UniProt ID: P77399 (1-714) Fatty acid oxidation complex subunit alpha
UniProt ID: P77399 (None-None) anaerobic Fatty acid oxidation complex subunit alpha
UniProt ID: P77399 (None-None) anaerobic Fatty acid oxidation complex subunit alpha
UniProt ID: P77399 (None-None) anaerobic Fatty acid oxidation complex subunit alpha
UniProt ID: P76503 (None-None) anaerobic 3-ketoacyl-CoA thiolase FadI beta subunit
UniProt ID: P76503 (None-None) anaerobic 3-ketoacyl-CoA thiolase FadI beta subunit
|
|
|

|
| Sample: |
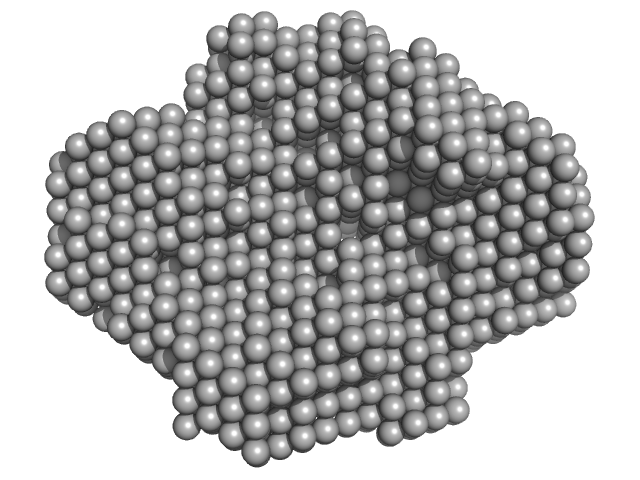
Fatty acid oxidation complex subunit alpha monomer, 77 kDa Escherichia coli (strain … protein
Anaerobic Fatty acid oxidation complex subunit alpha monomer, 77 kDa Escherichia coli protein
Anaerobic Fatty acid oxidation complex subunit alpha monomer, 77 kDa Escherichia coli protein
Anaerobic Fatty acid oxidation complex subunit alpha monomer, 77 kDa Escherichia coli protein
Anaerobic 3-ketoacyl-CoA thiolase FadI beta subunit dimer, 96 kDa Escherichia coli protein
Anaerobic 3-ketoacyl-CoA thiolase FadI beta subunit dimer, 96 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris, 500 mM NaCl, 5% glycerol, 0.05% C12E9 (1-O-(n-Dodecyl)-nonaethyleneglycol), 2.5 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Sep 22
|
Complementary substrate specificity and distinct quaternary assembly of the Escherichia coli aerobic and anaerobic beta-oxidation trifunctional enzyme complexes.
Biochem J (2019)
Sah-Teli SK, Hynönen MJ, Schmitz W, Geraets JA, Seitsonen J, Pedersen JS, Butcher SJ, Wierenga RK, Venkatesan R
|
| RgGuinier |
6.2 |
nm |
| Dmax |
19.6 |
nm |
| VolumePorod |
856 |
nm3 |
|
|
UniProt ID: Q7MVK4 (1-227) Hcp Transcriptional regulator
|
|
|
|
| Sample: |
Hcp Transcriptional regulator dimer, 51 kDa Porphyromonas gingivalis protein
|
| Buffer: |
25 mM Tris 150 mM NaCl 1mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Jan 24
|
Nitrosative stress sensing in Porphyromonas gingivalis: structure of and heme binding by the transcriptional regulator HcpR.
Acta Crystallogr D Struct Biol 75(Pt 4):437-450 (2019)
Belvin BR, Musayev FN, Burgner J, Scarsdale JN, Escalante CR, Lewis JP
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: Q8N884 (1-522) Cyclic GMP-AMP synthase
|
|
|
|
| Sample: |
Cyclic GMP-AMP synthase monomer, 61 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Apr 25
|
cGAS facilitates sensing of extracellular cyclic dinucleotides to activate innate immunity.
EMBO Rep (2019)
Liu H, Moura-Alves P, Pei G, Mollenkopf HJ, Hurwitz R, Wu X, Wang F, Liu S, Ma M, Fei Y, Zhu C, Koehler AB, Oberbeck-Mueller D, Hahnke K, Klemm M, Guhlich-Bornhof U, Ge B, Tuukkanen A, Kolbe M, Dorhoi A, Kaufmann SH
|
| RgGuinier |
3.1 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
UniProt ID: Q8N884 (1-522) Cyclic GMP-AMP synthase
UniProt ID: None (None-None) 2'-O,5'-O-((adenosine-3'-O,5'-O-diyl)bisphosphinico)guanosine
|
|
|
|
| Sample: |
Cyclic GMP-AMP synthase dimer, 123 kDa Homo sapiens protein
2'-O,5'-O-((adenosine-3'-O,5'-O-diyl)bisphosphinico)guanosine dimer, 1 kDa
|
| Buffer: |
20 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Apr 25
|
cGAS facilitates sensing of extracellular cyclic dinucleotides to activate innate immunity.
EMBO Rep (2019)
Liu H, Moura-Alves P, Pei G, Mollenkopf HJ, Hurwitz R, Wu X, Wang F, Liu S, Ma M, Fei Y, Zhu C, Koehler AB, Oberbeck-Mueller D, Hahnke K, Klemm M, Guhlich-Bornhof U, Ge B, Tuukkanen A, Kolbe M, Dorhoi A, Kaufmann SH
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
127 |
nm3 |
|
|
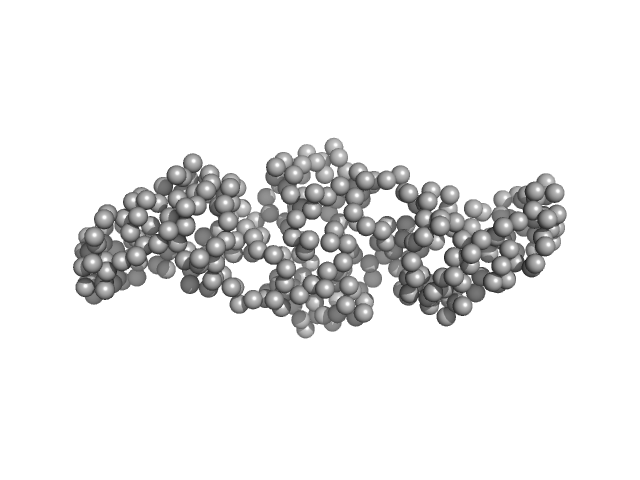
UniProt ID: Q1DB03 (1-159) Gliding motility protein MglB
|
|
|
|
| Sample: |
Gliding motility protein MglB dimer, 34 kDa Myxococcus xanthus protein
|
| Buffer: |
150 mM NaCl, 1 mM DTT, 20 mM Tris-HCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Oct 9
|
MglA functions as a three-state GTPase to control movement reversals of Myxococcus xanthus.
Nat Commun 10(1):5300 (2019)
Galicia C, Lhospice S, Varela PF, Trapani S, Zhang W, Navaza J, Herrou J, Mignot T, Cherfils J
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: Q5ZXN6 (None-None) Phosphocholinase AnkX
|
|
|

|
| Sample: |
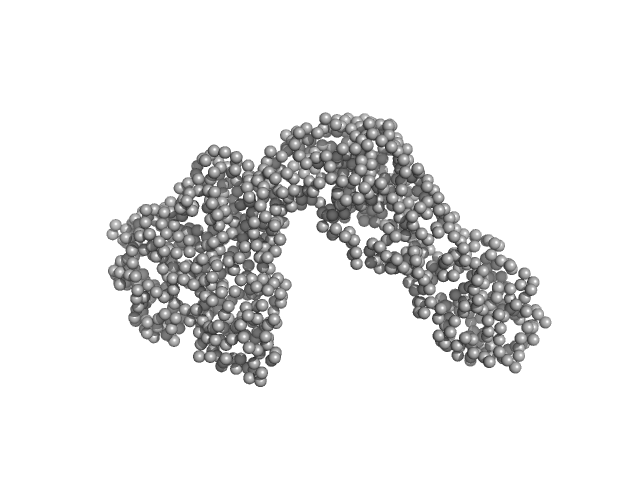
Phosphocholinase AnkX monomer, 107 kDa Legionella pneumophila protein
|
| Buffer: |
300 mM NaCl, 2 mM 2-mercaptoethanol and 30 mM Tris-HCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Oct 20
|
Legionella pneumophila effector AnkX
Wenhua Zhang
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.7 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
UniProt ID: Q99418 (58-400) Cytohesin-2; ARNO truncation mutant
|
|
|

|
| Sample: |
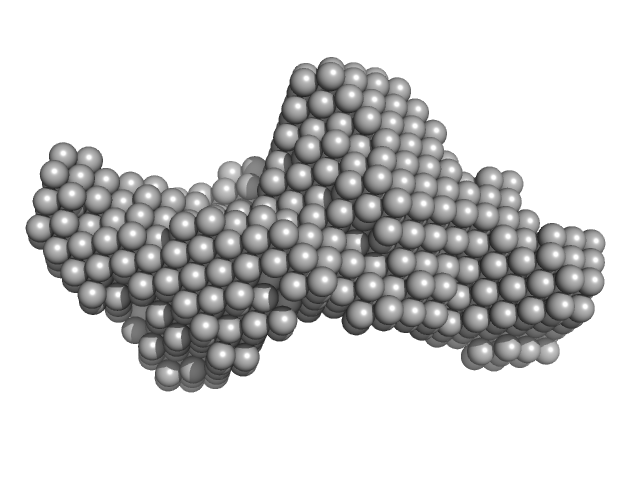
Cytohesin-2; ARNO truncation mutant monomer, 40 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 2 mM 2-mercaptoethanol and 30 mM Tris-HCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2015 Nov 25
|
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Structure (2019)
Das S, Malaby AW, Nawrotek A, Zhang W, Zeghouf M, Maslen S, Skehel M, Chakravarthy S, Irving TC, Bilsel O, Cherfils J, Lambright DG
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
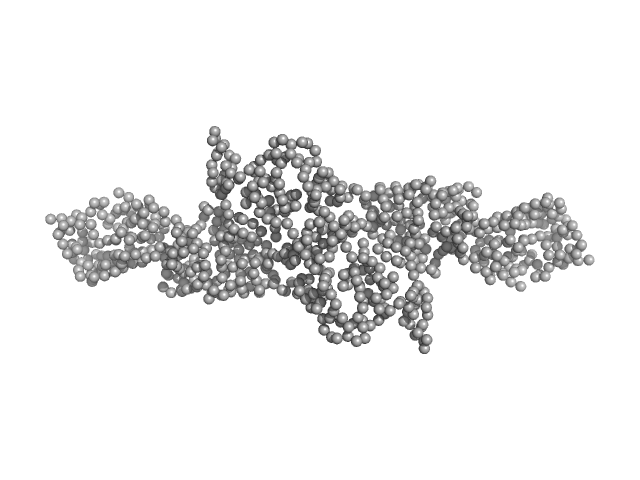
UniProt ID: Q99418 (1-400) Cytohesin-2 ARF nucleotide-binding site opener
|
|
|
|
| Sample: |
Cytohesin-2 ARF nucleotide-binding site opener dimer, 93 kDa Homo sapiens protein
|
| Buffer: |
300 mM NaCl, 2 mM 2-mercaptoethanol and 30 mM Tris-HCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jun 23
|
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Structure (2019)
Das S, Malaby AW, Nawrotek A, Zhang W, Zeghouf M, Maslen S, Skehel M, Chakravarthy S, Irving TC, Bilsel O, Cherfils J, Lambright DG
|
| RgGuinier |
4.8 |
nm |
| Dmax |
19.7 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
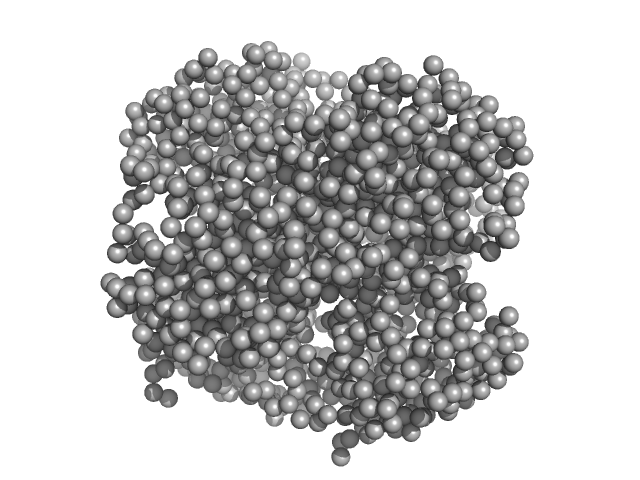
UniProt ID: None (None-None) HpcH/HpaI aldolase
|
|
|
|
| Sample: |
HpcH/HpaI aldolase hexamer, 165 kDa Rhizorhabdus wittichii RW1 protein
|
| Buffer: |
20 mM HEPES,, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Nov 23
|
EFAMIX
, a tool to decompose inline chromatography SAXS
data from partially overlapping components
Protein Science (2021)
Konarev P, Graewert M, Jeffries C, Fukuda M, Cheremnykh T, Volkov V, Svergun D
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
233 |
nm3 |
|
|
UniProt ID: P01241 (27-217) somapacitan
UniProt ID: P02768 (25-609) FcRn binding optimised human serum albumin V418M, T420A, E505G, V547A
UniProt ID: P61768 (21-119) neonatal Fc receptor ectodomain beta-microglogulin part with C-terminal His6 tag
UniProt ID: P55899 (24-297) neonatal Fc receptor ectodomain alpha-chain
|
|
|
|
| Sample: |
Somapacitan monomer, 22 kDa Homo sapiens protein
FcRn binding optimised human serum albumin V418M, T420A, E505G, V547A monomer, 66 kDa Homo sapiens protein
Neonatal Fc receptor ectodomain beta-microglogulin part with C-terminal His6 tag monomer, 13 kDa Homo sapiens protein
Neonatal Fc receptor ectodomain alpha-chain monomer, 30 kDa Homo sapiens protein
|
| Buffer: |
100 mM MES, 100 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 May 11
|
Identification of binding sites on human serum albumin for somapacitan - a long-acting growth hormone derivative.
Biochemistry (2020)
Johansson E, Nielsen AD, Demuth H, Wiberg C, Schjødt C, Huang T, Chen J, Jensen S, Petersen J, Thygesen P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.7 |
nm |
| VolumePorod |
227 |
nm3 |
|
|