|
|
|
|
|
| Sample: |
N-myc proto-oncogene protein, residues 1-100 monomer, 11 kDa Homo sapiens protein
Aurora kinase A mutant C290A:C393A monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20mM HEPES, 150mM NaCl, 5mM MgCl2, 3% v/v glycerol, 2mM TCEP, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Dec 9
|
The N-Myc MB0-MBI region interacts specifically and dynamically with the N-lobe of Aurora kinase A.
Nat Commun (2026)
Hultman J, Morad V, Tanner E, Kenney TMG, Pietras Z, Khare LP, Derbyshire D, Resetca D, Arrowsmith CH, Aili D, Ekström S, Penn LZ, Wallner B, Ahlner A, Sunnerhagen M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.4 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
|
|
|
|

|
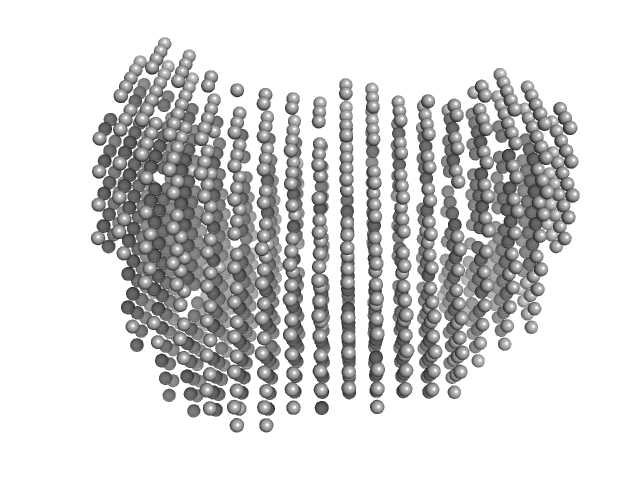
| Sample: |
Membrane primary amine oxidase dimer, 164 kDa Homo sapiens protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH: 7.4
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 Oct 16
|
Structural flexibility of domains in human AOC3
Gabriela Guedez
|
| RgGuinier |
4.6 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
372 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM EDTA, 3% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 4
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
|
|
|
|
|
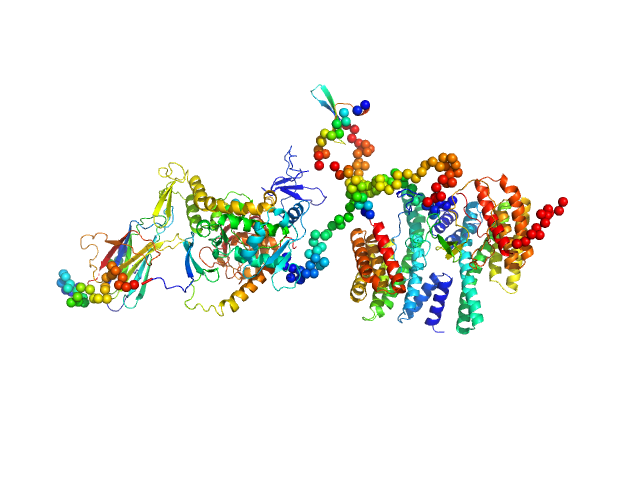
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM EDTA, 3% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Aug 17
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
5.4 |
nm |
| Dmax |
21.4 |
nm |
| VolumePorod |
281 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
14-3-3 protein eta dimer, 57 kDa Homo sapiens protein
Isoform 5 of E3 ubiquitin-protein ligase NEDD4-like monomer, 110 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 1 mM CaCl2, 3% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2023 Dec 4
|
Molecular basis of the regulation of full-length Nedd4-2 through calcium and 14-3-3 proteins
Veronika Obšilová
|
| RgGuinier |
5.5 |
nm |
| Dmax |
23.5 |
nm |
| VolumePorod |
290 |
nm3 |
|
|
|
|
|
|
|
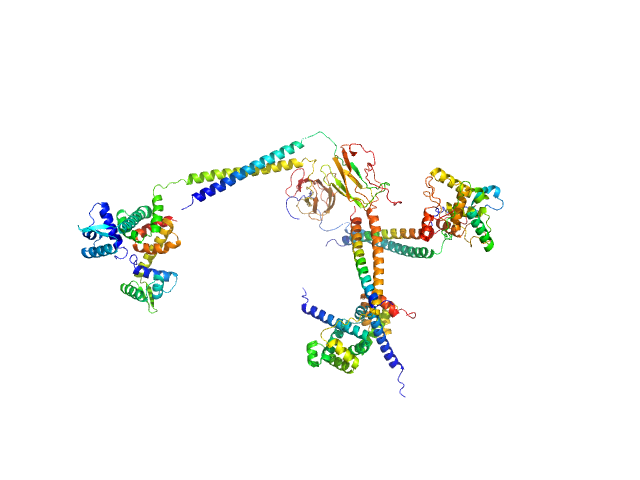
| Sample: |
Tumbleweed monomer, 133 kDa synthetic construct protein
|
| Buffer: |
10 mM HEPES, 150 mM NaCl, 5mM MgCl2, pH: 7.4
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Mar 15
|
An artificial motor protein that walks along a DNA track
Andrew Whitten
|
| RgGuinier |
6.7 |
nm |
| Dmax |
27.0 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Ras GTPase-activating protein-binding protein 1 dimer, 105 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl,, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Dec 13
|
SAXS reveals the molecular basis underlying pH-driven G3BP1 conformational dynamics: implications for stress granule formation
|
| RgGuinier |
6.6 |
nm |
| Dmax |
32.4 |
nm |
| VolumePorod |
221 |
nm3 |
|
|
|
|
|
|

|
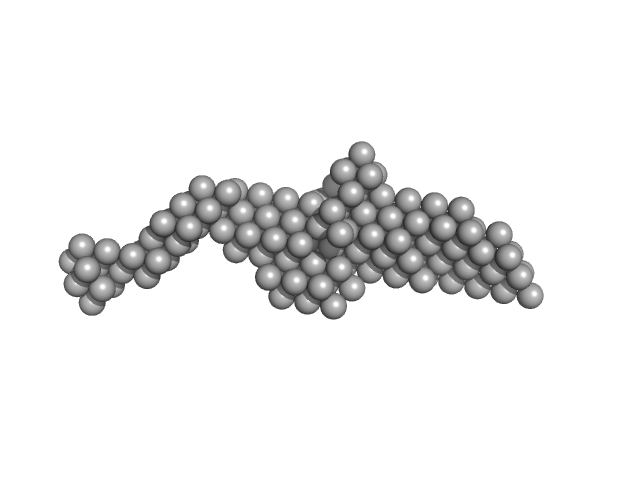
| Sample: |
Ras GTPase-activating protein-binding protein 1 dimer, 105 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl,, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 May 2
|
SAXS reveals the molecular basis underlying pH-driven G3BP1 conformational dynamics: implications for stress granule formation
|
| RgGuinier |
5.8 |
nm |
| Dmax |
28.7 |
nm |
| VolumePorod |
218 |
nm3 |
|
|
|
|
|
|

|
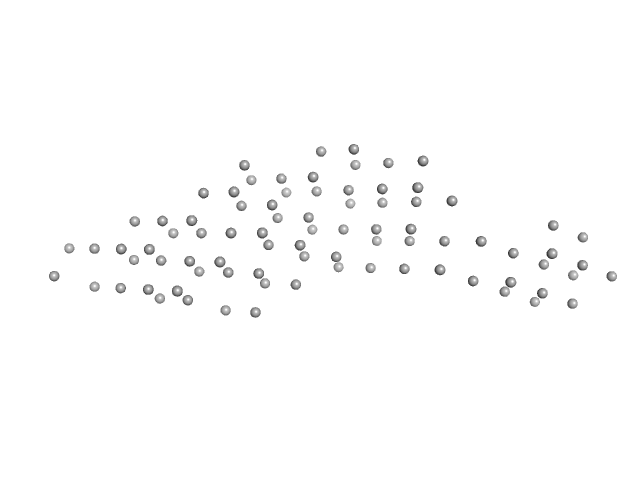
| Sample: |
Ras GTPase-activating protein-binding protein 1 dimer, 105 kDa Homo sapiens protein
|
| Buffer: |
50 mM MES, 50 mM NaCl,, pH: 6
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 May 2
|
SAXS reveals the molecular basis underlying pH-driven G3BP1 conformational dynamics: implications for stress granule formation
|
| RgGuinier |
4.9 |
nm |
| Dmax |
22.5 |
nm |
| VolumePorod |
163 |
nm3 |
|
|
|
|
|
|

|
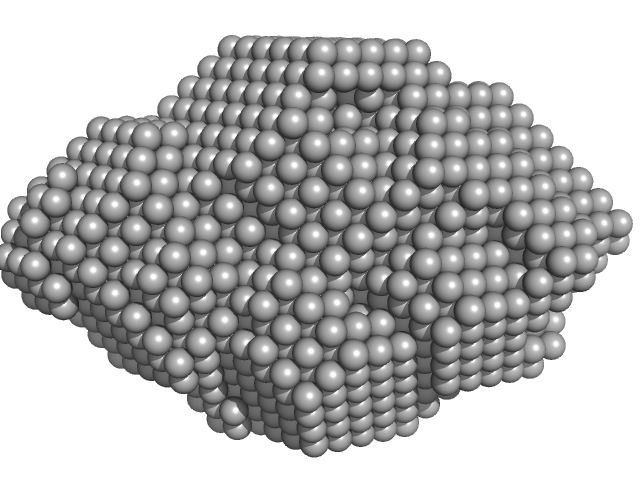
| Sample: |
NTF2L domain of Ras GTPase-activating protein-binding protein 1 dimer, 32 kDa protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl,, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 May 2
|
SAXS reveals the molecular basis underlying pH-driven G3BP1 conformational dynamics: implications for stress granule formation
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
40 |
nm3 |
|
|