UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
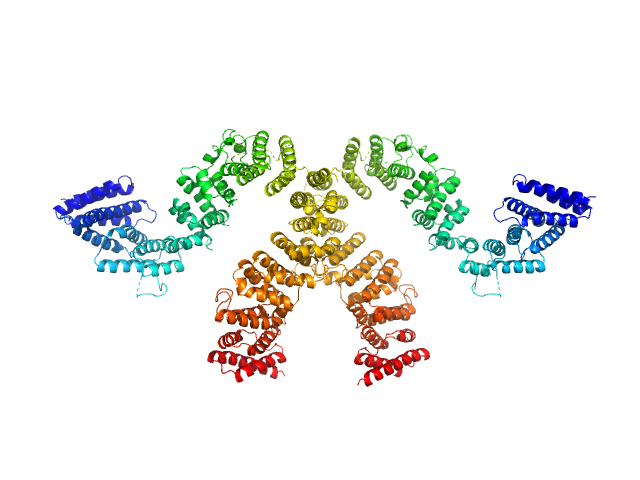
Condensin complex subunit 3-like protein dimer, 217 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
5.4 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
463 |
nm3 |
|
|
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
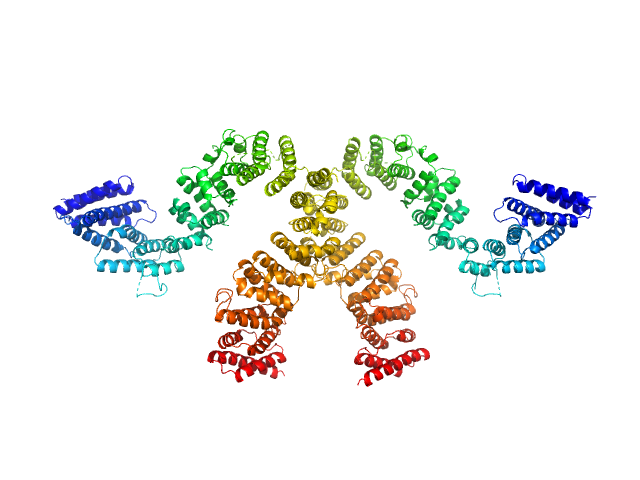
Condensin complex subunit 3-like protein monomer, 108 kDa Chaetomium thermophilum protein
Condensin complex subunit 3-like protein dimer, 217 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
5.3 |
nm |
| Dmax |
18.7 |
nm |
| VolumePorod |
400 |
nm3 |
|
|
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
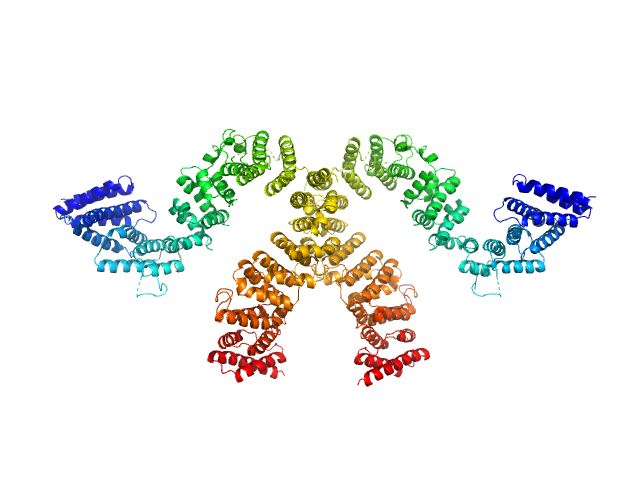
Condensin complex subunit 3-like protein monomer, 108 kDa Chaetomium thermophilum protein
Condensin complex subunit 3-like protein dimer, 217 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
4.7 |
nm |
| Dmax |
16.1 |
nm |
| VolumePorod |
301 |
nm3 |
|
|
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
UniProt ID: G0SHK2 (24-1006) Condensin complex subunit 3-like protein
|
|
|
|
| Sample: |
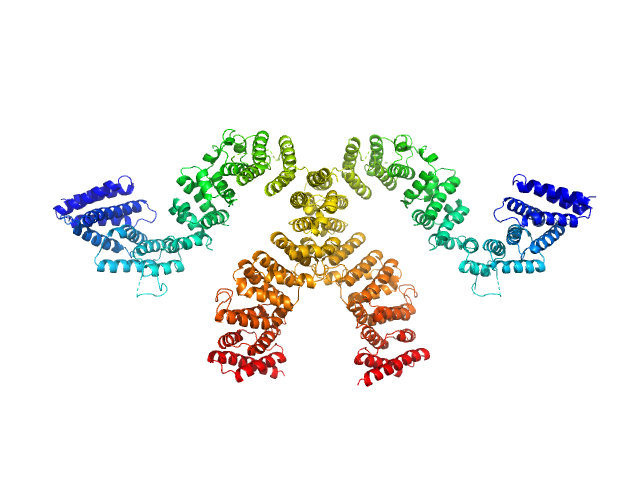
Condensin complex subunit 3-like protein monomer, 108 kDa Chaetomium thermophilum protein
Condensin complex subunit 3-like protein dimer, 217 kDa Chaetomium thermophilum protein
|
| Buffer: |
25 mM Tris, 300 mM NaCl, 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 20
|
Solution structure and flexibility of the condensin HEAT-repeat subunit Ycg1.
J Biol Chem 294(37):13822-13829 (2019)
Manalastas-Cantos K, Kschonsak M, Haering CH, Svergun DI
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
280 |
nm3 |
|
|
UniProt ID: Q7Q7F7 (25-177) AGAP005335-PA
UniProt ID: Q7Q7F8 (18-174) AGAP005334-PA
|
|
|
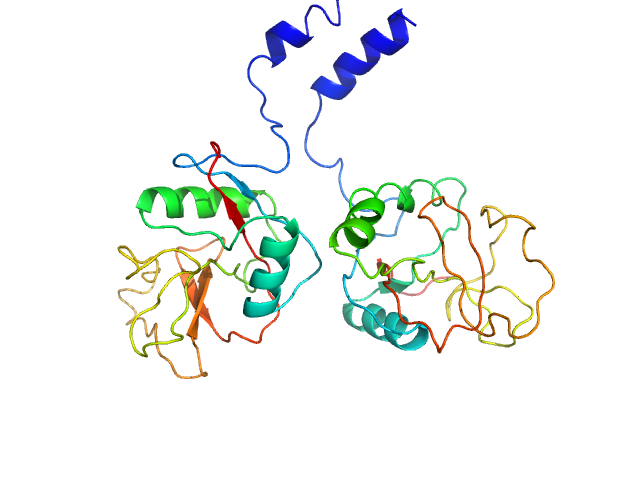
|
| Sample: |
AGAP005335-PA monomer, 18 kDa Anopheles gambiae protein
AGAP005334-PA monomer, 18 kDa Anopheles gambiae protein
|
| Buffer: |
500 mM NaCl, 20 mM CHES, 0.5 mM CaCl2, 1% glycerol, pH: 9 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Thomas Jefferson University on 2018 Aug 30
|
Solution structure, glycan specificity and of phenol oxidase inhibitory activity of Anopheles C-type lectins CTL4 and CTLMA2.
Sci Rep 9(1):15191 (2019)
Bishnoi R, Sousa GL, Contet A, Day CJ, Hou CD, Profitt LA, Singla D, Jennings MP, Valentine AM, Povelones M, Baxter RHG
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
UniProt ID: P03956 (100-469) Matrix metalloproteinase-1 (Interstitial collagenase)
|
|
|
|
| Sample: |
Matrix metalloproteinase-1 (Interstitial collagenase) monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM Sodium chloride, 10 mM Calcium chloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 May 14
|
Conformationally constrained mutant of human matrix metalloproteinase-1
Rob Holland
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
UniProt ID: P03956 (100-469) Matrix metalloproteinase-1 (Interstitial collagenase) (S243C, S318C)
|
|
|
|
| Sample: |
Matrix metalloproteinase-1 (Interstitial collagenase) (S243C, S318C) monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM Sodium chloride, 10 mM Calcium chloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 May 14
|
Conformationally constrained mutant of human matrix metalloproteinase-1
Rob Holland
|
| RgGuinier |
2.6 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
43 |
nm3 |
|
|
UniProt ID: P03956 (20-469) Pro-matrix metalloproteinase-1 (Interstitial collagenase)
|
|
|
|
| Sample: |
Pro-matrix metalloproteinase-1 (Interstitial collagenase) monomer, 52 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM Sodium chloride, 10 mM Calcium chloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2015 May 21
|
Conformationally constrained mutant of human matrix metalloproteinase-1
Rob Holland
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
73 |
nm3 |
|
|
UniProt ID: P03956 (20-469) Pro-matrix metalloproteinase-1 (Interstitial collagenase) (proMMP-1 S243C, S318C)
|
|
|
|
| Sample: |
Pro-matrix metalloproteinase-1 (Interstitial collagenase) (proMMP-1 S243C, S318C) monomer, 52 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM Sodium chloride, 10 mM Calcium chloride, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2015 May 21
|
Conformationally constrained mutant of human matrix metalloproteinase-1
Rob Holland
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
UniProt ID: P11532 (718-1973) Human dystrophin central domain R4-15 fragment
|
|
|
|
| Sample: |
Human dystrophin central domain R4-15 fragment monomer, 150 kDa protein
|
| Buffer: |
NaP 20 mM, NaCl 300 mM, EDTA 1 mM, Glycérol 2%, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2016 May 26
|
Dystrophin SAXS data
Raphael Dos Santos Morais
|
| RgGuinier |
9.7 |
nm |
| Dmax |
47.5 |
nm |
|
|