|
|
|
|
|
| Sample: |
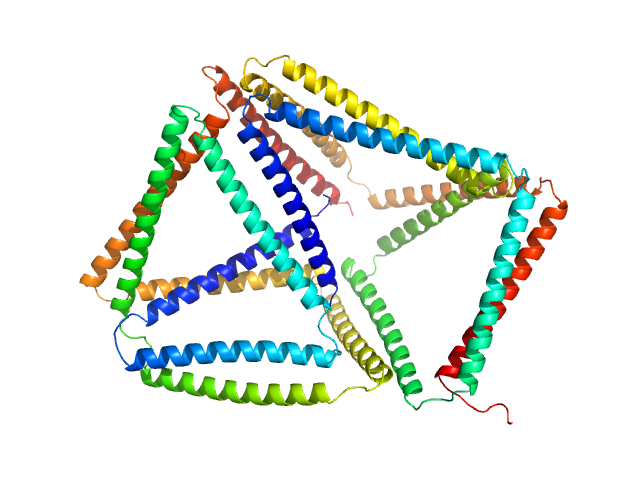
SBP1(9.b) monomer, 40 kDa synthetic construct protein
SBP2(9.b) monomer, 40 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 10% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Jun 19
|
Self-assembly and regulation of protein cages from pre-organised coiled-coil modules.
Nat Commun 12(1):939 (2021)
Lapenta F, Aupič J, Vezzoli M, Strmšek Ž, Da Vela S, Svergun DI, Carazo JM, Melero R, Jerala R
|
| RgGuinier |
4.0 |
nm |
| Dmax |
11.8 |
nm |
|
|
|
|
|
|
|
| Sample: |
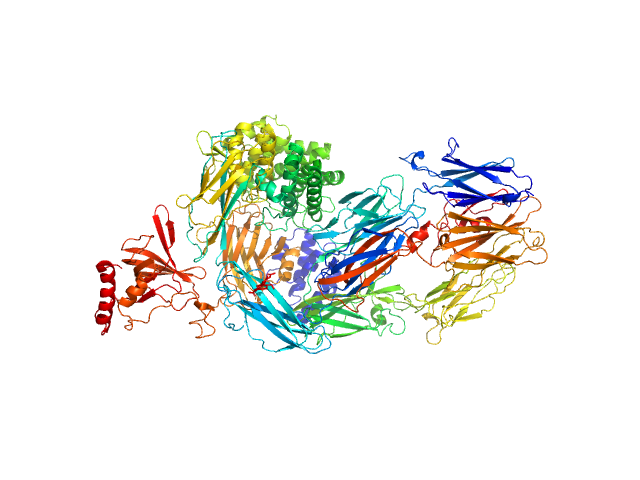
Complement C5 monomer, 186 kDa Homo sapiens protein
|
| Buffer: |
20mM Tris pH, 75mM NaCl, and 3% glycerol, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Sep 12
|
The allosteric modulation of Complement C5 by knob domain peptides.
Elife 10 (2021)
Macpherson A, Laabei M, Ahdash Z, Graewert MA, Birtley JR, Schulze ME, Crennell S, Robinson SA, Holmes B, Oleinikovas V, Nilsson PH, Snowden J, Ellis V, Mollnes TE, Deane CM, Svergun D, Lawson AD, van...
|
| RgGuinier |
4.8 |
nm |
| Dmax |
17.6 |
nm |
| VolumePorod |
392 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
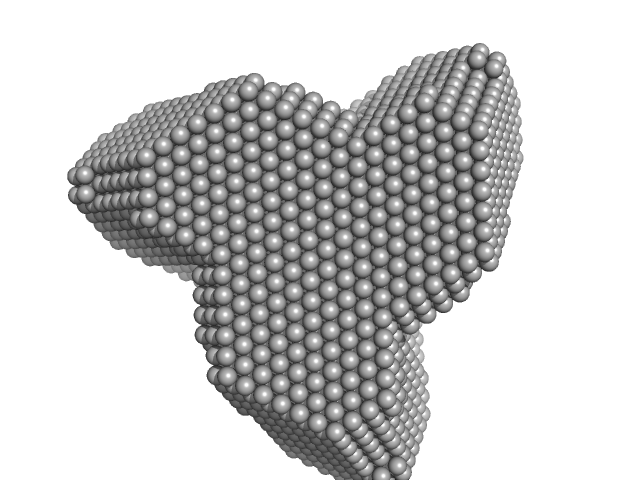
TET12(1.10)SN-f5(2CC) monomer, 55 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 10% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Dec 13
|
Designed folding pathway of modular coiled-coil-based proteins.
Nat Commun 12(1):940 (2021)
Aupič J, Strmšek Ž, Lapenta F, Pahovnik D, Pisanski T, Drobnak I, Ljubetič A, Jerala R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.6 |
nm |
| VolumePorod |
158 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
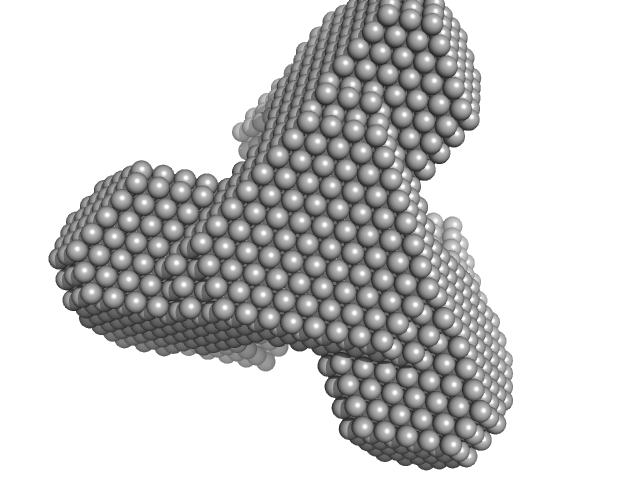
TET12(1.10)SN-f5(22CC) monomer, 55 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 10% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 5
|
Designed folding pathway of modular coiled-coil-based proteins.
Nat Commun 12(1):940 (2021)
Aupič J, Strmšek Ž, Lapenta F, Pahovnik D, Pisanski T, Drobnak I, Ljubetič A, Jerala R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.4 |
nm |
| VolumePorod |
165 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
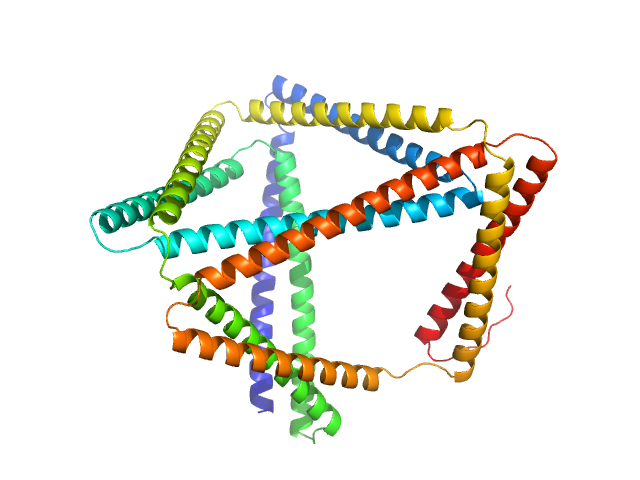
TET12(1.10)SN-f5(222CC) monomer, 55 kDa synthetic construct protein
|
| Buffer: |
20 mM Tris 150 mM NaCl 10% glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Sep 15
|
Designed folding pathway of modular coiled-coil-based proteins.
Nat Commun 12(1):940 (2021)
Aupič J, Strmšek Ž, Lapenta F, Pahovnik D, Pisanski T, Drobnak I, Ljubetič A, Jerala R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
170 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Phage-encoded SAM lyase Svi3-3 (including N-terminal His6-tag and Tev cleavage site) trimer, 56 kDa Unknown environmental phage protein
|
| Buffer: |
25 mM Tris-HCl, 150 mM NaCl, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Dec 9
|
Structure and mechanism of a phage-encoded SAM lyase revises catalytic function of enzyme family.
Elife 10 (2021)
Guo X, Söderholm A, Kanchugal P S, Isaksen GV, Warsi O, Eckhard U, Trigüis S, Gogoll A, Jerlström-Hultqvist J, Åqvist J, Andersson DI, Selmer M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Phage-encoded SAM lyase Svi3-3 (including N-terminal His6-tag and Tev cleavage site) trimer, 56 kDa Unknown environmental phage protein
|
| Buffer: |
25 mM Tris-HCl, 150 mM NaCl, 5 mM SAM, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2016 Dec 9
|
Structure and mechanism of a phage-encoded SAM lyase revises catalytic function of enzyme family.
Elife 10 (2021)
Guo X, Söderholm A, Kanchugal P S, Isaksen GV, Warsi O, Eckhard U, Trigüis S, Gogoll A, Jerlström-Hultqvist J, Åqvist J, Andersson DI, Selmer M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
89 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
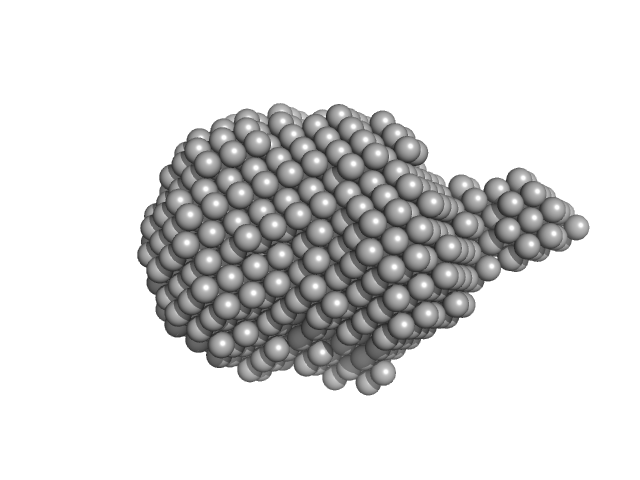
Ferric iron reductase protein FhuF (∆1-17) monomer, 28 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Phosphate, 200 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Apr 16
|
Conjuring up a ghost: structural and functional characterization of FhuF, a ferric siderophore reductase from E. coli
JBIC Journal of Biological Inorganic Chemistry (2021)
Trindade I, Hernandez G, Lebègue E, Barrière F, Cordeiro T, Piccioli M, Louro R
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
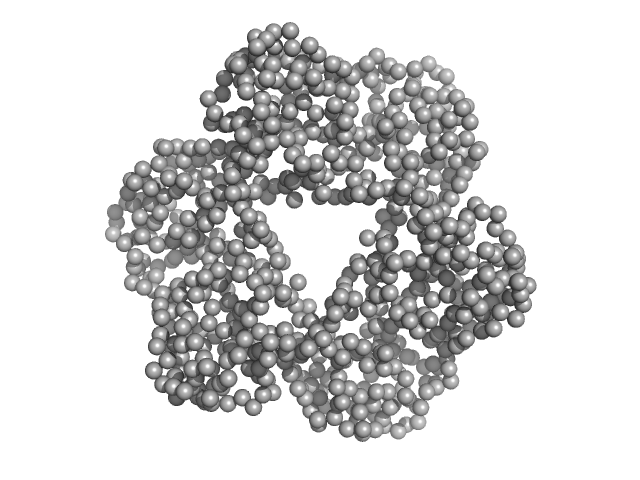
Proliferating cell nuclear antigen (C-terminal His-tagged) trimer, 90 kDa Candida albicans protein
|
| Buffer: |
20 mM Tris-HCl (pH 7.5), 100 mM NaCl, 1 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Sep 3
|
Structural analyses of PCNA from the fungal pathogen Candida albicans identify three regions with species-specific conformations.
FEBS Lett (2021)
Sundaram R, Manohar K, Patel SK, Acharya N, Vasudevan D
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
128 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
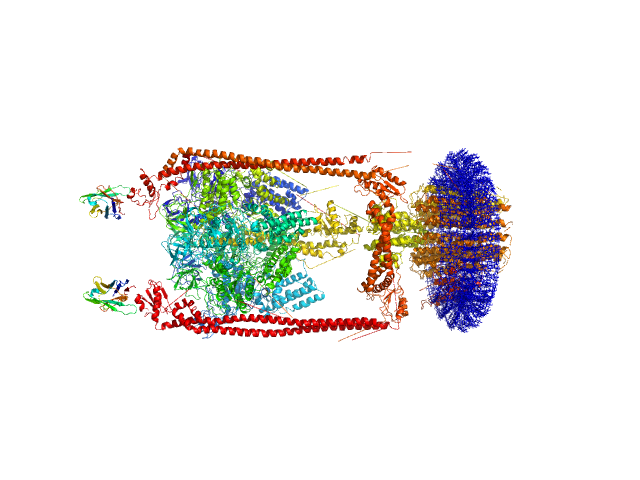
n-Dodecyl-β-D-Maltopyranoside , 77 kDa synthetic construct
A-type ATP synthase monomer, 655 kDa Thermus thermophilus protein
Monoclonal Antibody Fragment dimer, 33 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris/HCl, 100 mM sucrose, 100 mM NaCl, 2 mM MgCl2, 10% glycerol, 0.05% b-DDM, pH: 8
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2013 Mar 3
|
MPBuilder: A PyMOL Plugin for Building and Refinement of Solubilized Membrane Proteins Against Small Angle X-ray Scattering Data
Journal of Molecular Biology :166888 (2021)
Molodenskiy D, Svergun D, Mertens H
|
| RgGuinier |
8.0 |
nm |
| Dmax |
27.0 |
nm |
| VolumePorod |
1549 |
nm3 |
|
|