphate, pH: 7 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2017 Mar 5
|
CH2 domain orientation of human immunoglobulin G in solution: Structural comparison of glycosylated and aglycosylated Fc regions using small-angle X-ray scattering.
MAbs (2018)
Yageta S, Imamura H, Shibuya R, Honda S
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
UniProt ID: A0A2K3DZB3 (25-357) Calredoxin, Redox protein from Chlamydomonas reinhardtii
|
|
|
|
| Sample: |
Calredoxin, Redox protein from Chlamydomonas reinhardtii monomer, 40 kDa Chlamydomonas reinhardtii protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM DTT, 5 mM EGTA, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, Structural Biology Laboratory, Graduate School of Medical Life Science, Yokohama City University on 2015 Nov 17
|
Calcium sensing via EF-hand 4 enables thioredoxin activity in the sensor-responder protein calredoxin in the green alga Chlamydomonas reinhardtii.
J Biol Chem (2019)
Charoenwattanasatien R, Zinzius K, Scholz M, Wicke S, Tanaka H, Brandenburg JS, Marchetti GM, Ikegami T, Matsumoto T, Oda T, Sato M, Hippler M, Kurisu G
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.7 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
UniProt ID: A0A2K3DZB3 (25-357) Calredoxin, Redox protein from Chlamydomonas reinhardtii
|
|
|
|
| Sample: |
Calredoxin, Redox protein from Chlamydomonas reinhardtii monomer, 40 kDa Chlamydomonas reinhardtii protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM DTT, 5 mM CaCl2, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, Structural Biology Laboratory, Graduate School of Medical Life Science, Yokohama City University on 2015 Nov 17
|
Calcium sensing via EF-hand 4 enables thioredoxin activity in the sensor-responder protein calredoxin in the green alga Chlamydomonas reinhardtii.
J Biol Chem (2019)
Charoenwattanasatien R, Zinzius K, Scholz M, Wicke S, Tanaka H, Brandenburg JS, Marchetti GM, Ikegami T, Matsumoto T, Oda T, Sato M, Hippler M, Kurisu G
|
| RgGuinier |
3.1 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
68 |
nm3 |
|
|
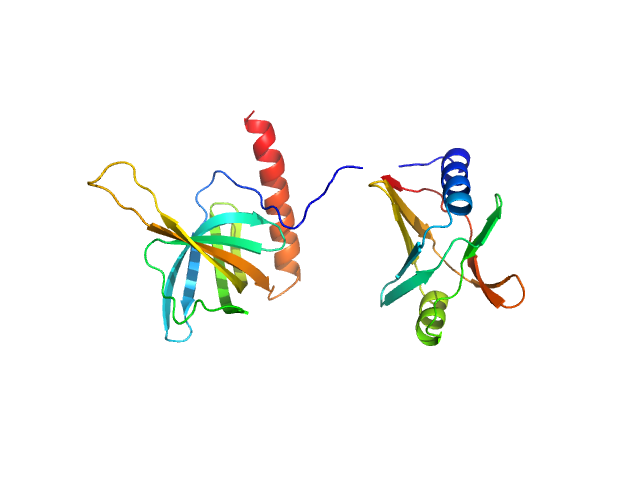
UniProt ID: P76010 (None-None) Flagellar brake protein YcgR
|
|
|
|
| Sample: |
Flagellar brake protein YcgR monomer, 29 kDa Escherichia coli protein
|
| Buffer: |
20 mM HEPES, 150mM NaCl, 10% glycerol,, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2016 Jan 4
|
Structural insights into the mechanism of c-di-GMP-bound YcgR regulating flagellar motility in Escherichia coli.
J Biol Chem 295(3):808-821 (2020)
Hou YJ, Yang WS, Hong Y, Zhang Y, Wang DC, Li DF
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
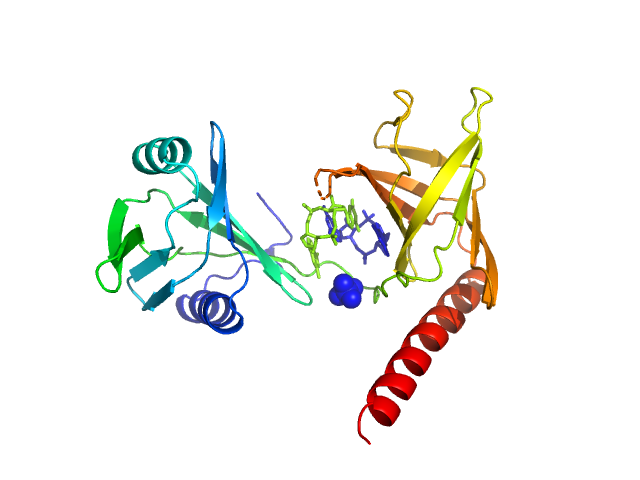
UniProt ID: P76010 (None-None) Flagellar brake protein YcgR in complex with c-di-GMP
|
|
|
|
| Sample: |
Flagellar brake protein YcgR in complex with c-di-GMP monomer, 29 kDa Escherichia coli protein
|
| Buffer: |
20 mM HEPES, 150mM NaCl, 10% glycerol,, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2016 Jan 4
|
Structural insights into the mechanism of c-di-GMP-bound YcgR regulating flagellar motility in Escherichia coli.
J Biol Chem 295(3):808-821 (2020)
Hou YJ, Yang WS, Hong Y, Zhang Y, Wang DC, Li DF
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.3 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
UniProt ID: Q5EPC7 (None-None) Relaxase (Tra_2) domain of TraI
|
|
|
|
| Sample: |
Relaxase (Tra_2) domain of TraI monomer, 46 kDa Neisseria gonorrhoeae protein
|
| Buffer: |
50 mM TRIS-HCl 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 11
|
DNA processing by the MOBH family relaxase TraI encoded within the gonococcal genetic island.
Nucleic Acids Res 47(15):8136-8153 (2019)
Heilers JH, Reiners J, Heller EM, Golzer A, Smits SHJ, van der Does C
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
UniProt ID: Q5EPC7 (674-850) TraI_2_C domain of TraI
|
|
|
|
| Sample: |
TraI_2_C domain of TraI monomer, 21 kDa Neisseria gonorrhoeae protein
|
| Buffer: |
50 mM TRIS-HCl 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 11
|
DNA processing by the MOBH family relaxase TraI encoded within the gonococcal genetic island.
Nucleic Acids Res 47(15):8136-8153 (2019)
Heilers JH, Reiners J, Heller EM, Golzer A, Smits SHJ, van der Does C
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
UniProt ID: Q5EPC7 (42-850) TraI
|
|
|

|
| Sample: |
TraI monomer, 91 kDa Neisseria gonorrhoeae protein
|
| Buffer: |
50 mM TRIS-HCl 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Mar 5
|
DNA processing by the MOBH family relaxase TraI encoded within the gonococcal genetic island.
Nucleic Acids Res 47(15):8136-8153 (2019)
Heilers JH, Reiners J, Heller EM, Golzer A, Smits SHJ, van der Does C
|
| RgGuinier |
7.3 |
nm |
| Dmax |
31.4 |
nm |
| VolumePorod |
293 |
nm3 |
|
|
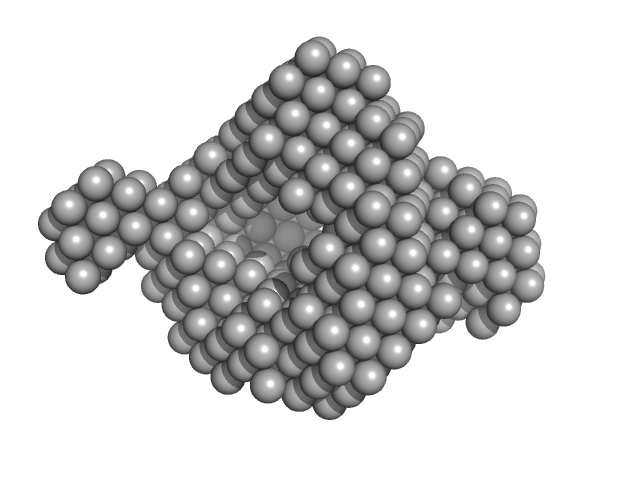
UniProt ID: P21177 (None-None) Fatty acid oxidation complex subunit alpha
UniProt ID: P21177 (None-None) Fatty acid oxidation complex subunit alpha
UniProt ID: P21151 (None-None) 3-ketoacyl-CoA thiolase FadA (beta subunit)
|
|
|

|
| Sample: |
Fatty acid oxidation complex subunit alpha monomer, 81 kDa Escherichia coli protein
Fatty acid oxidation complex subunit alpha monomer, 81 kDa Escherichia coli protein
3-ketoacyl-CoA thiolase FadA (beta subunit) dimer, 82 kDa Escherichia coli protein
|
| Buffer: |
20 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), 120 mM KCl, 2.5 mM DTT, pH: 7.2 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 May 30
|
Complementary substrate specificity and distinct quaternary assembly of the Escherichia coli aerobic and anaerobic beta-oxidation trifunctional enzyme complexes.
Biochem J (2019)
Sah-Teli SK, Hynönen MJ, Schmitz W, Geraets JA, Seitsonen J, Pedersen JS, Butcher SJ, Wierenga RK, Venkatesan R
|
| RgGuinier |
4.6 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
406 |
nm3 |
|
|