|
|
|
|
|
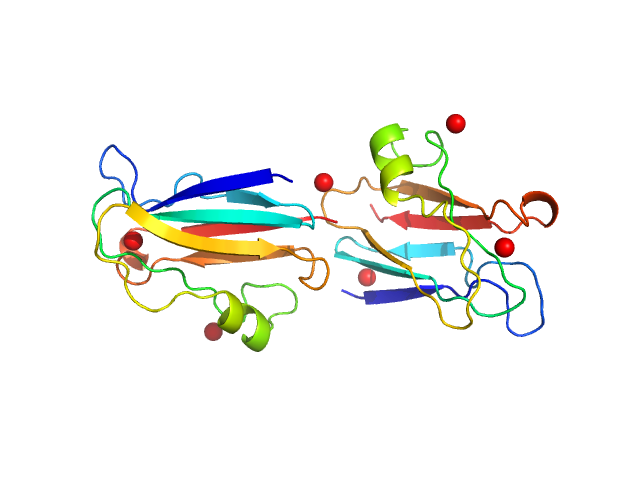
| Sample: |
Plastocyanin dimer, 23 kDa Phormidium laminosum protein
|
| Buffer: |
pure water, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2009 May 7
|
Metal-Mediated Self-Assembly of a β-Sandwich Protein
Chemistry - A European Journal 15(46):12672-12680 (2009)
Crowley P, Matias P, Khan A, Roessle M, Svergun D
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
|
|
|
|
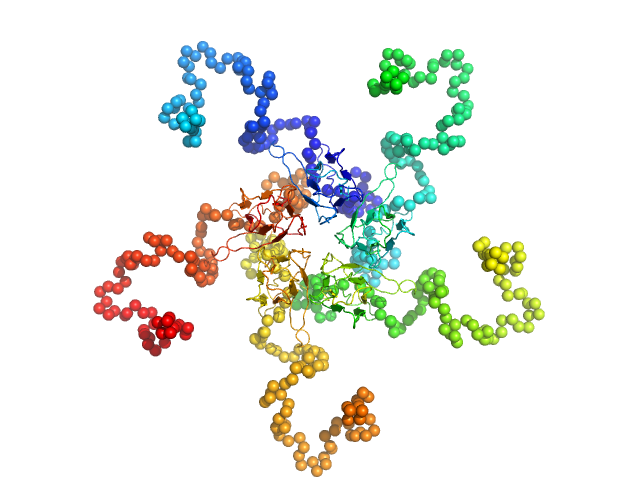
|
| Sample: |
Nucleoplasmin pentamer, 110 kDa Escherichia coli protein
|
| Buffer: |
20 mM Pipes buffer 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Dec 3
|
A mechanism for histone chaperoning activity of nucleoplasmin: thermodynamic and structural models.
J Mol Biol 393(2):448-63 (2009)
Taneva SG, Bañuelos S, Falces J, Arregi I, Muga A, Konarev PV, Svergun DI, Velázquez-Campoy A, Urbaneja MA
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
210 |
nm3 |
|
|
|
|
|
|

|
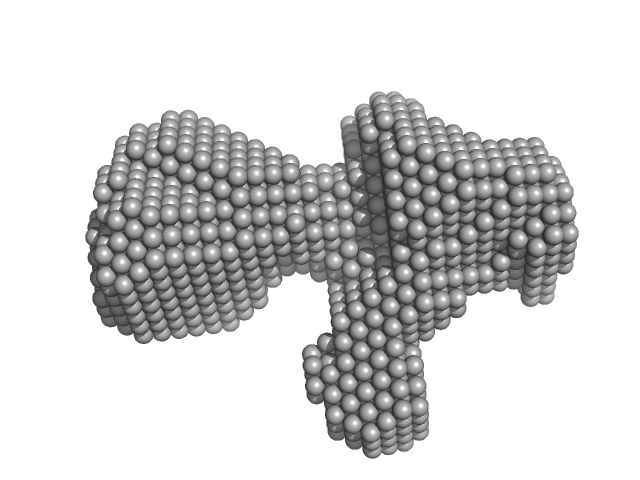
| Sample: |
DH-PH module of PDZRhoGEF monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCL 150 mM NaCl 1.0 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2008 Oct 30
|
The solution structure and dynamics of the DH-PH module of PDZRhoGEF in isolation and in complex with nucleotide-free RhoA.
Protein Sci 18(10):2067-79 (2009)
Cierpicki T, Bielnicki J, Zheng M, Gruszczyk J, Kasterka M, Petoukhov M, Zhang A, Fernandez EJ, Svergun DI, Derewenda U, Bushweller JH, Derewenda ZS
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
|
|
|
|
|
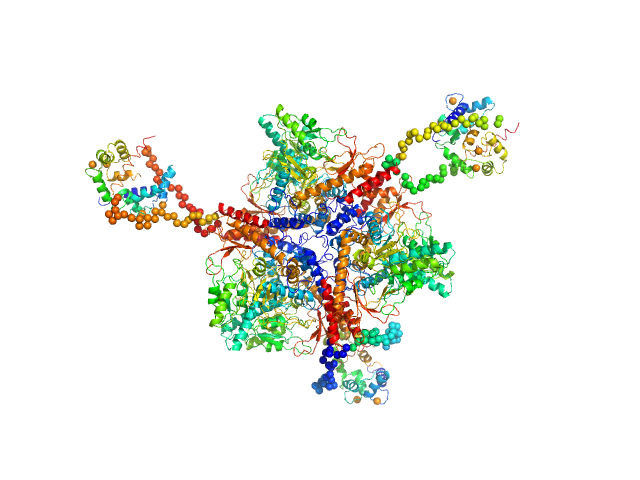
| Sample: |
Calmodulin monomer, 17 kDa Homo sapiens protein
Glutamate decarboxylase hexamer, 340 kDa Petunia x hybrida protein
|
| Buffer: |
50 mM HEPES 50 mM KCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2004 Nov 9
|
A common structural basis for pH- and calmodulin-mediated regulation in plant glutamate decarboxylase.
J Mol Biol 392(2):334-51 (2009)
Gut H, Dominici P, Pilati S, Astegno A, Petoukhov MV, Svergun DI, Grütter MG, Capitani G
|
| RgGuinier |
5.9 |
nm |
| Dmax |
23.0 |
nm |
| VolumePorod |
630 |
nm3 |
|
|
|
|
|
|

|
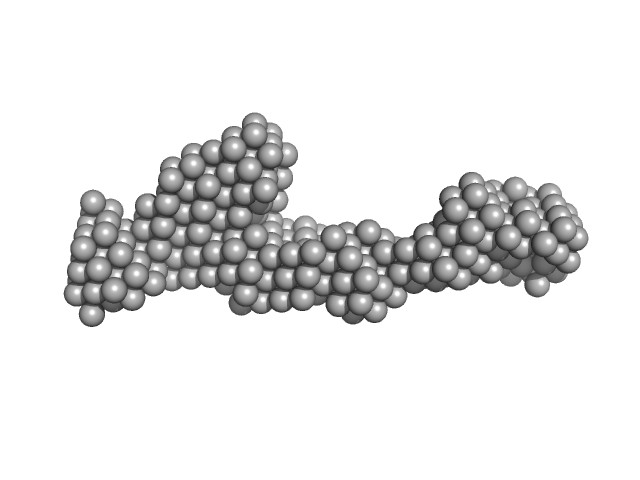
| Sample: |
Peroxisomal targeting signal 1 receptor monomer, 71 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES-KOH (pH 7.5), 100mM KCl, and 20mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 Feb 24
|
Solution structure of human Pex5.Pex14.PTS1 protein complexes obtained by small angle X-ray scattering.
J Biol Chem 284(37):25334-42 (2009)
Shiozawa K, Konarev PV, Neufeld C, Wilmanns M, Svergun DI
|
| RgGuinier |
5.0 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
181 |
nm3 |
|
|
|
|
|
|
|
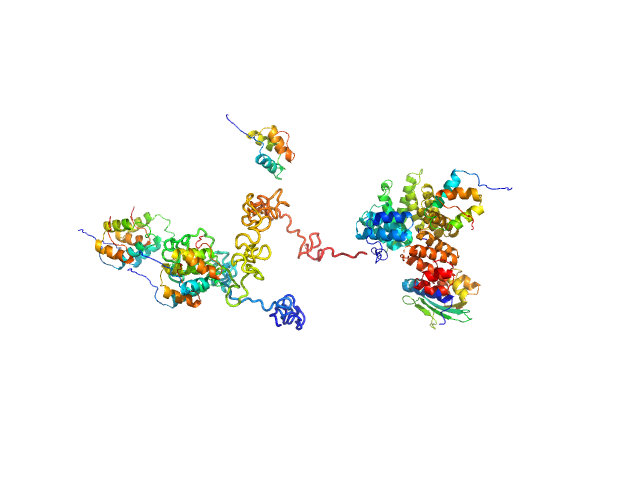
| Sample: |
Peroxisomal targeting signal 1 receptor monomer, 71 kDa Homo sapiens protein
Peroxisomal membrane protein PEX14 monomer, 7 kDa Homo sapiens protein
PTS1-BP monomer, 13 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES-KOH (pH 7.5), 100mM KCl, and 20mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 Mar 18
|
Solution structure of human Pex5.Pex14.PTS1 protein complexes obtained by small angle X-ray scattering.
J Biol Chem 284(37):25334-42 (2009)
Shiozawa K, Konarev PV, Neufeld C, Wilmanns M, Svergun DI
|
| RgGuinier |
6.0 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
267 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
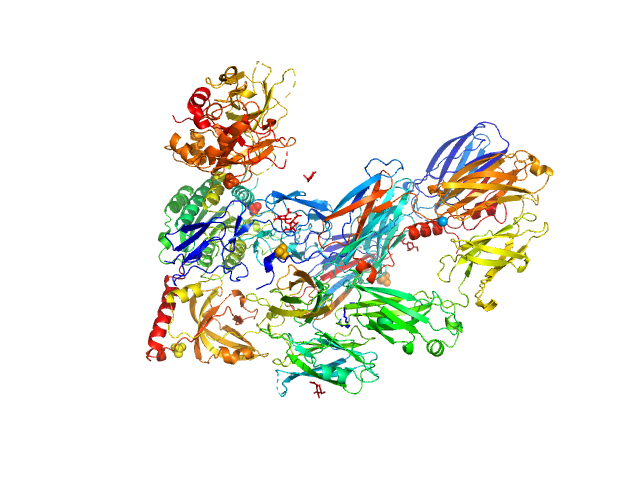
Cobra venom factor monomer, 185 kDa Naja kaouthia protein
|
| Buffer: |
10 mM Tris 5 mM MgCl2 10 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Dec 19
|
Insights into complement convertase formation based on the structure of the factor B-cobra venom factor complex
The EMBO Journal 28(16):2469-2478 (2009)
Janssen B, Gomes L, Koning R, Svergun D, Koster A, Fritzinger D, Vogel C, Gros P
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
384 |
nm3 |
|
|
|
|
|
|

|
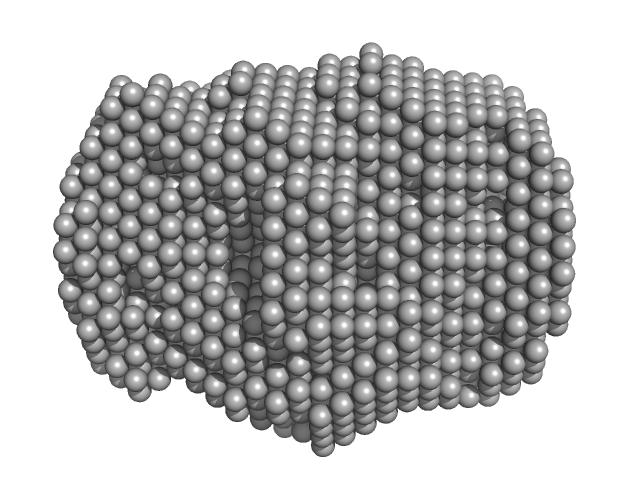
| Sample: |
ProNGF dimer, 45 kDa Mouse submandibulary glands protein
|
| Buffer: |
50 mM Naphosphat 0.5 M Ammonium Sulfate(NH4)2SO4, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Oct 25
|
Intrinsic structural disorder of mouse proNGF.
Proteins 75(4):990-1009 (2009)
Paoletti F, Covaceuszach S, Konarev PV, Gonfloni S, Malerba F, Schwarz E, Svergun DI, Cattaneo A, Lamba D
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Endoglin tetramer, 243 kDa Homo sapiens protein
|
| Buffer: |
10 mM Tris–HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2008 Oct 17
|
Structural and functional characterization of soluble endoglin receptor
Biochemical and Biophysical Research Communications 383(4):386-391 (2009)
Le B, Franke D, Svergun D, Han T, Hwang H, Kim K
|
| RgGuinier |
7.6 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
434 |
nm3 |
|
|
|
|
|
|

|
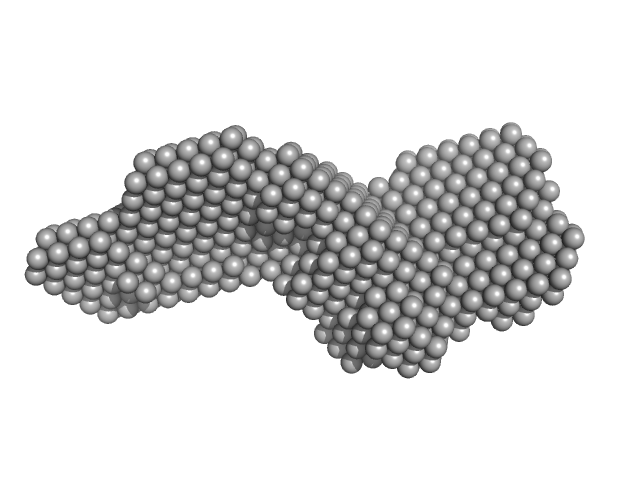
| Sample: |
Endoglin dimer, 121 kDa Homo sapiens protein
|
| Buffer: |
10 mM Tris–HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2008 Oct 17
|
Structural and functional characterization of soluble endoglin receptor
Biochemical and Biophysical Research Communications 383(4):386-391 (2009)
Le B, Franke D, Svergun D, Han T, Hwang H, Kim K
|
| RgGuinier |
4.7 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
173 |
nm3 |
|
|