|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG52_fit1_model1.png)
|
| Sample: |
1,2-dimyristoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG62_fit1_model1.png)
|
| Sample: |
1,2-dimyristoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG72_fit1_model1.png)
|
| Sample: |
1,2-dimyristoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG82_fit1_model1.png)
|
| Sample: |
1,2-dimyristoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDG92_fit1_model1.png)
|
| Sample: |
1,2-dipalmitoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDGA2_fit1_model1.png)
|
| Sample: |
1,2-dipalmitoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDGB2_fit1_model1.png)
|
| Sample: |
1,2-dipalmitoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDGC2_fit1_model1.png)
|
| Sample: |
1,2-dipalmitoyl-sn-glycero-3-phosphocholine monomer, 1 kDa
|
| Buffer: |
water, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 9
|
Restoring structural parameters of lipid mixtures from small-angle X-ray scattering data
Journal of Applied Crystallography 54(1) (2021)
Konarev P, Gruzinov A, Mertens H, Svergun D
|
|
|
|
|
|
|
|
| Sample: |
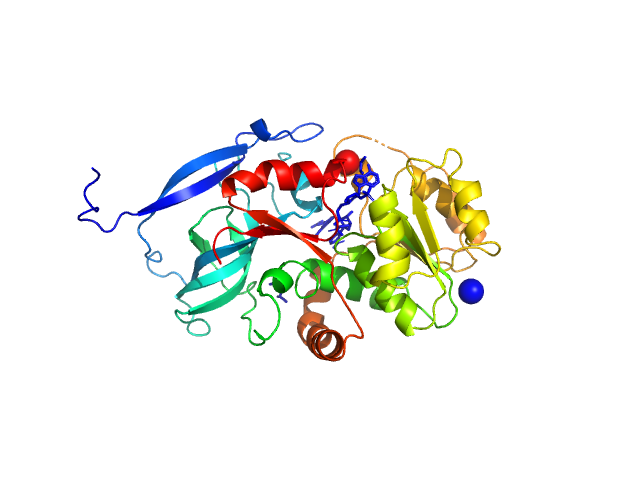
Malus domestica double bond reductase dimer, 77 kDa Malus domestica protein
|
| Buffer: |
50 mM Tris-HCl, 100 mM NaCl., pH: 7.5
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2020 Oct 23
|
The structural and functional characterization of Malus domestica double bond reductase MdDBR provides insights towards the identification of its substrates
International Journal of Biological Macromolecules 171:89-99 (2021)
Caliandro R, Polsinelli I, Demitri N, Musiani F, Martens S, Benini S
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.9 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
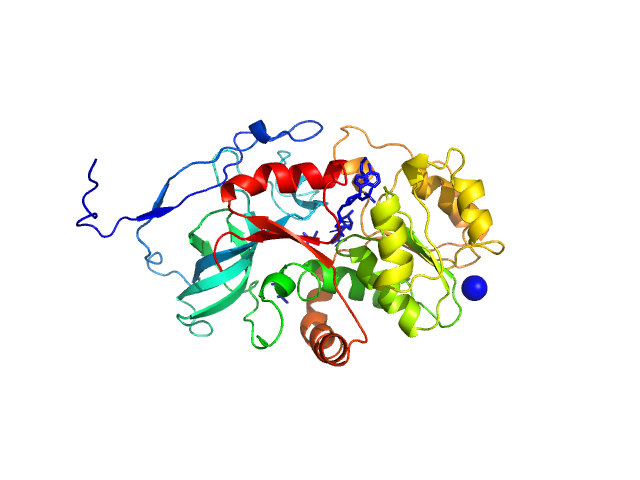
Malus domestica double bond reductase dimer, 77 kDa Malus domestica protein
|
| Buffer: |
50 mM Tris-HCl, 100 mM NaCl, 5 mM NADPH, pH: 7.5
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2020 Oct 23
|
The structural and functional characterization of Malus domestica double bond reductase MdDBR provides insights towards the identification of its substrates
International Journal of Biological Macromolecules 171:89-99 (2021)
Caliandro R, Polsinelli I, Demitri N, Musiani F, Martens S, Benini S
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
99 |
nm3 |
|
|