|
|
|
|
|
| Sample: |
Ox40 3 UTR Bulge monomer, 11 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 1
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
| RgGuinier |
1.8 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
17 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ox40 3 UTR Bulge-ADE monomer, 27 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 1
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
37 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ox40 3 UTR Bulge-ADE-CDE monomer, 35 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 1
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ox40 3 UTR CDEshort monomer, 5 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Jun 1
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
|
|
|
|
|
|
|
| Sample: |
Ox40 3 UTR Terminusuucg monomer, 10 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Aug 7
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.5 |
nm |
| VolumePorod |
16 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ox40 3 UTR ADEshort monomer, 7 kDa Mus musculus RNA
|
| Buffer: |
50 mM KCl, 25 mM sodium phosphate, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Aug 7
|
NMR-derived secondary structure of the full-length Ox40 mRNA 3'UTR and its multivalent binding to the immunoregulatory RBP Roquin.
Nucleic Acids Res (2022)
Tants JN, Becker LM, McNicoll F, Müller-McNicoll M, Schlundt A
|
| RgGuinier |
1.7 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
12 |
nm3 |
|
|
|
|
|
|
|
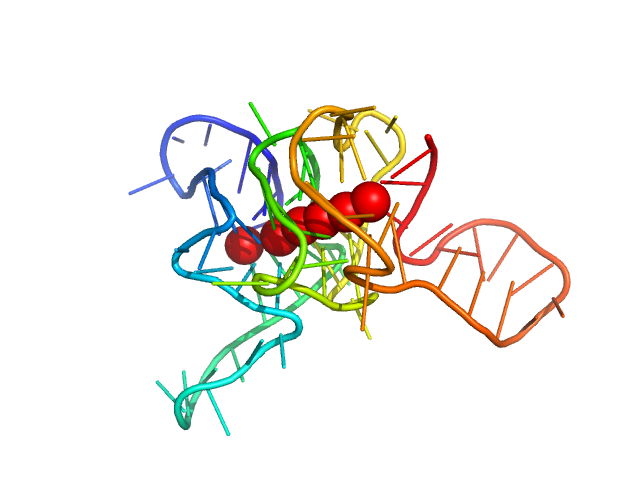
| Sample: |
c-Myc 12-tract G-quadruplex monomer Dnase I treated monomer, 22 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Sep 29
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
c-Kit 12-tract G-quadruplex monomer Dnase I treated monomer, 20 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Sep 29
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.7 |
nm |
| VolumePorod |
16 |
nm3 |
|
|
|
|
|
|
|
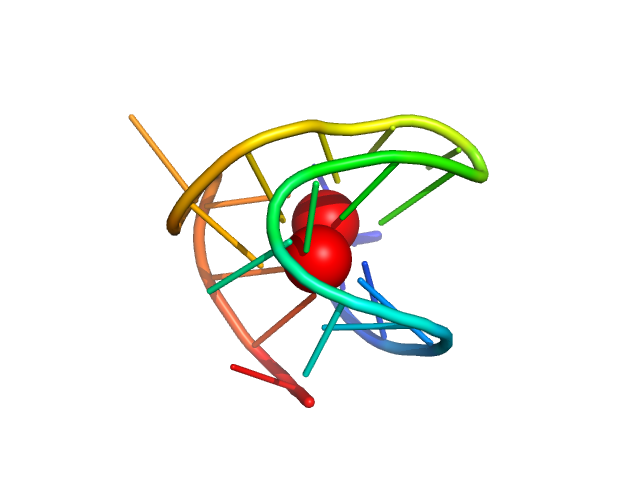
| Sample: |
TERT promoter G-quadruplex antiparallel monomer, 6 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Aug 13
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.2 |
nm |
| Dmax |
3.6 |
nm |
| VolumePorod |
7 |
nm3 |
|
|
|
|
|
|
|
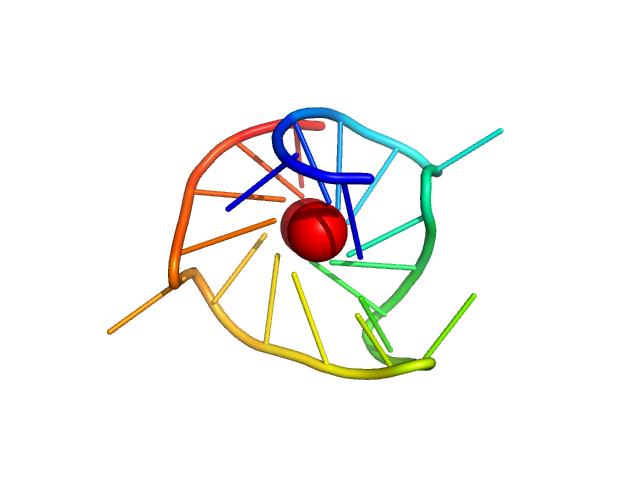
| Sample: |
TERT promoter G-quadruplex parallel monomer, 6 kDa synthetic construct DNA
|
| Buffer: |
6 mM Na2HPO4, 2 mM NaH2PO4, 1 mM Na2EDTA, 185 mM KCl, pH: 7.2
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Aug 11
|
Long promoter sequences form higher-order G-quadruplexes: an integrative structural biology study of c-Myc, k-Ras and c-Kit promoter sequences.
Nucleic Acids Res (2022)
Monsen RC, DeLeeuw LW, Dean WL, Gray RD, Chakravarthy S, Hopkins JB, Chaires JB, Trent JO
|
| RgGuinier |
1.3 |
nm |
| Dmax |
4.2 |
nm |
| VolumePorod |
8 |
nm3 |
|
|