UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
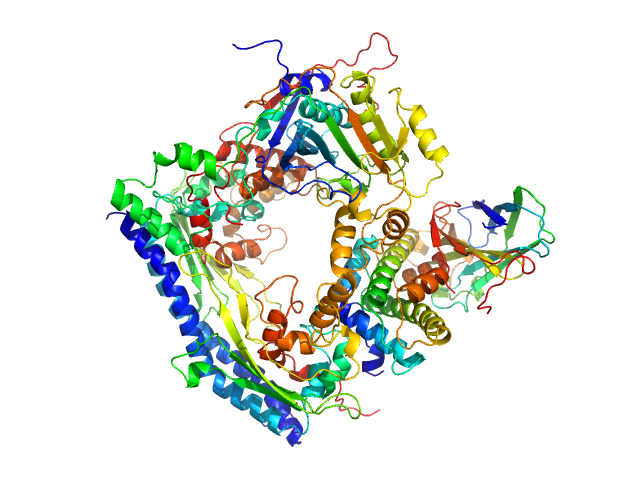
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 42% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Mar 5
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.0 |
nm |
|
|
UniProt ID: P53853 (1-225) Vacuolar protein sorting-associated protein 75 (1-225 aa)
UniProt ID: Q07794 (1-436) Histone acetyltransferase RTT109
UniProt ID: P32447 (1-169) Histone chaperone ASF1
UniProt ID: P84233 (35-135) Histone H3.2 (35-135 aa)
UniProt ID: P62799 (1-103) Histone H4
|
|
|
|
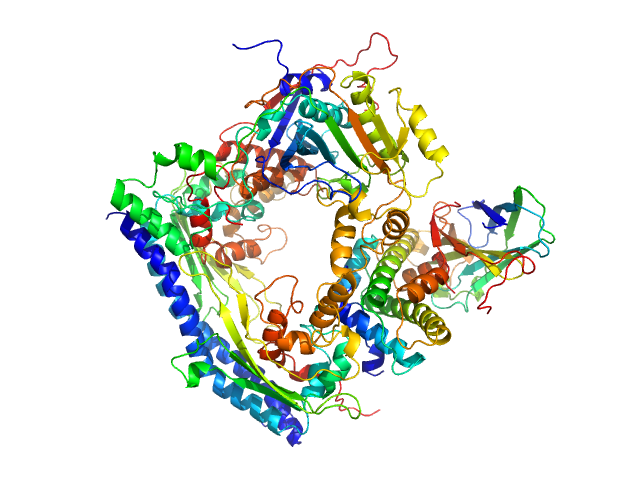
| Sample: |
Vacuolar protein sorting-associated protein 75 (1-225 aa) dimer, 53 kDa Saccharomyces cerevisiae protein
Histone acetyltransferase RTT109 monomer, 50 kDa Saccharomyces cerevisiae protein
Histone chaperone ASF1 monomer, 19 kDa protein
Histone H3.2 (35-135 aa) monomer, 12 kDa Xenopus laevis protein
Histone H4 monomer, 11 kDa Xenopus laevis protein
|
| Buffer: |
50 mM citrate, 150 mM NaCl, 5 mM BME, 42% D2O, pH: 6.5 |
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Mar 5
|
Histone chaperone exploits intrinsic disorder to switch acetylation specificity.
Nat Commun 10(1):3435 (2019)
Danilenko N, Lercher L, Kirkpatrick J, Gabel F, Codutti L, Carlomagno T
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.5 |
nm |
|
|
UniProt ID: Q62318 (58-418) Transcription intermediary factor 1-beta
|
|
|

|
| Sample: |
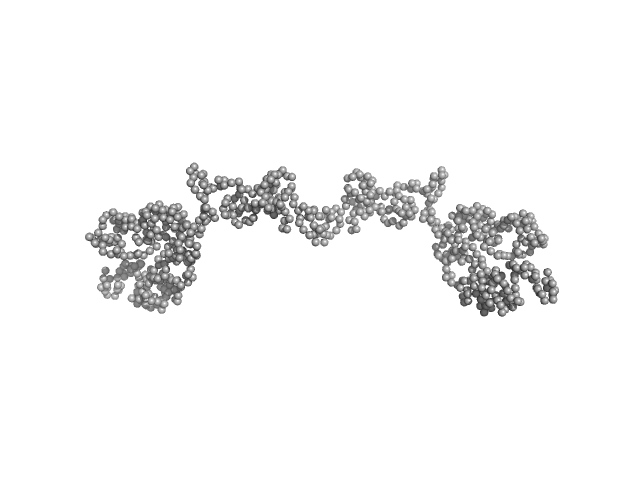
Transcription intermediary factor 1-beta dimer, 82 kDa Mus musculus protein
|
| Buffer: |
10 mM Tris 300 mM NaCl 0.1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Aug 10
|
A Dissection of Oligomerisation by the TRIM28 Tripartite Motif and the Interaction with Members of the Krab-ZFP Family.
J Mol Biol (2019)
Sun Y, Keown JR, Black MM, Raclot C, Demarais N, Trono D, Turelli P, Goldstone DC
|
| RgGuinier |
7.0 |
nm |
| Dmax |
23.2 |
nm |
| VolumePorod |
232 |
nm3 |
|
|
UniProt ID: Q62318 (58-418) Transcription intermediary factor 1-beta
UniProt ID: G3X9G7 (1-74) Zinc finger protein 809 N-terminal MBP fusion
|
|
|
|
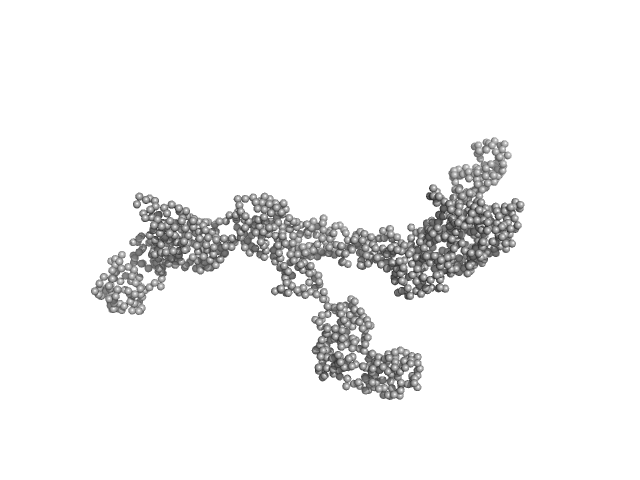
| Sample: |
Transcription intermediary factor 1-beta dimer, 82 kDa Mus musculus protein
Zinc finger protein 809 N-terminal MBP fusion monomer, 52 kDa Mus musculus protein
|
| Buffer: |
10 mM Tris 300 mM NaCl 0.1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Aug 10
|
A Dissection of Oligomerisation by the TRIM28 Tripartite Motif and the Interaction with Members of the Krab-ZFP Family.
J Mol Biol (2019)
Sun Y, Keown JR, Black MM, Raclot C, Demarais N, Trono D, Turelli P, Goldstone DC
|
| RgGuinier |
6.4 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
252 |
nm3 |
|
|
UniProt ID: Q8NB78 (31-822) Lysyne-specific Demethylase LSD2
|
|
|
|
| Sample: |
Lysyne-specific Demethylase LSD2 monomer, 89 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 200 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 11
|
A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex.
Cell Rep 27(2):387-399.e7 (2019)
Marabelli C, Marrocco B, Pilotto S, Chittori S, Picaud S, Marchese S, Ciossani G, Forneris F, Filippakopoulos P, Schoehn G, Rhodes D, Subramaniam S, Mattevi A
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
124 |
nm3 |
|
|
UniProt ID: Q49A26 (261-553) NPAC dehydrogenase domain
|
|
|
|
| Sample: |
NPAC dehydrogenase domain tetramer, 125 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 200 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jan 26
|
A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex.
Cell Rep 27(2):387-399.e7 (2019)
Marabelli C, Marrocco B, Pilotto S, Chittori S, Picaud S, Marchese S, Ciossani G, Forneris F, Filippakopoulos P, Schoehn G, Rhodes D, Subramaniam S, Mattevi A
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
167 |
nm3 |
|
|
UniProt ID: Q49A26 (205-553) NPAC linker+DH (delta-205)
|
|
|
|
| Sample: |
NPAC linker+DH (delta-205) tetramer, 150 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 200 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jan 26
|
A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex.
Cell Rep 27(2):387-399.e7 (2019)
Marabelli C, Marrocco B, Pilotto S, Chittori S, Picaud S, Marchese S, Ciossani G, Forneris F, Filippakopoulos P, Schoehn G, Rhodes D, Subramaniam S, Mattevi A
|
| RgGuinier |
4.0 |
nm |
| Dmax |
11.4 |
nm |
| VolumePorod |
240 |
nm3 |
|
|
UniProt ID: None (2-136) Histone H3
UniProt ID: P62799 (2-103) Histone H4
UniProt ID: Q6AZJ8 (2-130) Histone H2a
UniProt ID: Q92130 (5-126) Histone H2b
UniProt ID: None (None-None) 147bp 601 Widom sequence
|
|
|
|
| Sample: |
Histone H3 dimer, 31 kDa Xenopus laevis protein
Histone H4 dimer, 22 kDa Xenopus laevis protein
Histone H2a dimer, 28 kDa Xenopus laevis protein
Histone H2b dimer, 27 kDa Xenopus laevis protein
147bp 601 Widom sequence monomer, 45 kDa synthetic construct DNA
|
| Buffer: |
15 mM HEPES, 200 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 11
|
A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex.
Cell Rep 27(2):387-399.e7 (2019)
Marabelli C, Marrocco B, Pilotto S, Chittori S, Picaud S, Marchese S, Ciossani G, Forneris F, Filippakopoulos P, Schoehn G, Rhodes D, Subramaniam S, Mattevi A
|
| RgGuinier |
4.4 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
353 |
nm3 |
|
|
UniProt ID: Q8NB78 (31-822) Lysyne-specific Demethylase LSD2
UniProt ID: Q49A26 (205-553) NPAC linker+DH (delta-205)
UniProt ID: None (2-136) Histone H3
UniProt ID: P62799 (2-103) Histone H4
UniProt ID: Q6AZJ8 (2-130) Histone H2a
UniProt ID: Q92130 (5-126) Histone H2b
UniProt ID: None (None-None) 147bp 601 Widom sequence
|
|
|
|
| Sample: |
Lysyne-specific Demethylase LSD2 monomer, 89 kDa Homo sapiens protein
NPAC linker+DH (delta-205) tetramer, 150 kDa Homo sapiens protein
Histone H3 dimer, 31 kDa Xenopus laevis protein
Histone H4 dimer, 22 kDa Xenopus laevis protein
Histone H2a dimer, 28 kDa Xenopus laevis protein
Histone H2b dimer, 27 kDa Xenopus laevis protein
147bp 601 Widom sequence monomer, 45 kDa synthetic construct DNA
|
| Buffer: |
15 mM HEPES, 200 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 11
|
A Tail-Based Mechanism Drives Nucleosome Demethylation by the LSD2/NPAC Multimeric Complex.
Cell Rep 27(2):387-399.e7 (2019)
Marabelli C, Marrocco B, Pilotto S, Chittori S, Picaud S, Marchese S, Ciossani G, Forneris F, Filippakopoulos P, Schoehn G, Rhodes D, Subramaniam S, Mattevi A
|
| RgGuinier |
7.8 |
nm |
| Dmax |
30.4 |
nm |
| VolumePorod |
1100 |
nm3 |
|
|
UniProt ID: P98170 (1-497) E3 ubiquitin-protein ligase XIAP
|
|
|
|
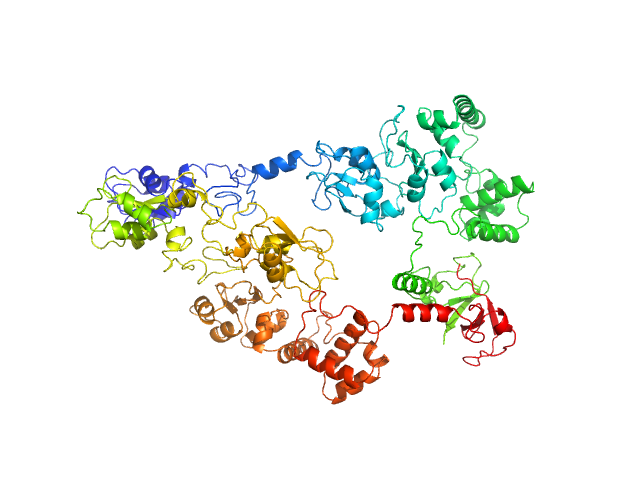
| Sample: |
E3 ubiquitin-protein ligase XIAP dimer, 113 kDa Homo sapiens protein
|
| Buffer: |
Xiap buffer, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 15
|
Conformational characterization of full-length X-chromosome-linked inhibitor of apoptosis protein (XIAP) through an integrated approach.
IUCrJ 6(Pt 5):948-957 (2019)
Polykretis P, Luchinat E, Bonucci A, Giachetti A, Graewert MA, Svergun DI, Banci L
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
207 |
nm3 |
|
|