|
|
|
|
|
| Sample: |
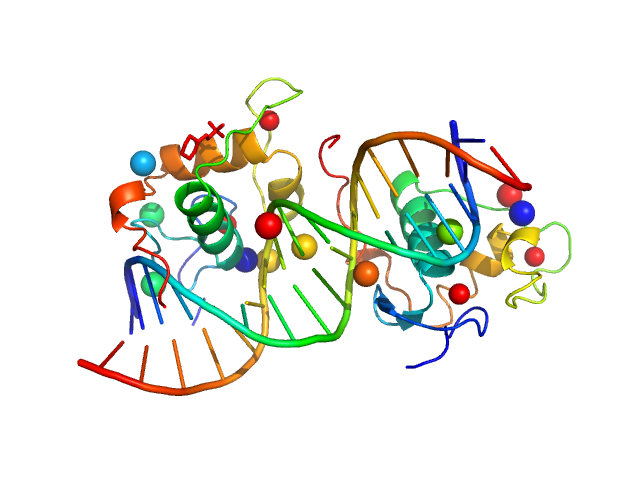
Ramp2 DNA monomer, 11 kDa DNA
Retinoic acid receptor RXR-alpha dimer, 20 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 50 mM KCl, 5% glycerol, 2 mM Chaps, and 5 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Oct 7
|
Solution Behavior of the Intrinsically Disordered N-Terminal Domain of Retinoid X Receptor α in the Context of the Full-Length Protein
Biochemistry 55(12):1741-1748 (2016)
Belorusova A, Osz J, Petoukhov M, Peluso-Iltis C, Kieffer B, Svergun D, Rochel N
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.8 |
nm |
|
|
|
|
|
|

|
| Sample: |
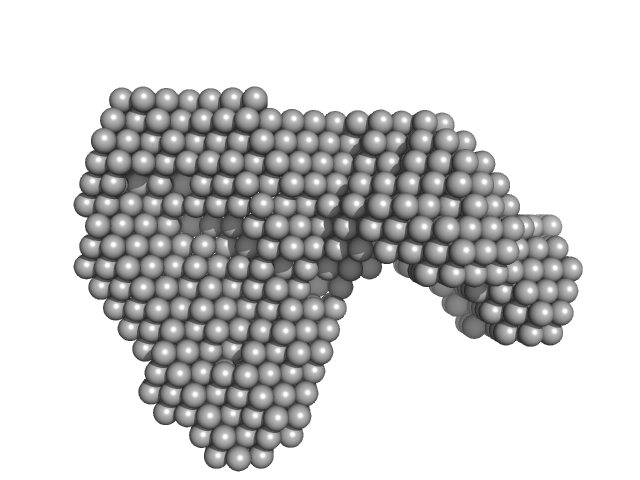
Contactin-associated protein-like 2 extracellular domains (1-1261) monomer, 140 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSess, University of Utah on 2010 Oct 4
|
Structural Characterization of the Extracellular Domain of CASPR2 and Insights into Its Association with the Novel Ligand Contactin1.
J Biol Chem 291(11):5788-802 (2016)
Rubio-Marrero EN, Vincelli G, Jeffries CM, Shaikh TR, Pakos IS, Ranaivoson FM, von Daake S, Demeler B, De Jaco A, Perkins G, Ellisman MH, Trewhella J, Comoletti D
|
| RgGuinier |
4.4 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
282 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
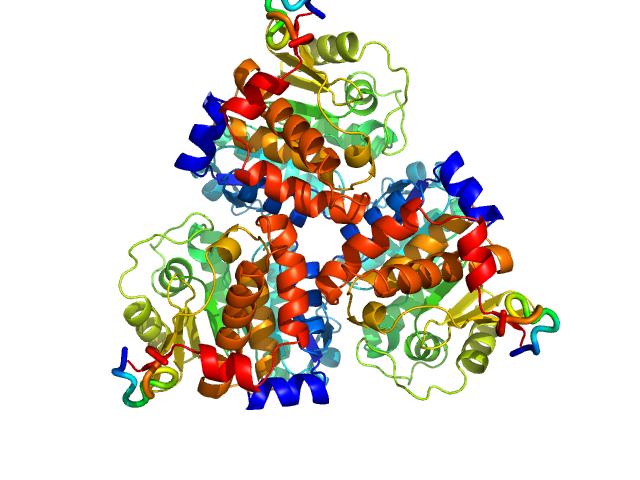
Ribokinase ThiM trimer, 89 kDa Staphylococcus aureus protein
|
| Buffer: |
50 mM Potassium phosphate 10 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2010 Nov 19
|
Structure of ThiM from Vitamin B1 biosynthetic pathway of Staphylococcus aureus - Insights into a novel pro-drug approach addressing MRSA infections.
Sci Rep 6:22871 (2016)
Drebes J, Künz M, Windshügel B, Kikhney AG, Müller IB, Eberle RJ, Oberthür D, Cang H, Svergun DI, Perbandt M, Betzel C, Wrenger C
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
145 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
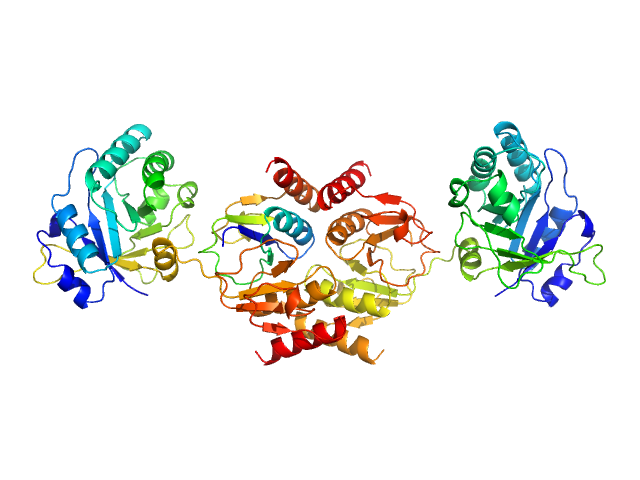
Endolysin , 33 kDa Clostridium phage phiCTP1 protein
|
| Buffer: |
20 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Mar 17
|
Crystal Structure of the CTP1L Endolysin Reveals How Its Activity Is Regulated by a Secondary Translation Product.
J Biol Chem 291(10):4882-93 (2016)
Dunne M, Leicht S, Krichel B, Mertens HD, Thompson A, Krijgsveld J, Svergun DI, Gómez-Torres N, Garde S, Uetrecht C, Narbad A, Mayer MJ, Meijers R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.8 |
nm |
| VolumePorod |
95 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
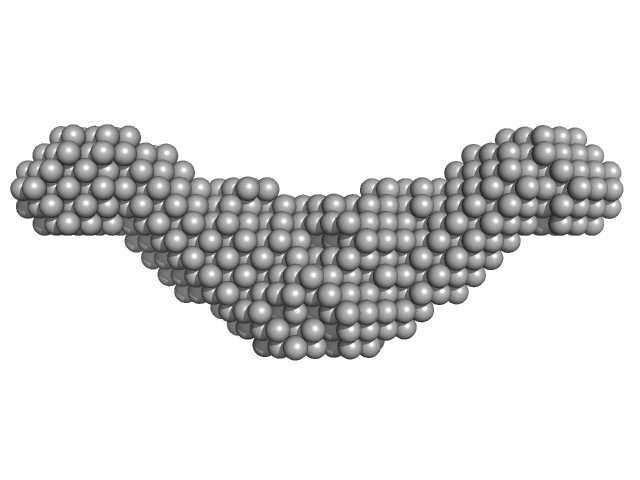
Endolysin CS74L , 31 kDa Clostridium phage phi8074-B1 protein
|
| Buffer: |
20 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Mar 17
|
Crystal Structure of the CTP1L Endolysin Reveals How Its Activity Is Regulated by a Secondary Translation Product.
J Biol Chem 291(10):4882-93 (2016)
Dunne M, Leicht S, Krichel B, Mertens HD, Thompson A, Krijgsveld J, Svergun DI, Gómez-Torres N, Garde S, Uetrecht C, Narbad A, Mayer MJ, Meijers R
|
| RgGuinier |
3.6 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
79 |
nm3 |
|
|
|
|
|
|
|
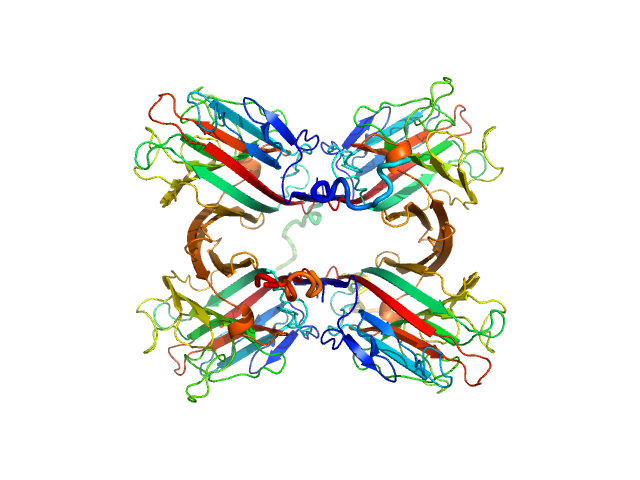
| Sample: |
Recombinant Tn antigen-binding lectin tetramer, 105 kDa Vatairea macrocarpa protein
|
| Buffer: |
100 mM sodium phosphate 150 mM NaCl 5% (v/v) glycerol, pH: 5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2014 Jan 11
|
Structural characterization of a Vatairea macrocarpa lectin in complex with a tumor-associated antigen: A new tool for cancer research.
Int J Biochem Cell Biol 72:27-39 (2016)
Sousa BL, Silva-Filho JC, Kumar P, Graewert MA, Pereira RI, Cunha RMS, Nascimento KS, Bezerra GA, Delatorre P, Djinovic-Carugo K, Nagano CS, Gruber K, Cavada BS
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
168 |
nm3 |
|
|
|
|
|
|
|
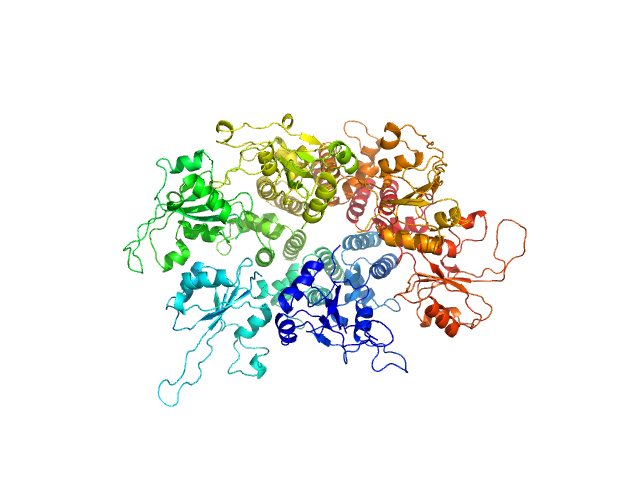
| Sample: |
Replication initiator protein of a promiscuous streptococcal plasmid pMV158. hexamer, Streptococcus sp. protein
|
| Buffer: |
10 mM TRIS 5 mM EDTA 1.0 M KCl, pH: 8.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Jun 16
|
Conformational plasticity of RepB, the replication initiator protein of promiscuous streptococcal plasmid pMV158.
Sci Rep 6:20915 (2016)
Boer DR, Ruiz-Masó JA, Rueda M, Petoukhov MV, Machón C, Svergun DI, Orozco M, del Solar G, Coll M
|
| RgGuinier |
3.8 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
282 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
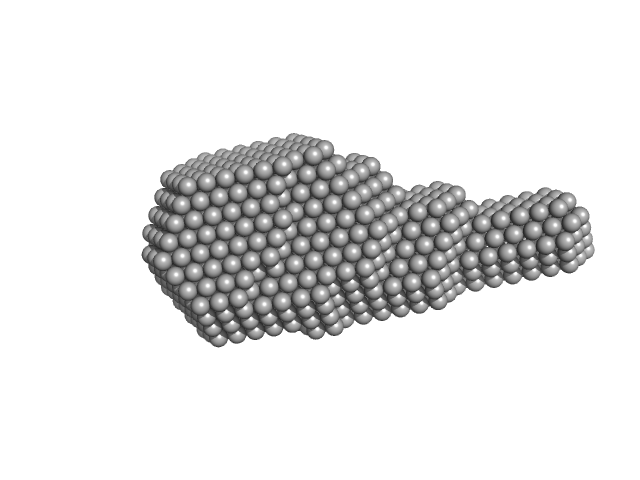
Middle East Respiratory Syndrome (MERS) coronavirus nucleocapsid N-Protein (N-terminal domain 1-164) monomer, 18 kDa Middle East respiratory … protein
|
| Buffer: |
10 mM HEPES 300 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Oct 5
|
Structural characterization of the N-terminal part of the MERS-CoV nucleocapsid by X-ray diffraction and small-angle X-ray scattering.
Acta Crystallogr D Struct Biol 72(Pt 2):192-202 (2016)
Papageorgiou N, Lichière J, Baklouti A, Ferron F, Sévajol M, Canard B, Coutard B
|
| RgGuinier |
2.0 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
|
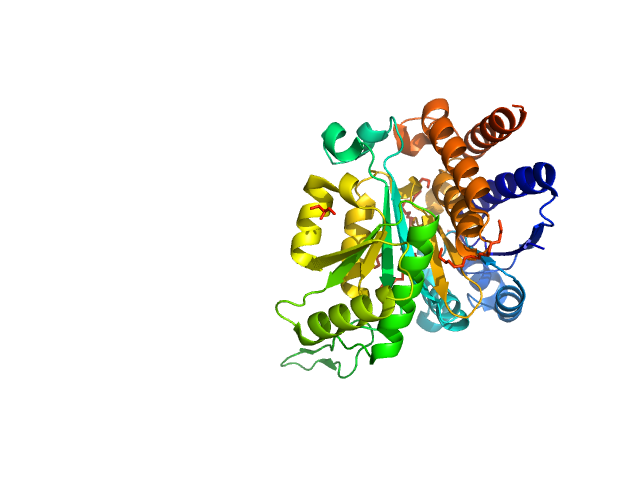
| Sample: |
3-isopropylmalate dehydrogenase dimer, 73 kDa Thermus thermophilus protein
|
| Buffer: |
25 mM MOPS/NaOH, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2012 Nov 16
|
Dual Role of the Active Site Residues of Thermus thermophilus 3-Isopropylmalate Dehydrogenase: Chemical Catalysis and Domain Closure.
Biochemistry 55(3):560-74 (2016)
Gráczer É, Szimler T, Garamszegi A, Konarev PV, Lábas A, Oláh J, Palló A, Svergun DI, Merli A, Závodszky P, Weiss MS, Vas M
|
|
|
|
|
|
|
|
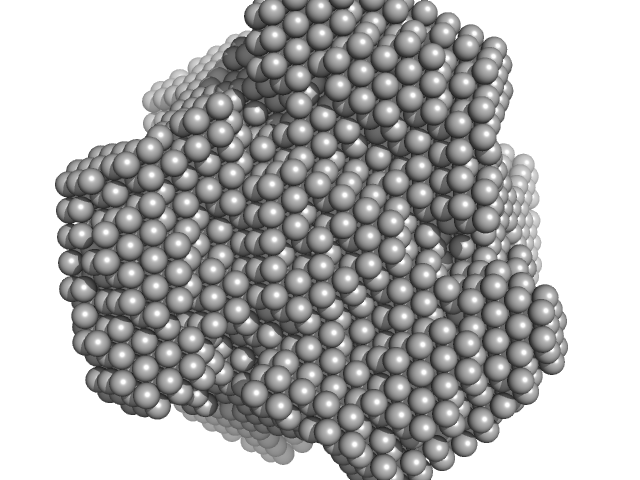
| Sample: |
Inorganic pyrophosphatase (PPase) from E. coli hexamer, 117 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris 10 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Jun 30
|
X-Ray Solution Scattering Study of Four Escherichia coli Enzymes Involved in Stationary-Phase Metabolism.
PLoS One 11(5):e0156105 (2016)
Dadinova LA, Shtykova EV, Konarev PV, Rodina EV, Snalina NE, Vorobyeva NN, Kurilova SA, Nazarova TI, Jeffries CM, Svergun DI
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
166 |
nm3 |
|
|