|
|
|
|
|
| Sample: |
Ras GTPase-activating protein 1 monomer, 101 kDa Homo sapiens protein
Docking protein 1 monomer, 3 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 150 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 4
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.6 |
nm |
| VolumePorod |
196 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S) monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 350 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 12
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
42 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ephrin type-B receptor 4 monomer, 2 kDa Homo sapiens protein
Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S) monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 350 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 12
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Rho GTPase-activating protein 35 monomer, 3 kDa Homo sapiens protein
Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S) monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 350 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 12
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDQP8_ensemble_example.png)
|
| Sample: |
Stress response regulating small RNA OxyS monomer, 36 kDa Escherichia coli RNA
|
| Buffer: |
50 mM HEPES pH 6.8, 50 mM NaCl, pH: 6.8 |
| Experiment: |
SAXS
data collected at Austrian SAXS beamline 5.2L, ELETTRA on 2022 Jan 31
|
Spatial arrangement of functional domains in OxyS stress response sRNA
RNA :rna.079618.123 (2023)
Stih V, Amenitsch H, Plavec J, Podbevsek P
|
| RgGuinier |
4.8 |
nm |
| Dmax |
15.5 |
nm |
| VolumePorod |
68 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
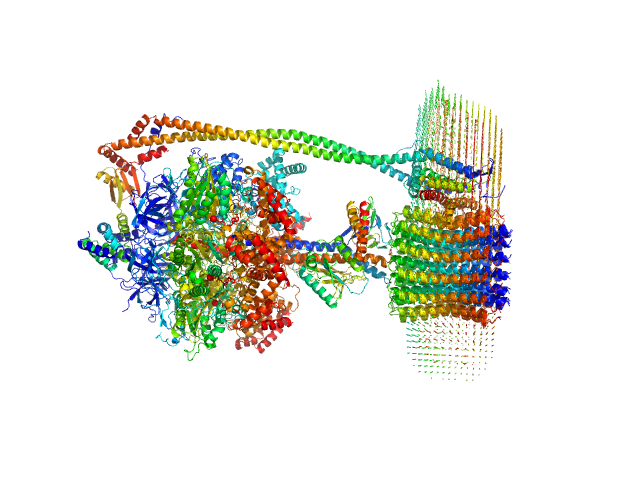
ATP synthase subunit alpha, chloroplastic trimer, 166 kDa Spinacia oleracea protein
ATP synthase subunit beta, chloroplastic trimer, 161 kDa Spinacia oleracea protein
ATP synthase gamma chain, chloroplastic monomer, 40 kDa Spinacia oleracea protein
ATP synthase delta chain, chloroplastic monomer, 28 kDa Spinacia oleracea protein
ATP synthase epsilon chain, chloroplastic monomer, 15 kDa Spinacia oleracea protein
ATP synthase subunit a, chloroplastic monomer, 27 kDa Spinacia oleracea protein
ATP synthase subunit b, chloroplastic monomer, 21 kDa Spinacia oleracea protein
ATP synthase subunit b', chloroplastic monomer, 24 kDa Spinacia oleracea protein
ATP synthase subunit c, chloroplastic 14-mer, 112 kDa Spinacia oleracea protein
4-trans-(4-trans-Propylcyclohexyl)-cyclohexyl α-maltoside 0, 283 kDa
|
| Buffer: |
300 mM NaCl, 30 mM HEPES, 2 mM MgCl2, 0.04% (w/v) tPCC-α-M, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2020 Oct 3
|
I-Shaped Dimers of a Plant Chloroplast FOF1-ATP Synthase in Response to Changes in Ionic Strength
International Journal of Molecular Sciences 24(13):10720 (2023)
Osipov S, Ryzhykau Y, Zinovev E, Minaeva A, Ivashchenko S, Verteletskiy D, Sudarev V, Kuklina D, Nikolaev M, Semenov Y, Zagryadskaya Y, Okhrimenko I, Gette M, Dronova E, Shishkin A, Dencher N, Kuklin A, Ivanovich V, Uversky V, Vlasov A
|
| RgGuinier |
6.6 |
nm |
| Dmax |
27.5 |
nm |
| VolumePorod |
927 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDRS8_fit1_model1.png)
|
| Sample: |
ATP synthase subunit alpha, chloroplastic trimer, 166 kDa Spinacia oleracea protein
ATP synthase subunit beta, chloroplastic trimer, 161 kDa Spinacia oleracea protein
ATP synthase gamma chain, chloroplastic monomer, 40 kDa Spinacia oleracea protein
ATP synthase delta chain, chloroplastic monomer, 28 kDa Spinacia oleracea protein
ATP synthase epsilon chain, chloroplastic monomer, 15 kDa Spinacia oleracea protein
ATP synthase subunit a, chloroplastic monomer, 27 kDa Spinacia oleracea protein
ATP synthase subunit b, chloroplastic monomer, 21 kDa Spinacia oleracea protein
ATP synthase subunit b', chloroplastic monomer, 24 kDa Spinacia oleracea protein
ATP synthase subunit c, chloroplastic 14-mer, 112 kDa Spinacia oleracea protein
4-trans-(4-trans-Propylcyclohexyl)-cyclohexyl α-maltoside 0, 283 kDa
|
| Buffer: |
150 mM NaCl, 30 mM HEPES, 2 mM MgCl2, 0.04% (w/v) tPCC-α-M, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2020 Oct 3
|
I-Shaped Dimers of a Plant Chloroplast FOF1-ATP Synthase in Response to Changes in Ionic Strength
International Journal of Molecular Sciences 24(13):10720 (2023)
Osipov S, Ryzhykau Y, Zinovev E, Minaeva A, Ivashchenko S, Verteletskiy D, Sudarev V, Kuklina D, Nikolaev M, Semenov Y, Zagryadskaya Y, Okhrimenko I, Gette M, Dronova E, Shishkin A, Dencher N, Kuklin A, Ivanovich V, Uversky V, Vlasov A
|
| RgGuinier |
9.6 |
nm |
| Dmax |
41.5 |
nm |
| VolumePorod |
1506 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDRT8_fit1_model1.png)
|
| Sample: |
ATP synthase subunit alpha, chloroplastic trimer, 166 kDa Spinacia oleracea protein
ATP synthase subunit beta, chloroplastic trimer, 161 kDa Spinacia oleracea protein
ATP synthase gamma chain, chloroplastic monomer, 40 kDa Spinacia oleracea protein
ATP synthase delta chain, chloroplastic monomer, 28 kDa Spinacia oleracea protein
ATP synthase epsilon chain, chloroplastic monomer, 15 kDa Spinacia oleracea protein
ATP synthase subunit a, chloroplastic monomer, 27 kDa Spinacia oleracea protein
ATP synthase subunit b, chloroplastic monomer, 21 kDa Spinacia oleracea protein
ATP synthase subunit b', chloroplastic monomer, 24 kDa Spinacia oleracea protein
ATP synthase subunit c, chloroplastic 14-mer, 112 kDa Spinacia oleracea protein
4-trans-(4-trans-Propylcyclohexyl)-cyclohexyl α-maltoside 0, 283 kDa
|
| Buffer: |
250 mM NaCl, 30 mM HEPES, 2 mM MgCl2, 0.04% (w/v) tPCC-α-M, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2020 Oct 3
|
I-Shaped Dimers of a Plant Chloroplast FOF1-ATP Synthase in Response to Changes in Ionic Strength
International Journal of Molecular Sciences 24(13):10720 (2023)
Osipov S, Ryzhykau Y, Zinovev E, Minaeva A, Ivashchenko S, Verteletskiy D, Sudarev V, Kuklina D, Nikolaev M, Semenov Y, Zagryadskaya Y, Okhrimenko I, Gette M, Dronova E, Shishkin A, Dencher N, Kuklin A, Ivanovich V, Uversky V, Vlasov A
|
| RgGuinier |
7.4 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
949 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDRU8_fit1_model1.png)
|
| Sample: |
ATP synthase subunit alpha, chloroplastic trimer, 166 kDa Spinacia oleracea protein
ATP synthase subunit beta, chloroplastic trimer, 161 kDa Spinacia oleracea protein
ATP synthase gamma chain, chloroplastic monomer, 40 kDa Spinacia oleracea protein
ATP synthase delta chain, chloroplastic monomer, 28 kDa Spinacia oleracea protein
ATP synthase epsilon chain, chloroplastic monomer, 15 kDa Spinacia oleracea protein
ATP synthase subunit a, chloroplastic monomer, 27 kDa Spinacia oleracea protein
ATP synthase subunit b, chloroplastic monomer, 21 kDa Spinacia oleracea protein
ATP synthase subunit b', chloroplastic monomer, 24 kDa Spinacia oleracea protein
ATP synthase subunit c, chloroplastic 14-mer, 112 kDa Spinacia oleracea protein
4-trans-(4-trans-Propylcyclohexyl)-cyclohexyl α-maltoside 0, 283 kDa
|
| Buffer: |
300 mM NaCl, 30 mM HEPES, 2 mM MgCl2, 0.04% (w/v) tPCC-α-M, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2020 Oct 3
|
I-Shaped Dimers of a Plant Chloroplast FOF1-ATP Synthase in Response to Changes in Ionic Strength
International Journal of Molecular Sciences 24(13):10720 (2023)
Osipov S, Ryzhykau Y, Zinovev E, Minaeva A, Ivashchenko S, Verteletskiy D, Sudarev V, Kuklina D, Nikolaev M, Semenov Y, Zagryadskaya Y, Okhrimenko I, Gette M, Dronova E, Shishkin A, Dencher N, Kuklin A, Ivanovich V, Uversky V, Vlasov A
|
| RgGuinier |
7.8 |
nm |
| Dmax |
39.5 |
nm |
| VolumePorod |
1127 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDRV8_fit1_model1.png)
|
| Sample: |
ATP synthase subunit alpha, chloroplastic trimer, 166 kDa Spinacia oleracea protein
ATP synthase subunit beta, chloroplastic trimer, 161 kDa Spinacia oleracea protein
ATP synthase gamma chain, chloroplastic monomer, 40 kDa Spinacia oleracea protein
ATP synthase delta chain, chloroplastic monomer, 28 kDa Spinacia oleracea protein
ATP synthase epsilon chain, chloroplastic monomer, 15 kDa Spinacia oleracea protein
ATP synthase subunit a, chloroplastic monomer, 27 kDa Spinacia oleracea protein
ATP synthase subunit b, chloroplastic monomer, 21 kDa Spinacia oleracea protein
ATP synthase subunit b', chloroplastic monomer, 24 kDa Spinacia oleracea protein
ATP synthase subunit c, chloroplastic 14-mer, 112 kDa Spinacia oleracea protein
4-trans-(4-trans-Propylcyclohexyl)-cyclohexyl α-maltoside 0, 283 kDa
|
| Buffer: |
350 mM NaCl, 30 mM HEPES, 2 mM MgCl2, 0.04% (w/v) tPCC-α-M, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2020 Oct 3
|
I-Shaped Dimers of a Plant Chloroplast FOF1-ATP Synthase in Response to Changes in Ionic Strength
International Journal of Molecular Sciences 24(13):10720 (2023)
Osipov S, Ryzhykau Y, Zinovev E, Minaeva A, Ivashchenko S, Verteletskiy D, Sudarev V, Kuklina D, Nikolaev M, Semenov Y, Zagryadskaya Y, Okhrimenko I, Gette M, Dronova E, Shishkin A, Dencher N, Kuklin A, Ivanovich V, Uversky V, Vlasov A
|
| RgGuinier |
8.9 |
nm |
| Dmax |
44.5 |
nm |
| VolumePorod |
1150 |
nm3 |
|
|