|
|
|
|
|
| Sample: |
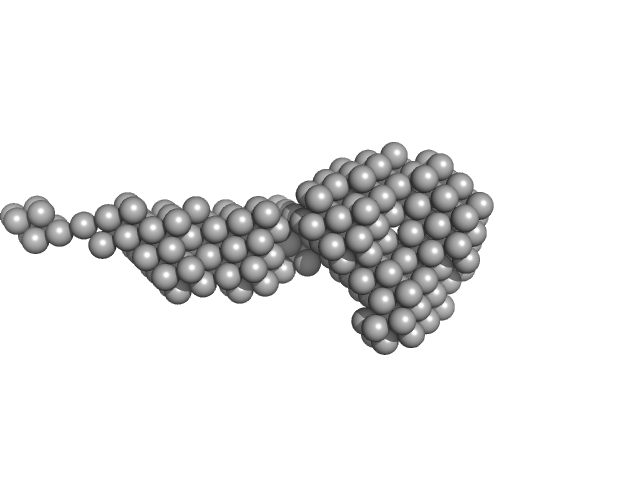
Lectin nano-block WA20-ΔN3ACG dimer, 58 kDa protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5% glycerol,, pH: 7.5
|
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Oct 31
|
Self-Assembling Lectin Nano-Block Oligomers Enhance Binding Avidity to Glycans
International Journal of Molecular Sciences 23(2):676 (2022)
Irumagawa S, Hiemori K, Saito S, Tateno H, Arai R
|
| RgGuinier |
4.1 |
nm |
| Dmax |
21.6 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cytochrome c' monomer, 14 kDa Achromobacter xylosoxidans protein
|
| Buffer: |
Phosphate Buffer pD 1.7, pH: 1.7
|
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Aug 12
|
Open-Bundle Structure as the Unfolding Intermediate of Cytochrome c′ Revealed by Small Angle Neutron Scattering
Biomolecules 12(1):95 (2022)
Yamaguchi T, Akao K, Koutsioubas A, Frielinghaus H, Kohzuma T
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
13 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cytochrome c' dimer, 27 kDa Alcaligenes protein
|
| Buffer: |
Phosphate Buffer pD 6.4, pH: 6.4
|
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Aug 12
|
Open-Bundle Structure as the Unfolding Intermediate of Cytochrome c′ Revealed by Small Angle Neutron Scattering
Biomolecules 12(1):95 (2022)
Yamaguchi T, Akao K, Koutsioubas A, Frielinghaus H, Kohzuma T
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.5 |
nm |
| VolumePorod |
11 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cytochrome c' dimer, 27 kDa Alcaligenes protein
|
| Buffer: |
Phosphate Buffer pD 9.6, pH: 9.6
|
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Aug 12
|
Open-Bundle Structure as the Unfolding Intermediate of Cytochrome c′ Revealed by Small Angle Neutron Scattering
Biomolecules 12(1):95 (2022)
Yamaguchi T, Akao K, Koutsioubas A, Frielinghaus H, Kohzuma T
|
| RgGuinier |
1.9 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
10 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cytochrome c' monomer, 14 kDa Achromobacter xylosoxidans protein
|
| Buffer: |
Phosphate Buffer pD 13, pH: 13
|
| Experiment: |
SANS
data collected at KWS1, FRM2 on 2017 Aug 12
|
Open-Bundle Structure as the Unfolding Intermediate of Cytochrome c′ Revealed by Small Angle Neutron Scattering
Biomolecules 12(1):95 (2022)
Yamaguchi T, Akao K, Koutsioubas A, Frielinghaus H, Kohzuma T
|
| RgGuinier |
4.8 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
20 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
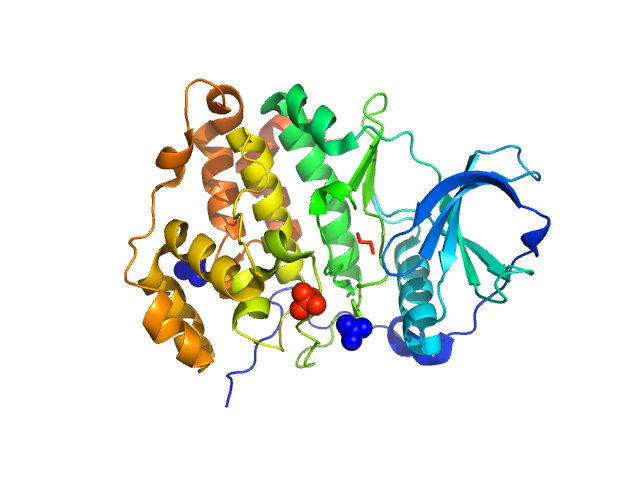
Casein kinase II subunit alpha monomer, 39 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 500 mM NaCl, pH: 8.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Oct 10
|
Mechanism of CK2 Inhibition by a Ruthenium-Based Polyoxometalate.
Front Mol Biosci 9:906390 (2022)
Fabbian S, Giachin G, Bellanda M, Borgo C, Ruzzene M, Spuri G, Campofelice A, Veneziano L, Bonchio M, Carraro M, Battistutta R
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
59 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
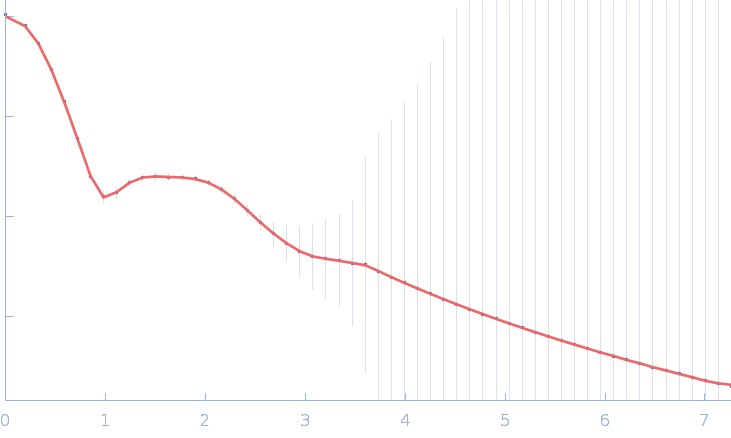
Francisella tularensis outer membrane protein A dimer, 80 kDa Francisella tularensis subsp. … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.05% B-DDM, pH: 7.5
|
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Mar 20
|
Structural and biophysical properties of FopA, a major outer membrane protein of Francisella tularensis.
PLoS One 17(8):e0267370 (2022)
Nagaratnam N, Martin-Garcia JM, Yang JH, Goode MR, Ketawala G, Craciunescu FM, Zook JD, Sonowal M, Williams D, Grant TD, Fromme R, Hansen DT, Fromme P
|
| RgGuinier |
4.4 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
330 |
nm3 |
|
|
|
|
|
|
|
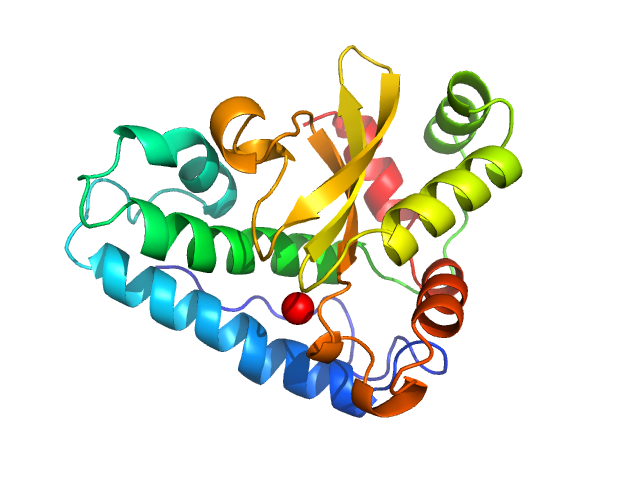
| Sample: |
Superoxide dismutase [Mn] , 23 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 1
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.3 |
nm |
| VolumePorod |
54 |
nm3 |
|
|
|
|
|
|
|
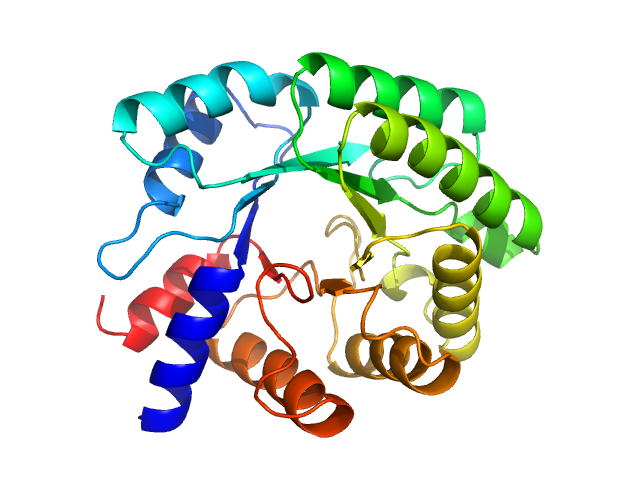
| Sample: |
Deoxyribose-phosphate aldolase , 28 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 1
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
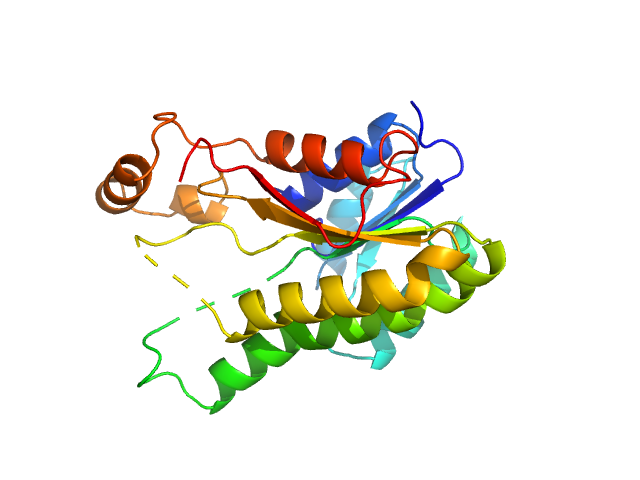
3-oxoacyl-[acyl-carrier-protein] reductase FabG , 26 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 1
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
3.6 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
212 |
nm3 |
|
|