|
|
|
|
|
| Sample: |
ABC transporter ATP-binding protein dimer, 62 kDa Streptococcus agalactiae protein
ABC transporter, permease protein monomer, 74 kDa Streptococcus agalactiae serotype … protein
|
| Buffer: |
50 mM Tris, 300 mM NaCl, 10 % Glycerol, 0.005 % LMNG,, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Apr 25
|
The extracellular domain of SaNSrFP binds bacitracin and allows the identification of new members of the BceAB transporter family
Frontiers in Microbiology 16 (2025)
Mammen C, Gottstein J, Cea P, Tantsur K, Reiners J, Bonus M, Gohlke H, Smits S
|
| RgGuinier |
5.8 |
nm |
| Dmax |
20.5 |
nm |
| VolumePorod |
735 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Non structural Protein 15 in Complex with ADPRP Macrodomain monomer, 254 kDa Severe acute respiratory … protein
|
| Buffer: |
20mM HEPES, 150mM Nacl, pH: 7.5 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2025 Mar 20
|
Non Structural Protein 15 in Complex with ADPRP Macrodomain of SARS CoV 2
Hira Singh Gariya
|
|
|
|
|
|
|
|
| Sample: |
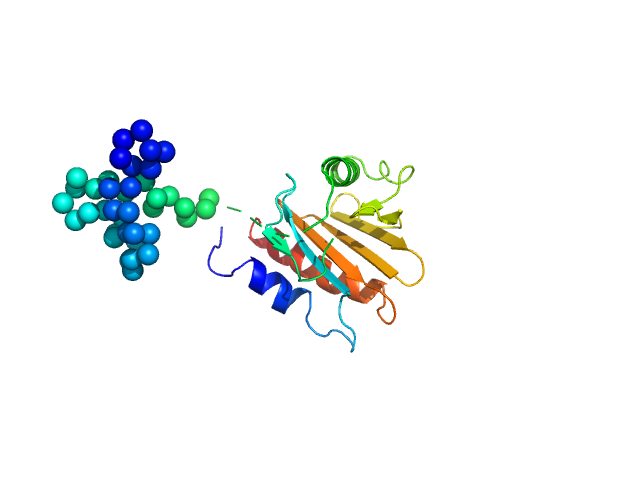
Profilin-2 monomer, 18 kDa Hevea brasiliensis protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 3
|
Allergen-induced structural rearrangements in IgE: insights from SAXS and molecular dynamics.
Int J Biol Macromol :147658 (2025)
Gómez-Velasco H, García-Ramírez B, Siliqi D, Graewert MA, Quintero-Martinez A, Ortega E, Rodríguez-Romero A
|
| RgGuinier |
2.0 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Profilin-2 monomer, 18 kDa Hevea brasiliensis protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2025 Apr 11
|
Allergen-induced structural rearrangements in IgE: insights from SAXS and molecular dynamics.
Int J Biol Macromol :147658 (2025)
Gómez-Velasco H, García-Ramírez B, Siliqi D, Graewert MA, Quintero-Martinez A, Ortega E, Rodríguez-Romero A
|
| RgGuinier |
1.7 |
nm |
| Dmax |
4.5 |
nm |
| VolumePorod |
21 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
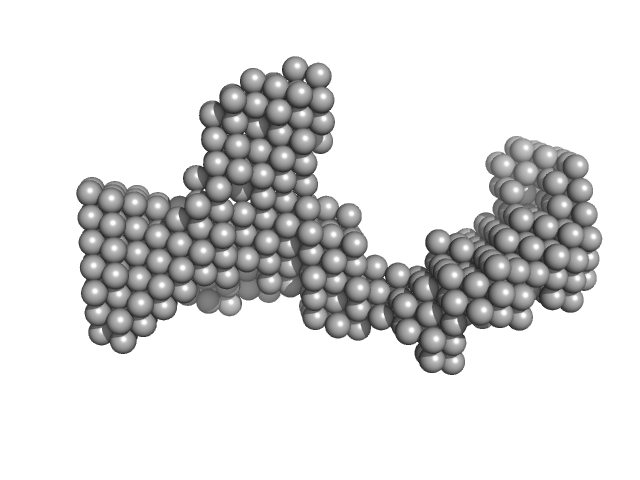
DENV2 5' untranslated terminal region monomer, 57 kDa RNA
|
| Buffer: |
10 mM Bis-Tris, 100 mM NaCl, 5 mM MgCl2, 15 mM KCl, 5% glycerol, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Sep 10
|
Structural Dynamics of Dengue Virus UTRs and Their Cyclization.
Biophys J (2025)
Robinson ZE, Pereira HS, D'Souza MH, Patel TR
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
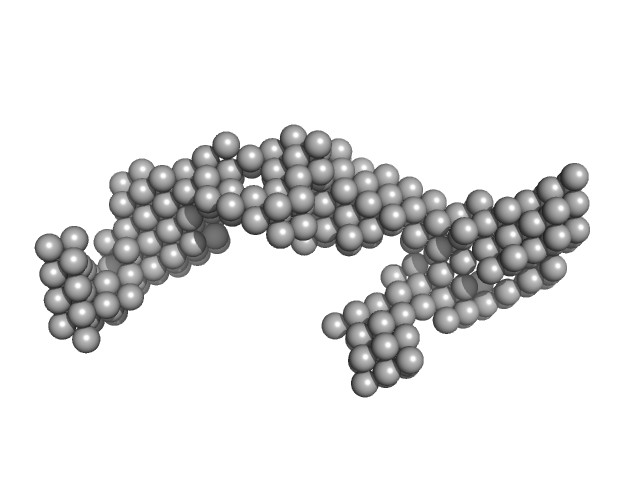
DENV2 3' untranslated region monomer, 160 kDa RNA
|
| Buffer: |
10 mM Bis-Tris, 100 mM NaCl, 5 mM MgCl2, 15 mM KCl, 5% glycerol, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Sep 10
|
Structural Dynamics of Dengue Virus UTRs and Their Cyclization.
Biophys J (2025)
Robinson ZE, Pereira HS, D'Souza MH, Patel TR
|
| RgGuinier |
6.0 |
nm |
| Dmax |
18.7 |
nm |
| VolumePorod |
155 |
nm3 |
|
|
|
|
|
|
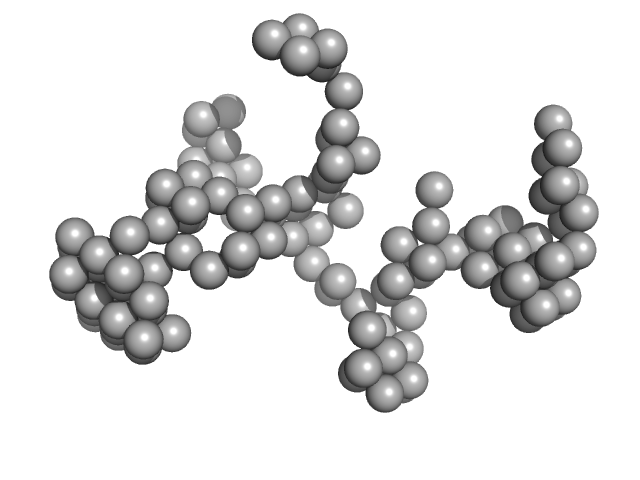
|
| Sample: |
DENV2 complex 5'-3' monomer, 217 kDa RNA
|
| Buffer: |
10 mM Bis-Tris, 100 mM NaCl, 5 mM MgCl2, 15 mM KCl, 5% glycerol, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Sep 10
|
Structural Dynamics of Dengue Virus UTRs and Their Cyclization.
Biophys J (2025)
Robinson ZE, Pereira HS, D'Souza MH, Patel TR
|
| RgGuinier |
7.5 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
320 |
nm3 |
|
|
|
|
|
|
|
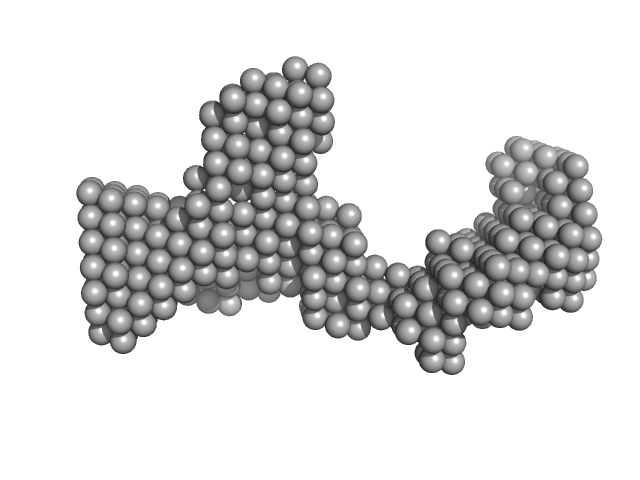
| Sample: |
DENV2 RNA, 5' Untranslated region monomer, 57 kDa RNA
|
| Buffer: |
10 mM Bis-Tris, 100 mM NaCl, 5 mM MgCl2, 15 mM KCl, 5% glycerol, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Sep 8
|
Structural Dynamics of Dengue Virus UTRs and Their Cyclization.
Biophys J (2025)
Robinson ZE, Pereira HS, D'Souza MH, Patel TR
|
| RgGuinier |
3.7 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
|
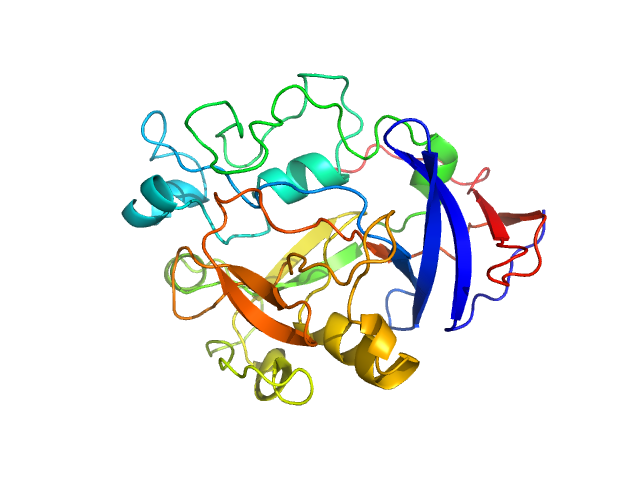
| Sample: |
IgA protease monomer, 32 kDa Thomasclavelia ramosa protein
|
| Buffer: |
25 mM HEPES, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2023 Jun 26
|
Structure of the Thomasclavelia ramosa immunoglobulin A protease reveals a modular and minimizable architecture distinct from other immunoglobulin A proteases.
Proc Natl Acad Sci U S A 122(35):e2503549122 (2025)
Tran N, Frenette A, Holyoak T
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.0 |
nm |
| VolumePorod |
38 |
nm3 |
|
|
|
|
|
|
|
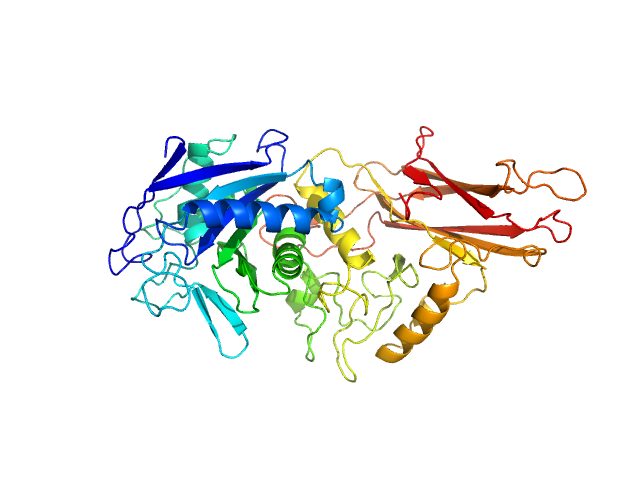
| Sample: |
IgA protease monomer, 55 kDa Thomasclavelia ramosa protein
|
| Buffer: |
25 mM HEPES, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2023 Jun 26
|
Structure of the Thomasclavelia ramosa immunoglobulin A protease reveals a modular and minimizable architecture distinct from other immunoglobulin A proteases.
Proc Natl Acad Sci U S A 122(35):e2503549122 (2025)
Tran N, Frenette A, Holyoak T
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
82 |
nm3 |
|
|