|
|
|
|
|
| Sample: |
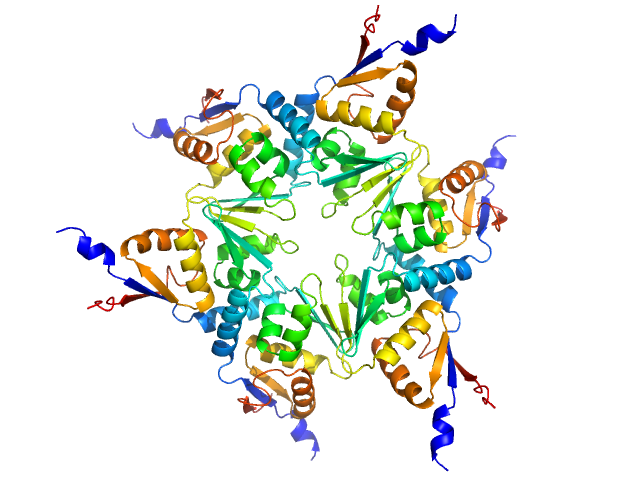
Longitudinals lacking protein, isoform G hexamer, 92 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM Tris, pH 7.4, 200 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 8
|
BTB domains: A structural view of evolution, multimerization, and protein-protein interactions.
Bioessays :e2200179 (2022)
Bonchuk A, Balagurov K, Georgiev P
|
| RgGuinier |
4.1 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
213 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Longitudinals lacking protein, isoform G hexamer, 92 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM Tris, pH 7.4, 200 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 8
|
BTB domains: A structural view of evolution, multimerization, and protein-protein interactions.
Bioessays :e2200179 (2022)
Bonchuk A, Balagurov K, Georgiev P
|
| RgGuinier |
4.8 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
263 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
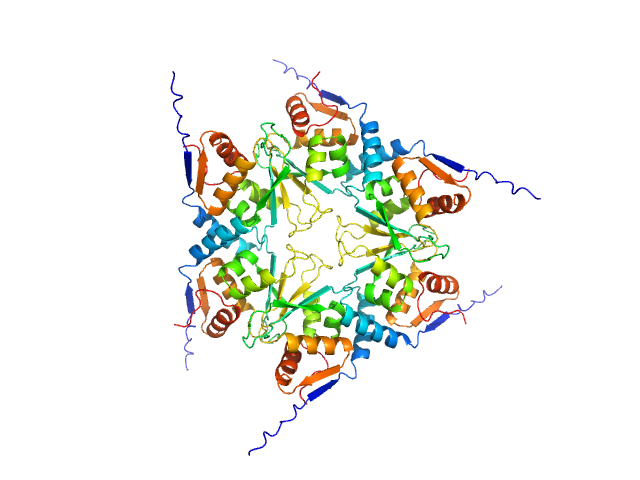
Uncharacterized protein, isoform A hexamer, 92 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM Tris, pH 7.4, 200 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 8
|
BTB domains: A structural view of evolution, multimerization, and protein-protein interactions.
Bioessays :e2200179 (2022)
Bonchuk A, Balagurov K, Georgiev P
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
167 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Uncharacterized protein, isoform A hexamer, 92 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM Tris, pH 7.4, 200 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jul 8
|
BTB domains: A structural view of evolution, multimerization, and protein-protein interactions.
Bioessays :e2200179 (2022)
Bonchuk A, Balagurov K, Georgiev P
|
| RgGuinier |
4.1 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
192 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
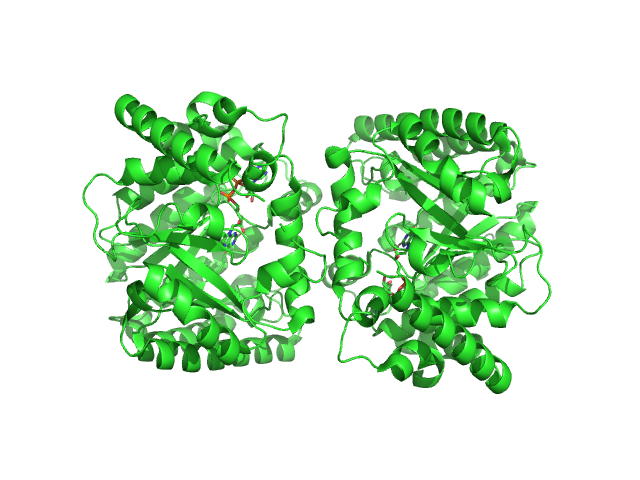
Minimal proline dehydrogenase domain of proline utilization A (design #2) dimer, 87 kDa Sinorhizobium meliloti protein
|
| Buffer: |
25 mM HEPES pH 7.6, 150 mM NaCl, and 1mM TCEP, pH: 7.6 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2022 Apr 12
|
Structure-based engineering of minimal Proline dehydrogenase domains for inhibitor discovery.
Protein Eng Des Sel (2022)
Bogner AN, Ji J, Tanner JJ
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
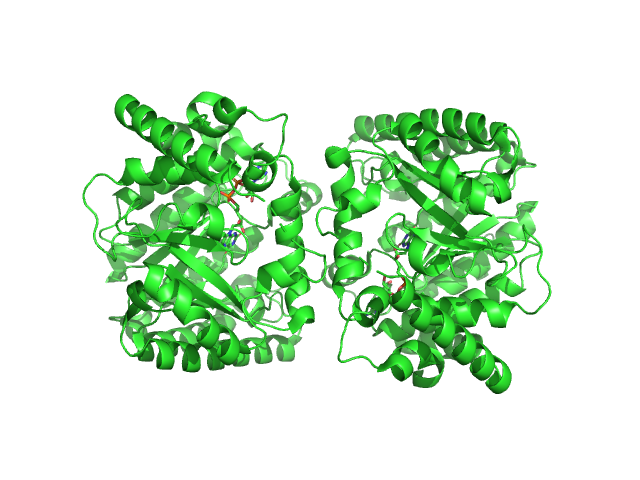
Minimal proline dehydrogenase domain of proline utilization A (design #2) dimer, 87 kDa Sinorhizobium meliloti protein
|
| Buffer: |
25 mM HEPES pH 7.6, 150 mM NaCl, and 1mM TCEP, pH: 7.6 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2022 Apr 12
|
Structure-based engineering of minimal Proline dehydrogenase domains for inhibitor discovery.
Protein Eng Des Sel (2022)
Bogner AN, Ji J, Tanner JJ
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Minimal proline dehydrogenase domain of proline utilization A (design #2) dimer, 87 kDa Sinorhizobium meliloti protein
|
| Buffer: |
25 mM HEPES pH 7.6, 150 mM NaCl, and 1mM TCEP, pH: 7.6 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2022 Apr 12
|
Structure-based engineering of minimal Proline dehydrogenase domains for inhibitor discovery.
Protein Eng Des Sel (2022)
Bogner AN, Ji J, Tanner JJ
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
108 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2020 Mar 7
|
Inline small‐angle X‐ray scattering‐coupled chromatography under extreme hydrostatic pressure
Protein Science 31(12) (2022)
Miller R, Cummings C, Huang Q, Ando N, Gillilan R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
253 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2020 Mar 7
|
Inline small‐angle X‐ray scattering‐coupled chromatography under extreme hydrostatic pressure
Protein Science 31(12) (2022)
Miller R, Cummings C, Huang Q, Ando N, Gillilan R
|
| RgGuinier |
3.5 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
267 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
DHPC - 1,2-dihexanoyl-sn-glycero-3-phosphocholine None, lipid
DMPC - 1,2-dimyristoyl-sn-glycero-3-phosphocholine None, lipid
|
| Buffer: |
Tris buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Jun 1
|
Expanding the Toolbox for Bicelle-Forming Surfactant–Lipid Mixtures
Molecules 27(21):7628 (2022)
Giudice R, Paracini N, Laursen T, Blanchet C, Roosen-Runge F, Cárdenas M
|
|
|