|
|
|
|

|
| Sample: |
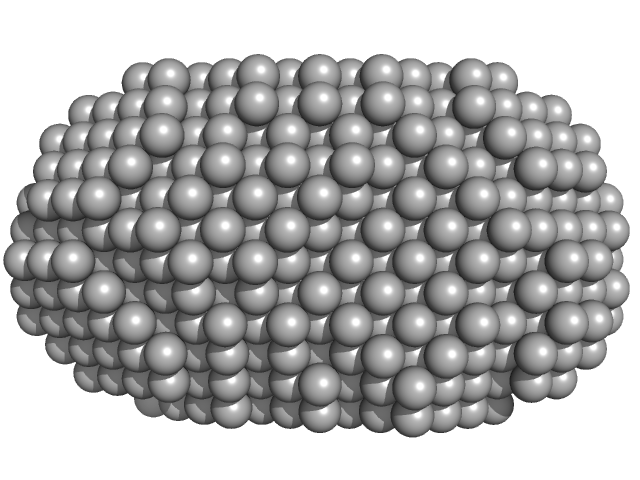
Very long-chain specific acyl-CoA dehydrogenase, mitochondrial dimer, 126 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Dec 2
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
215 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
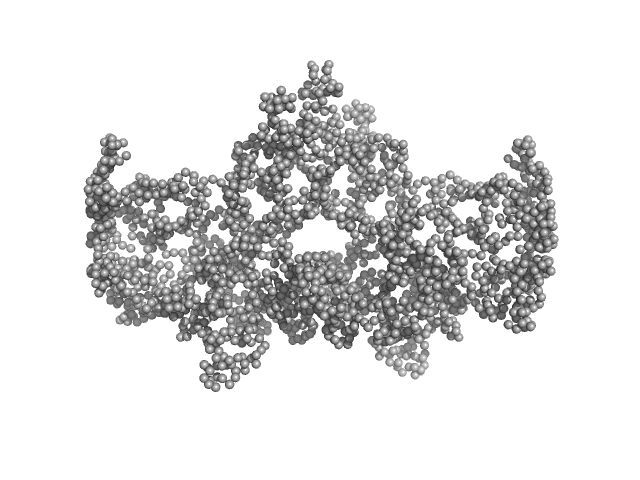
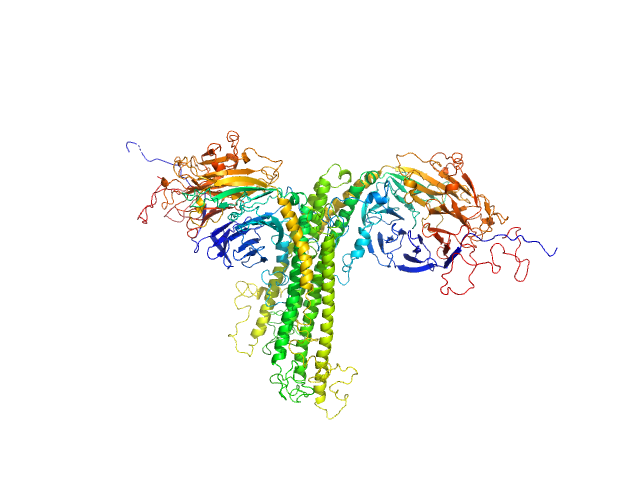
Complex I assembly factor ACAD9, mitochondrial dimer, 132 kDa Homo sapiens protein
Evolutionarily conserved signaling intermediate in Toll pathway, mitochondrial tetramer, 88 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 250 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Oct 4
|
Assembly of the mitochondrial Complex I assembly complex suggests a regulatory role for deflavination.
Angew Chem Int Ed Engl (2020)
Giachin G, Jessop M, Bouverot R, Acajjaoui S, Saidi M, Chretien A, Bacia-Verloop M, Signor L, Mas PJ, Favier A, Borel Meneroud E, Hons M, Hart DJ, Kandiah E, Boeri Erba E, Buisson A, Leonard G, Gutsch...
|
| RgGuinier |
6.0 |
nm |
| Dmax |
16.5 |
nm |
| VolumePorod |
683 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
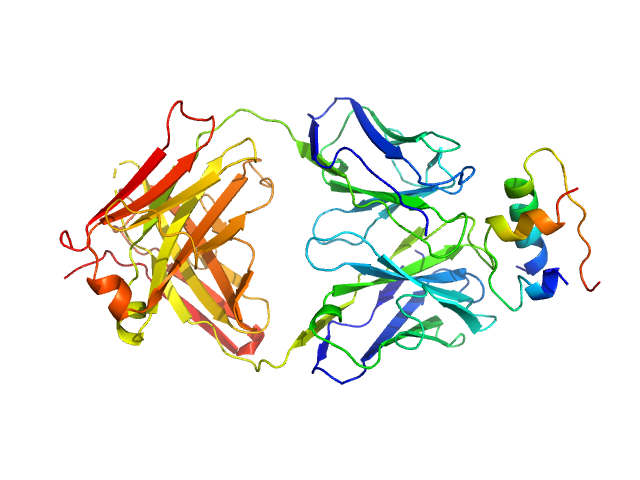
Anaphase Promoting Complex/Cyclosome Subunit 4 dimer, 184 kDa Arabidopsis thaliana protein
|
| Buffer: |
10 mM Tris 150 mM NaCl 1 mM TCEP, pH: 8
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Mar 6
|
Structural Characterisation of the Arabidopsis thaliana APC4
Steven De Gieter
|
| RgGuinier |
4.6 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
259 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
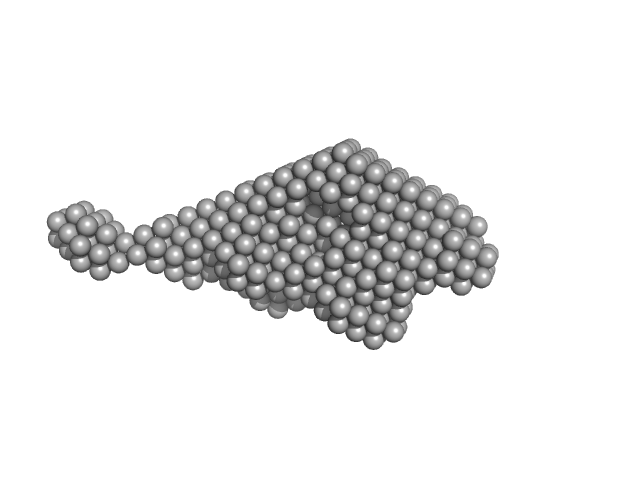
Testican-2 monomer, 40 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES (Na), 130 mM NaCl, 5 % (v/v) glycerol, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 18
|
The Central Region of Testican-2 Forms a Compact Core and Promotes Cell Migration
International Journal of Molecular Sciences 21(24):9413 (2020)
Krajnc A, Gaber A, Lenarčič B, Pavšič M
|
| RgGuinier |
3.4 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Testican-2 monomer, 40 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES (Na), 130 mM NaCl, 3 mM CaCl2, 5 % (v/v) glycerol, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 18
|
The Central Region of Testican-2 Forms a Compact Core and Promotes Cell Migration
International Journal of Molecular Sciences 21(24):9413 (2020)
Krajnc A, Gaber A, Lenarčič B, Pavšič M
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
78 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
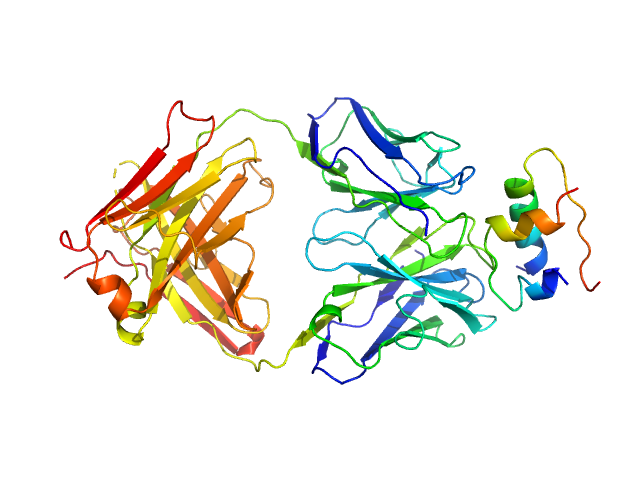
Insulin (Insulin B chain and Insulin A chain) monomer, 6 kDa Homo sapiens protein
HUI-018 Fab monomer, 47 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2013 Mar 7
|
Insulin Binding to the Analytical Antibody Sandwich Pair OXI
‐005 and HUI
‐018 – Epitope Mapping and Binding Properties
Protein Science (2020)
Johansson E, Wu X, Yu B, Yang Z, Cao Z, Wiberg C, Jeppesen C, Poulsen F
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
86 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Insulin (Insulin B chain and Insulin A chain) monomer, 6 kDa Homo sapiens protein
HUI-018 Fab monomer, 47 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2013 Mar 7
|
Insulin Binding to the Analytical Antibody Sandwich Pair OXI
‐005 and HUI
‐018 – Epitope Mapping and Binding Properties
Protein Science (2020)
Johansson E, Wu X, Yu B, Yang Z, Cao Z, Wiberg C, Jeppesen C, Poulsen F
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
105 |
nm3 |
|
|
|
|
|
|
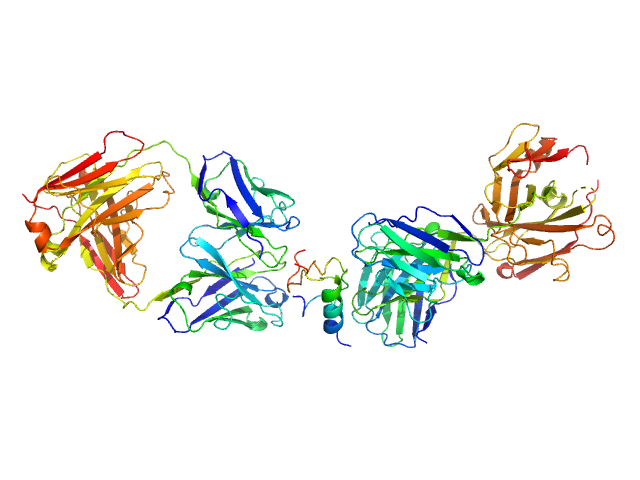
|
| Sample: |
HUI-018 Fab monomer, 47 kDa Mus musculus protein
Insulin (Insulin B chain and Insulin A chain) monomer, 6 kDa Sus scrofa protein
OXI-005 Fab monomer, 47 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 140 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Novo Nordisk A/S on 2019 Mar 28
|
Insulin Binding to the Analytical Antibody Sandwich Pair OXI
‐005 and HUI
‐018 – Epitope Mapping and Binding Properties
Protein Science (2020)
Johansson E, Wu X, Yu B, Yang Z, Cao Z, Wiberg C, Jeppesen C, Poulsen F
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.2 |
nm |
| VolumePorod |
138 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
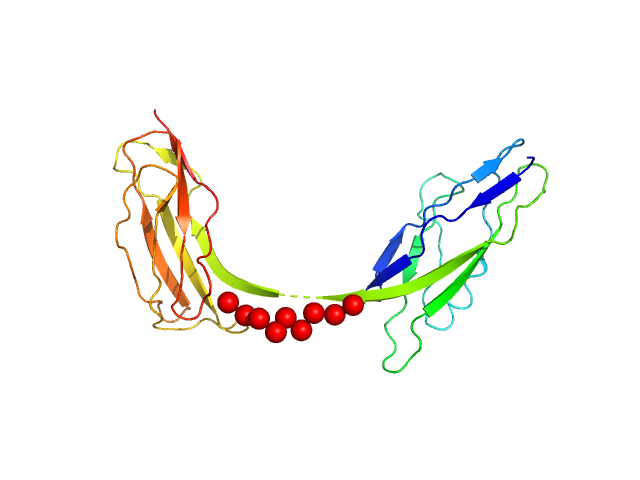
Intimin, D00-D0 domain (6xHis tagged) monomer, 23 kDa Escherichia coli O127:H6 … protein
|
| Buffer: |
10 mM HEPES, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Jul 12
|
The extracellular juncture domains in the intimin passenger adopt a constitutively extended conformation inducing restraints to its sphere of action.
Sci Rep 10(1):21249 (2020)
Weikum J, Kulakova A, Tesei G, Yoshimoto S, Jægerum LV, Schütz M, Hori K, Skepö M, Harris P, Leo JC, Morth JP
|
| RgGuinier |
3.1 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Intimin (D0-D1 domain, 6xHis tagged) monomer, 23 kDa Escherichia coli O127:H6 … protein
|
| Buffer: |
10 mM HEPES, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Jul 12
|
The extracellular juncture domains in the intimin passenger adopt a constitutively extended conformation inducing restraints to its sphere of action.
Sci Rep 10(1):21249 (2020)
Weikum J, Kulakova A, Tesei G, Yoshimoto S, Jægerum LV, Schütz M, Hori K, Skepö M, Harris P, Leo JC, Morth JP
|
| RgGuinier |
2.2 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
34 |
nm3 |
|
|