UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jun 20
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
4.7 |
nm |
| Dmax |
17.7 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
UniProt ID: P28700 (230-467) Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD)
UniProt ID: P10276 (173-421) Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB)
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD) monomer, 26 kDa Mus musculus protein
Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB) monomer, 28 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Jul 23
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
4.8 |
nm |
| Dmax |
19.4 |
nm |
| VolumePorod |
167 |
nm3 |
|
|
UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
UniProt ID: P28700 (230-467) Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD)
UniProt ID: P28700 (227-439) Retinoid-X receptor alpha (RXR-alpha) Δ helix12
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD) monomer, 26 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Δ helix12 monomer, 24 kDa Mus musculus protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Jul 23
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
15.7 |
nm |
| VolumePorod |
183 |
nm3 |
|
|
UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
UniProt ID: P28700 (230-467) Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD)
UniProt ID: P10276 (173-421) Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB)
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD) monomer, 26 kDa Mus musculus protein
Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB) monomer, 28 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Jul 23
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
4.8 |
nm |
| Dmax |
19.5 |
nm |
| VolumePorod |
178 |
nm3 |
|
|
UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
UniProt ID: P28700 (230-467) Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD)
UniProt ID: P10276 (173-421) Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB) mutant I396E
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD) monomer, 26 kDa Mus musculus protein
Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB) mutant I396E monomer, 28 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Mar 9
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
5.3 |
nm |
| Dmax |
22.4 |
nm |
| VolumePorod |
171 |
nm3 |
|
|
UniProt ID: Q60974 (2059-2325) Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID)
UniProt ID: P28700 (230-467) Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD)
UniProt ID: P10276 (173-421) Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB)
|
|
|
|
| Sample: |
Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) monomer, 29 kDa Mus musculus protein
Retinoid-X receptor alpha (RXR-alpha) Ligand Binding Domain (LBD) monomer, 26 kDa Mus musculus protein
Retinoic acid receptor alpha (RAR-alpha) Ligand binding domain (LDB) monomer, 28 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Jul 23
|
Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression.
Structure (2019)
Cordeiro TN, Sibille N, Germain P, Barthe P, Boulahtouf A, Allemand F, Bailly R, Vivat V, Ebel C, Barducci A, Bourguet W, le Maire A, Bernadó P
|
| RgGuinier |
4.2 |
nm |
| Dmax |
17.2 |
nm |
| VolumePorod |
131 |
nm3 |
|
|
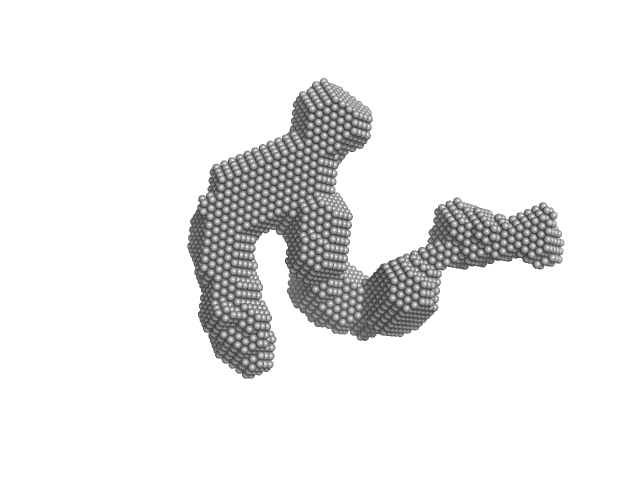
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|

|
| Sample: |
Condensin complex subunit 3-like protein monomer, 108 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
236 |
nm3 |
|
|
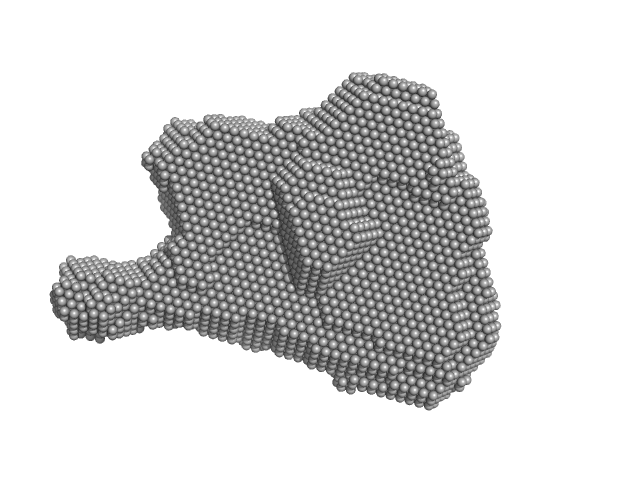
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
UniProt ID: G0SBJ6 (515-634) Condensin complex subunit 2
|
|
|

|
| Sample: |
Condensin complex subunit 3-like protein monomer, 108 kDa Chaetomium thermophilum protein
Condensin complex subunit 2 monomer, 17 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
4.3 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
230 |
nm3 |
|
|
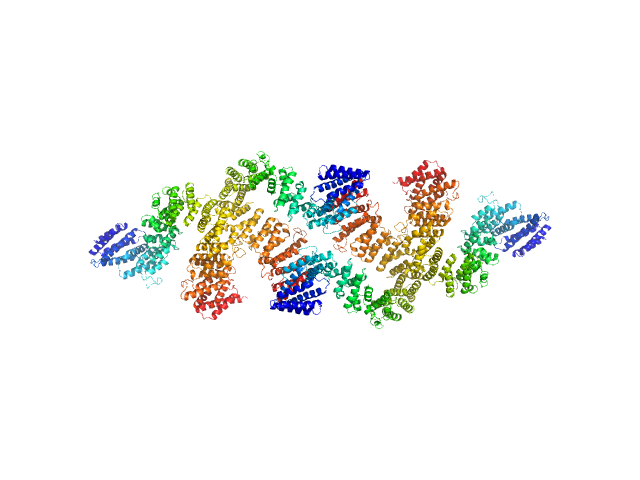
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
Condensin complex subunit 3-like protein tetramer, 433 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
8.3 |
nm |
| Dmax |
32.7 |
nm |
| VolumePorod |
1080 |
nm3 |
|
|
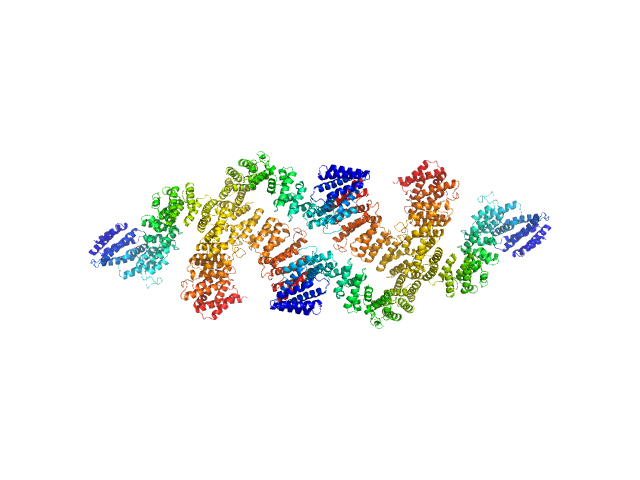
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
Condensin complex subunit 3-like protein tetramer, 433 kDa Chaetomium thermophilum protein
Condensin complex subunit 3-like protein dimer, 217 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
6.8 |
nm |
| Dmax |
29.8 |
nm |
| VolumePorod |
720 |
nm3 |
|
|