|
|
|
|
|
| Sample: |
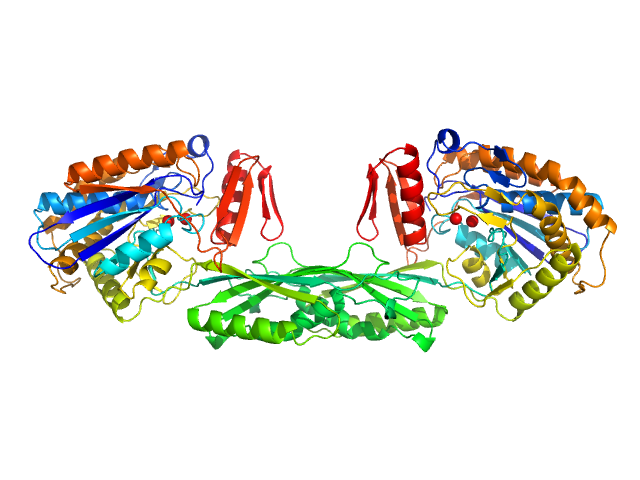
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 2 dimer, 100 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 May 8
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
126 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
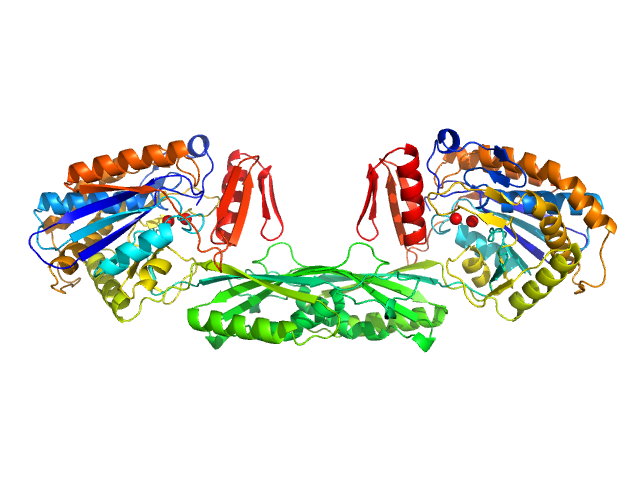
Methotrexate dimer, 1 kDa
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 2 dimer, 100 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 May 8
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.1 |
nm |
| VolumePorod |
125 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
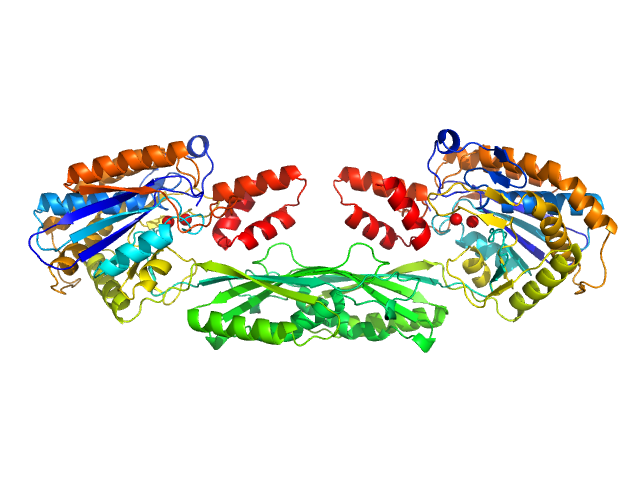
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 3 dimer, 100 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 May 8
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
127 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
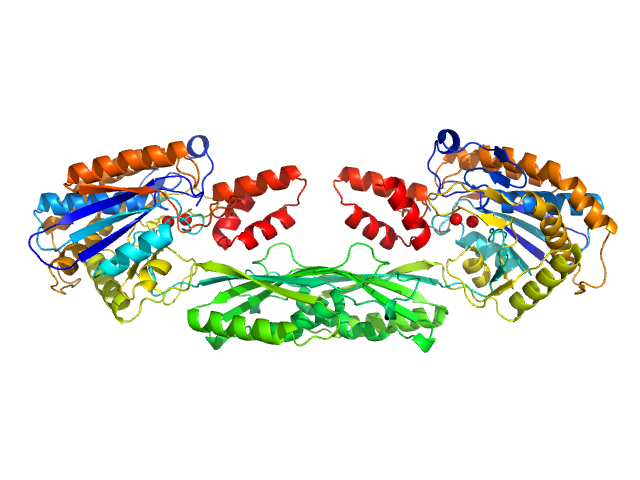
Methotrexate dimer, 1 kDa
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 3 dimer, 100 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 May 8
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.3 |
nm |
| VolumePorod |
125 |
nm3 |
|
|
|
|
|
|
|
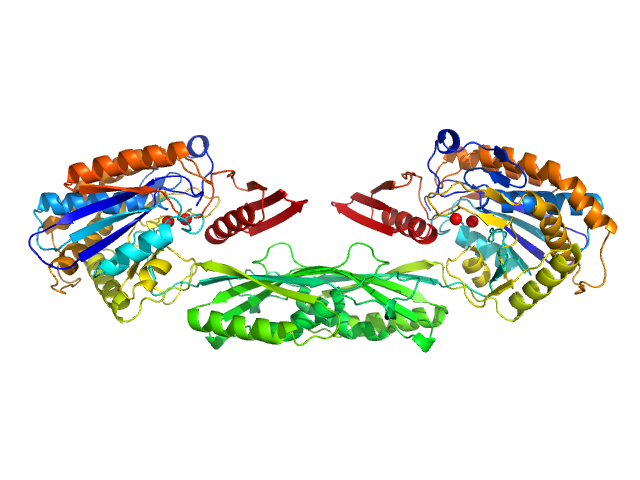
| Sample: |
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 1 Disulfide Variant dimer, 96 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Dec 6
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
|
|
|
|
|
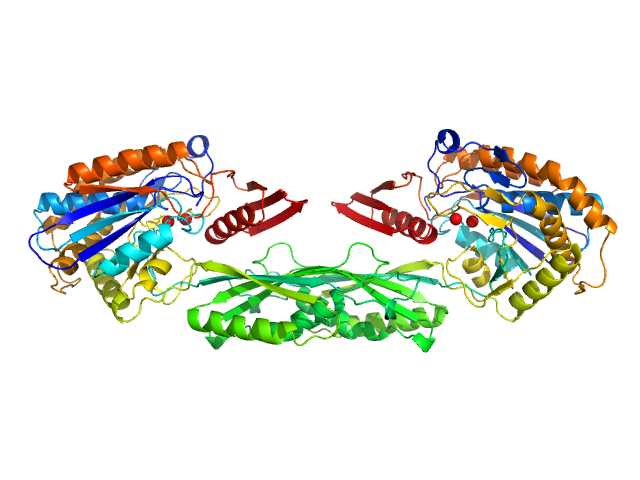
| Sample: |
Methotrexate dimer, 1 kDa
Pro-Carboxypeptidase G2 (circular permutant CP-N89) K177A Design 1 Disulfide Variant dimer, 96 kDa Pseudomonas sp. (strain … protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Dec 6
|
Massively parallel, computationally guided design of a proenzyme.
Proc Natl Acad Sci U S A 119(15):e2116097119 (2022)
Yachnin BJ, Azouz LR, White RE 3rd, Minetti CASA, Remeta DP, Tan VM, Drake JM, Khare SD
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.3 |
nm |
| VolumePorod |
125 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Gag-Pol polyprotein (HIV-1 reverse transcriptase C38V/C280S, L289K truncated to 1 - 556) monomer, 65 kDa Human immunodeficiency virus … protein
|
| Buffer: |
50 mM Bis-Tris pH 6.8, 100 mM NaCl, 1% glycerol, 0.02% w/v NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Jun 20
|
Relative domain orientation of the L289K HIV
‐1 reverse transcriptase monomer
Protein Science 31(5) (2022)
Xi Z, Ilina T, Guerrero M, Fan L, Sluis‐Cremer N, Wang Y, Ishima R
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
91 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Gag-Pol polyprotein (HIV-1 reverse transcriptase C38V/C280S, L289K truncated to 1 - 556) monomer, 65 kDa Human immunodeficiency virus … protein
|
| Buffer: |
50 mM Bis-Tris pH 6.8, 100 mM NaCl, 1% glycerol, 0.02% w/v NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Jun 20
|
Relative domain orientation of the L289K HIV
‐1 reverse transcriptase monomer
Protein Science 31(5) (2022)
Xi Z, Ilina T, Guerrero M, Fan L, Sluis‐Cremer N, Wang Y, Ishima R
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.8 |
nm |
| VolumePorod |
91 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Gag-Pol polyprotein (HIV-1 reverse transcriptase C38V/C280S, L289K truncated to 1 - 556) monomer, 65 kDa Human immunodeficiency virus … protein
|
| Buffer: |
50 mM Bis-Tris pH 6.8, 100 mM NaCl, 1% glycerol, 0.02% w/v NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Jun 20
|
Relative domain orientation of the L289K HIV
‐1 reverse transcriptase monomer
Protein Science 31(5) (2022)
Xi Z, Ilina T, Guerrero M, Fan L, Sluis‐Cremer N, Wang Y, Ishima R
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
93 |
nm3 |
|
|
|
|
|
|
|
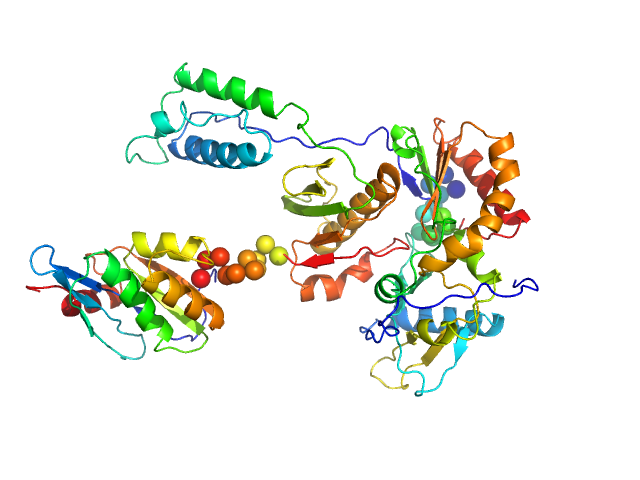
| Sample: |
Gag-Pol polyprotein (HIV-1 reverse transcriptase C38V/C280S, L289K truncated to 1 - 556) monomer, 65 kDa Human immunodeficiency virus … protein
|
| Buffer: |
50 mM Bis-Tris pH 6.8, 100 mM NaCl, 1% glycerol, 0.02% w/v NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Jun 20
|
Relative domain orientation of the L289K HIV
‐1 reverse transcriptase monomer
Protein Science 31(5) (2022)
Xi Z, Ilina T, Guerrero M, Fan L, Sluis‐Cremer N, Wang Y, Ishima R
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
90 |
nm3 |
|
|