|
|
|
|
|
| Sample: |
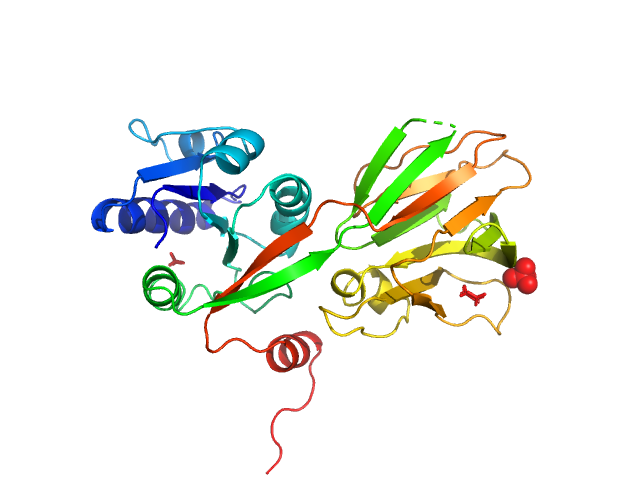
NAD kinase , 33 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 2
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Anti-CD20 IgG antibody (Rituximab heavy chain chimeric) dimer, 98 kDa Homo sapiens protein
Anti-CD20 IgG antibody (Rituximab light chain chimeric) dimer, 46 kDa Homo sapiens protein
|
| Buffer: |
100 mM HEPES, pH: 7.7
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Sep 20
|
Structural Characterization of the Full-Length Anti-CD20 Antibody Rituximab.
Front Mol Biosci 9:823174 (2022)
Belviso BD, Mangiatordi GF, Alberga D, Mangini V, Carrozzini B, Caliandro R
|
| RgGuinier |
5.2 |
nm |
| Dmax |
19.7 |
nm |
|
|
|
|
|
|
|
| Sample: |
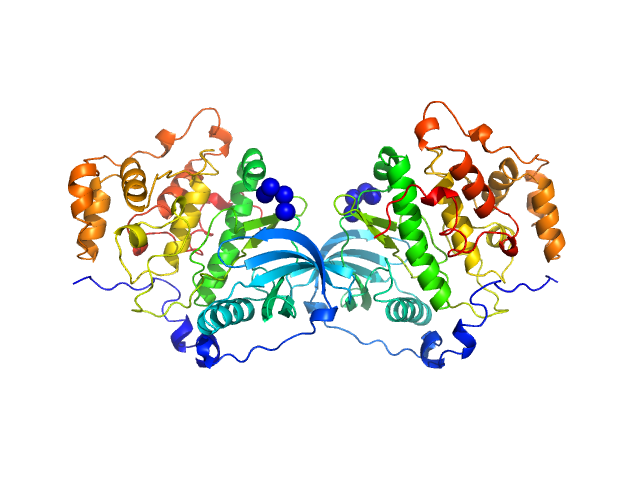
Casein kinase II subunit alpha dimer, 78 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 500 mM NaCl, pH: 8.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Oct 10
|
Mechanism of CK2 Inhibition by a Ruthenium-Based Polyoxometalate.
Front Mol Biosci 9:906390 (2022)
Fabbian S, Giachin G, Bellanda M, Borgo C, Ruzzene M, Spuri G, Campofelice A, Veneziano L, Bonchio M, Carraro M, Battistutta R
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
122 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Bovine Serum Albumin trimer, 199 kDa Bos taurus protein
|
| Buffer: |
20 mM Hepes, 150mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Sep 24
|
SEC-SAXS: Experimental set-up and software developments build up a powerful tool.
Methods Enzymol 677:221-249 (2022)
Pérez J, Thureau A, Vachette P
|
| RgGuinier |
5.2 |
nm |
| Dmax |
17.6 |
nm |
| VolumePorod |
370 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Eukaryotic initiation factor 4F subunit p150 , 28 kDa Saccharomyces cerevisiae (strain … protein
|
| Buffer: |
25 mM potassium phosphate, 25 mM NaCl, pH: 6.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 14
|
eIF4G1 N-terminal intrinsically disordered domain is a multi-docking station for RNA, Pab1, Pub1, and self-assembly.
Front Mol Biosci 9:986121 (2022)
Chaves-Arquero B, Martínez-Lumbreras S, Sibille N, Camero S, Bernadó P, Jiménez MÁ, Zorrilla S, Pérez-Cañadillas JM
|
| RgGuinier |
5.2 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
137 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
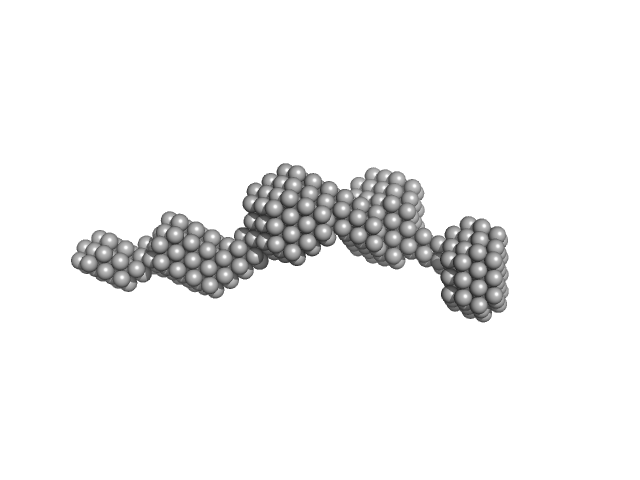
centrosomal-P4.1-associated-protein monomer, 12 kDa Homo sapiens protein
Centriolar coiled-coil protein of 110 kDa dimer, 20 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, pH 7.5, 100 mM NaCl, 1 mM DTT, 1 mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Feb 23
|
Centriolar cap proteins CP110 and CPAP control slow elongation of microtubule plus ends.
J Cell Biol 224(3) (2025)
Iyer SS, Chen F, Ogunmolu FE, Moradi S, Volkov VA, van Grinsven EJ, van Hoorn C, Wu J, Andrea N, Hua S, Jiang K, Vakonakis I, Potočnjak M, Herzog F, Gigant B, Gudimchuk N, Stecker KE, Dogterom M, Stei...
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.2 |
nm |
| VolumePorod |
35 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
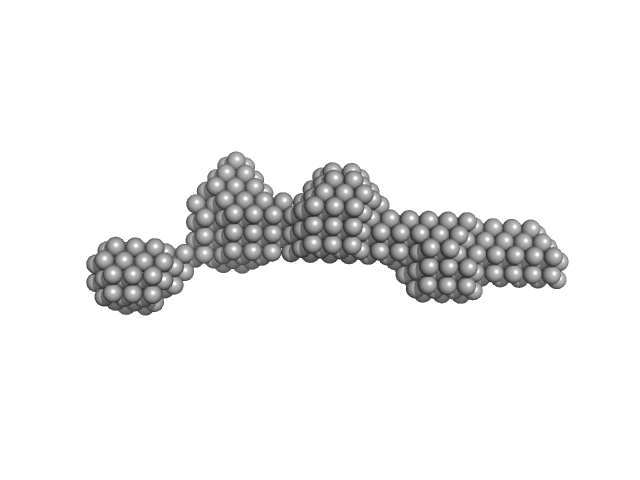
Centriolar coiled-coil protein of 110 kDa dimer, 20 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, pH 7.5, 100 mM NaCl, 1 mM DTT, 1 mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Feb 23
|
Centriolar cap proteins CP110 and CPAP control slow elongation of microtubule plus ends.
J Cell Biol 224(3) (2025)
Iyer SS, Chen F, Ogunmolu FE, Moradi S, Volkov VA, van Grinsven EJ, van Hoorn C, Wu J, Andrea N, Hua S, Jiang K, Vakonakis I, Potočnjak M, Herzog F, Gigant B, Gudimchuk N, Stecker KE, Dogterom M, Stei...
|
| RgGuinier |
3.5 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
33 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
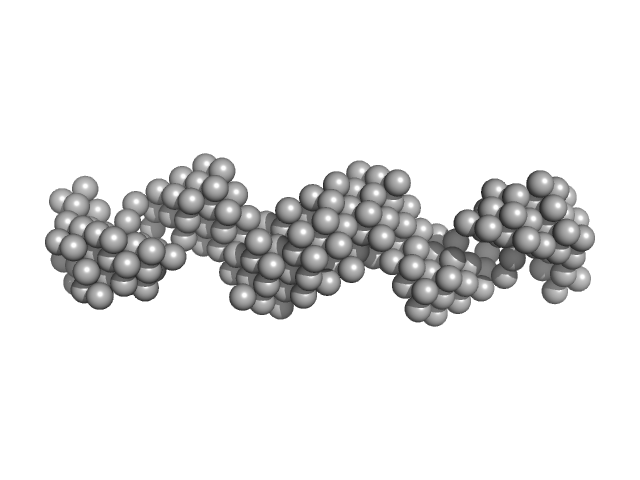
HomA outer membrane protein dimer, 146 kDa Helicobacter pylori protein
|
| Buffer: |
20 mM Tris-Cl, 200 mM NaCl, 5 mM β-mercaptoethanol, pH: 8
|
| Experiment: |
SAXS
data collected at BL-18, INDUS-2 on 2021 Feb 2
|
Biophysical characterization of the homodimers of HomA and HomB, outer membrane proteins of Helicobacter pylori.
Sci Rep 11(1):24471 (2021)
Tamrakar A, Singh R, Kumar A, Makde RD, Ashish, Kodgire P
|
| RgGuinier |
8.3 |
nm |
| Dmax |
28.2 |
nm |
| VolumePorod |
426 |
nm3 |
|
|
|
|
|
|
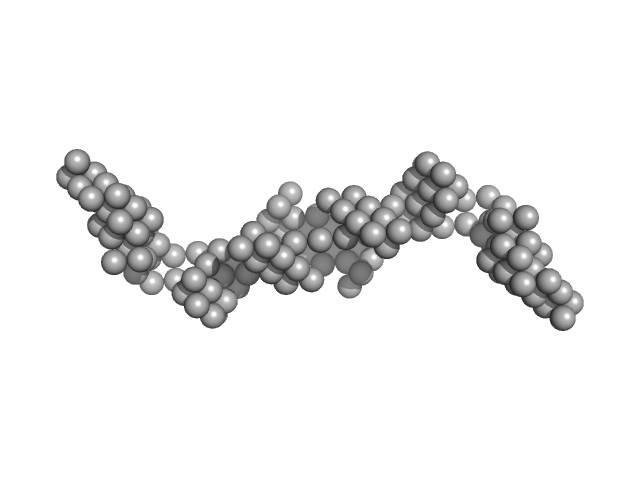
|
| Sample: |
HomB outer membrane protein dimer, 148 kDa Helicobacter pylori protein
|
| Buffer: |
20 mM Tris-Cl, 200 mM NaCl, 5 mM β-mercaptoethanol, pH: 8
|
| Experiment: |
SAXS
data collected at BL-18, INDUS-2 on 2021 Feb 2
|
Biophysical characterization of the homodimers of HomA and HomB, outer membrane proteins of Helicobacter pylori.
Sci Rep 11(1):24471 (2021)
Tamrakar A, Singh R, Kumar A, Makde RD, Ashish, Kodgire P
|
| RgGuinier |
8.2 |
nm |
| Dmax |
27.1 |
nm |
| VolumePorod |
500 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Synthetic FRET biosensor Twitch 2B monomer, 62 kDa synthetic construct protein
|
| Buffer: |
20 mM MOPS 100 mM NaCl 5 mM CaCl2, pH: 7
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Mar 17
|
Synthetic FRET biosensor Twitch 2B
Pablo Trigo Mourino
|
|
|