|
|
|
|
|
| Sample: |
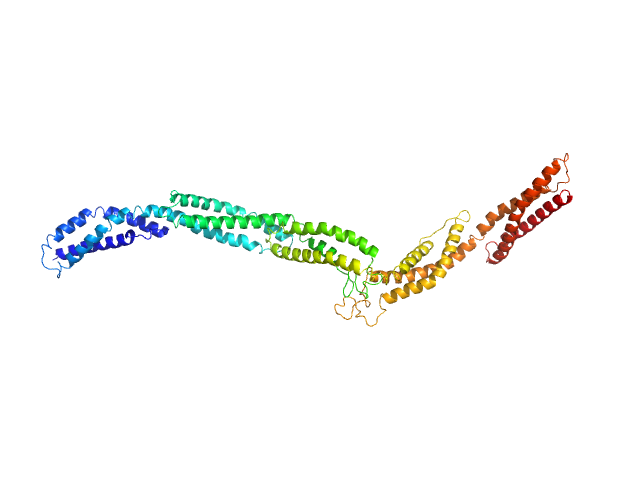
Dystrophin central domain repeats 20 to 24. monomer, 67 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 1 mM EDTA 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2011 Jul 10
|
Dystrophin's central domain forms a complex filament that becomes disorganized by in-frame deletions.
J Biol Chem 293(18):6637-6646 (2018)
Delalande O, Molza AE, Dos Santos Morais R, Chéron A, Pollet É, Raguenes-Nicol C, Tascon C, Giudice E, Guilbaud M, Nicolas A, Bondon A, Leturcq F, Férey N, Baaden M, Perez J, Roblin P, Piétri-Rouxel F, Hubert JF, Czjzek M, Le Rumeur E
|
| RgGuinier |
5.8 |
nm |
| Dmax |
22.5 |
nm |
| VolumePorod |
107 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
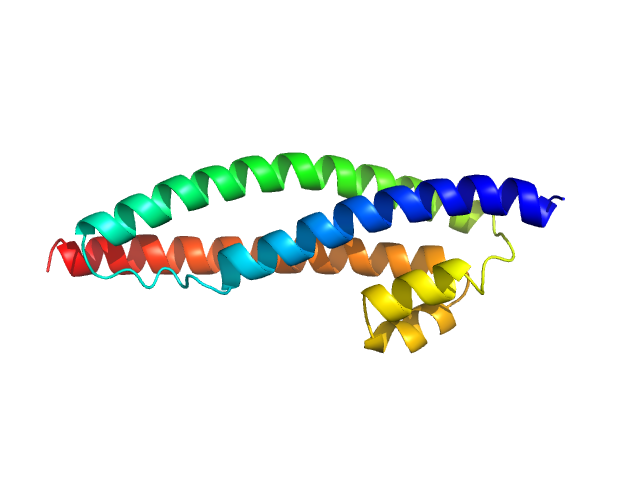
Dystrophin central domain single repeat 23 monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 1 mM EDTA 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2011 Oct 7
|
Dystrophin's central domain forms a complex filament that becomes disorganized by in-frame deletions.
J Biol Chem 293(18):6637-6646 (2018)
Delalande O, Molza AE, Dos Santos Morais R, Chéron A, Pollet É, Raguenes-Nicol C, Tascon C, Giudice E, Guilbaud M, Nicolas A, Bondon A, Leturcq F, Férey N, Baaden M, Perez J, Roblin P, Piétri-Rouxel F, Hubert JF, Czjzek M, Le Rumeur E
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
20 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
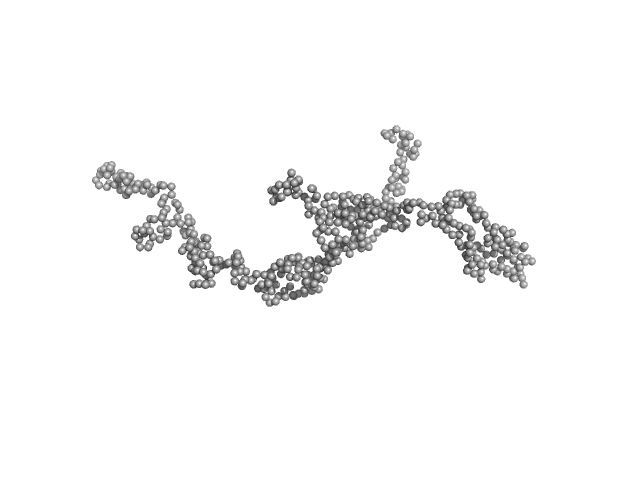
Dystrophin central domain repeats 16 to 21 (Δ2146-2305; Becker muscular dystrophy variant, deletion of exons 45-47) monomer, 64 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 1 mM EDTA 2% glycerol 5% acetonitrile, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2014 Feb 5
|
Dystrophin's central domain forms a complex filament that becomes disorganized by in-frame deletions.
J Biol Chem 293(18):6637-6646 (2018)
Delalande O, Molza AE, Dos Santos Morais R, Chéron A, Pollet É, Raguenes-Nicol C, Tascon C, Giudice E, Guilbaud M, Nicolas A, Bondon A, Leturcq F, Férey N, Baaden M, Perez J, Roblin P, Piétri-Rouxel F, Hubert JF, Czjzek M, Le Rumeur E
|
| RgGuinier |
6.0 |
nm |
| Dmax |
21.0 |
nm |
| VolumePorod |
184 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
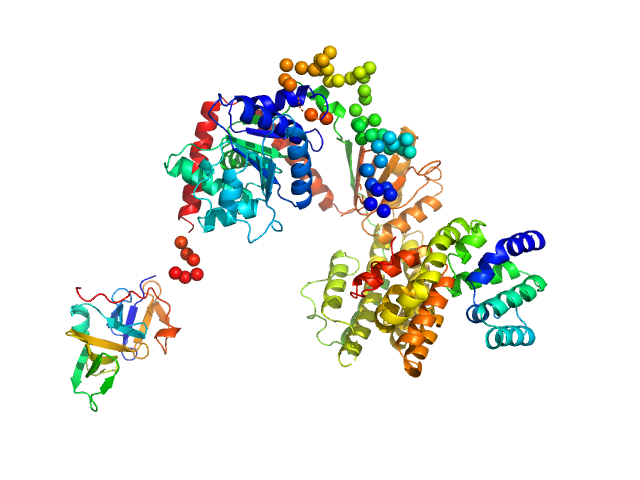
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2012 Apr 6
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
186 |
nm3 |
|
|
|
|
|
|
|
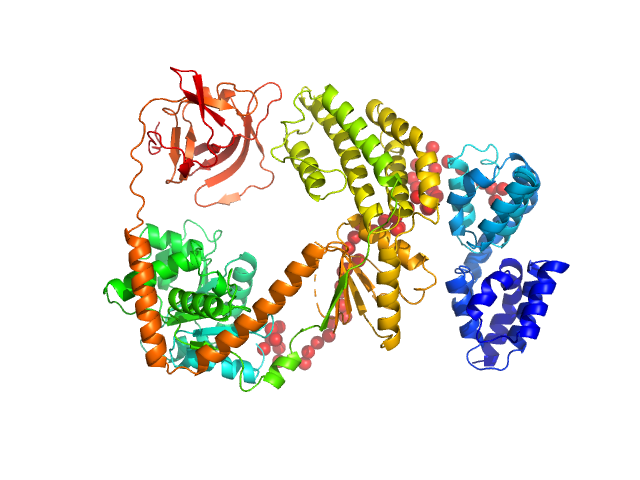
| Sample: |
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, 2mM ADP-AlFx, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 20
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.2 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
|
|
|
|
|
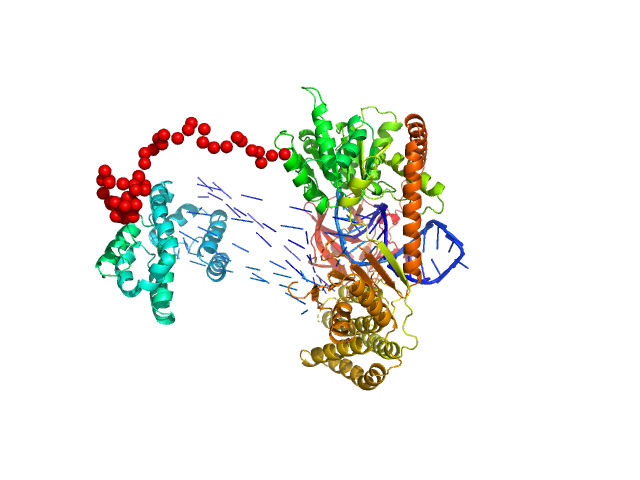
| Sample: |
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
5´ppp 10mer hairpin dsRNA monomer, 8 kDa RNA
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 May 29
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.1 |
nm |
| Dmax |
16.1 |
nm |
| VolumePorod |
160 |
nm3 |
|
|
|
|
|
|
|
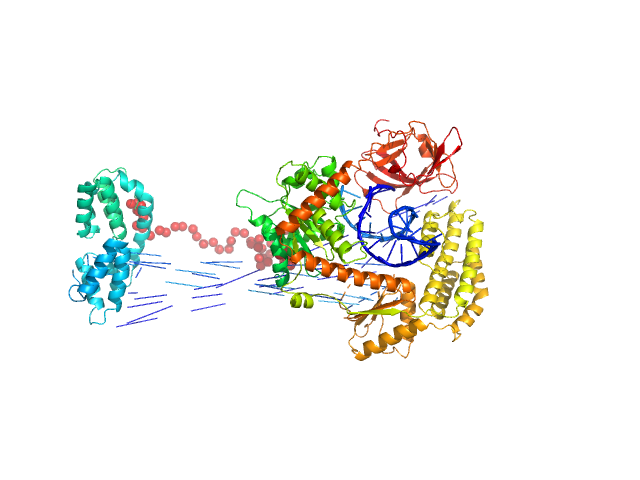
| Sample: |
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
5´ppp 10mer hairpin dsRNA monomer, 8 kDa RNA
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, 0.5 mM AMP-PNP, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 May 28
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.2 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
163 |
nm3 |
|
|
|
|
|
|
|
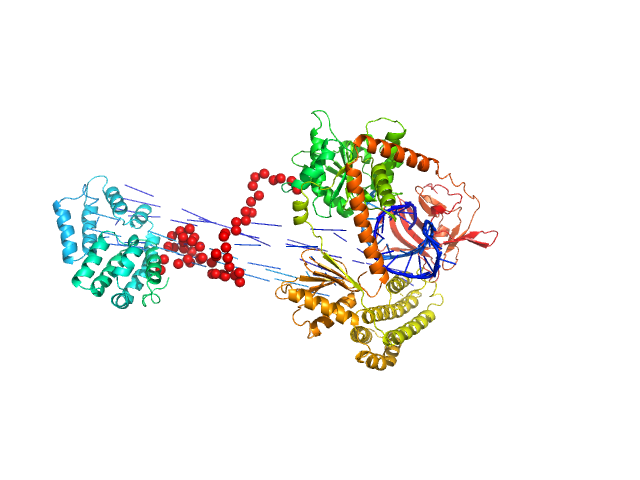
| Sample: |
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
5´ppp 10mer hairpin dsRNA monomer, 8 kDa RNA
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, 2mM ADP-AlFx, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 20
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.0 |
nm |
| Dmax |
18.3 |
nm |
| VolumePorod |
156 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
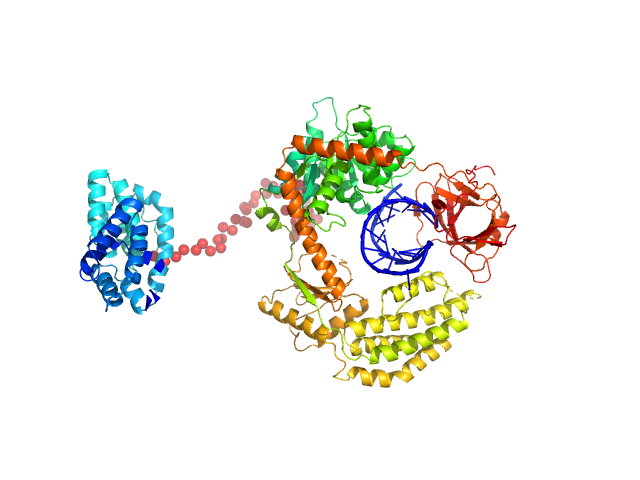
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
5´ppp 8mer hairpin dsRNA monomer, 6 kDa RNA
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2015 Nov 20
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.3 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
179 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
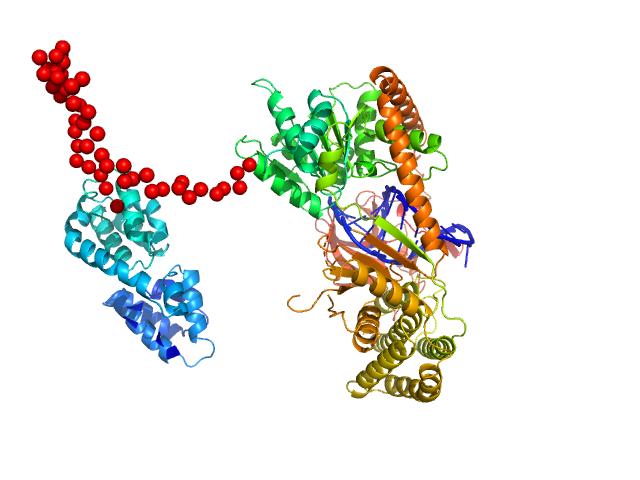
Probable ATP-dependent RNA helicase DDX58 monomer, 108 kDa Homo sapiens protein
5´ppp 8mer hairpin dsRNA monomer, 6 kDa RNA
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 2.5 mM MgCl2, 10% glycerol and 1mM DTT, 0.5 mM AMP-PNP, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 Apr 27
|
Combined roles of ATP and small hairpin RNA in the activation of RIG-I revealed by solution-based analysis.
Nucleic Acids Res 46(6):3169-3186 (2018)
Shah N, Beckham SA, Wilce JA, Wilce MCJ
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
188 |
nm3 |
|
|