|
|
|
|

|
| Sample: |
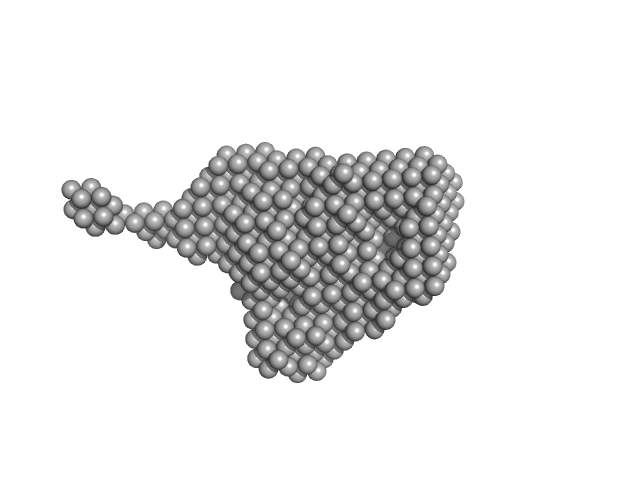
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
1.9 |
nm |
| Dmax |
8.1 |
nm |
|
|
|
|
|
|

|
| Sample: |
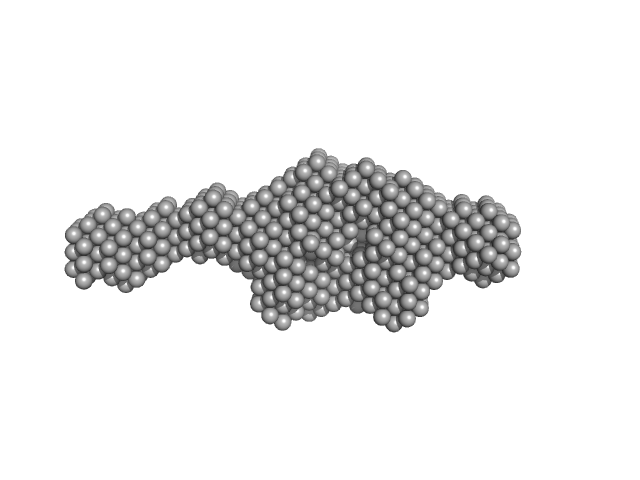
Aryl hydrocarbon receptor nuclear translocator-like protein 1 monomer, 10 kDa Mus musculus protein
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.5 |
nm |
|
|
|
|
|
|

|
| Sample: |
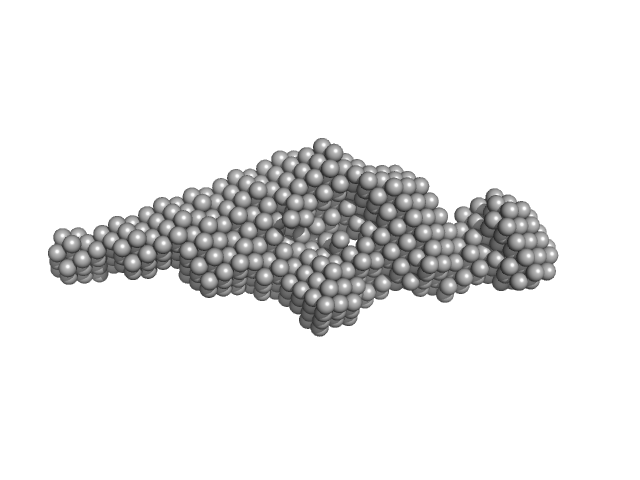
Brain and muscle ARNT-like 1 (G517-L625) including transactivation domain (TAD) monomer, 11 kDa Mus musculus protein
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP) monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
3.1 |
nm |
| Dmax |
13.7 |
nm |
|
|
|
|
|
|

|
| Sample: |
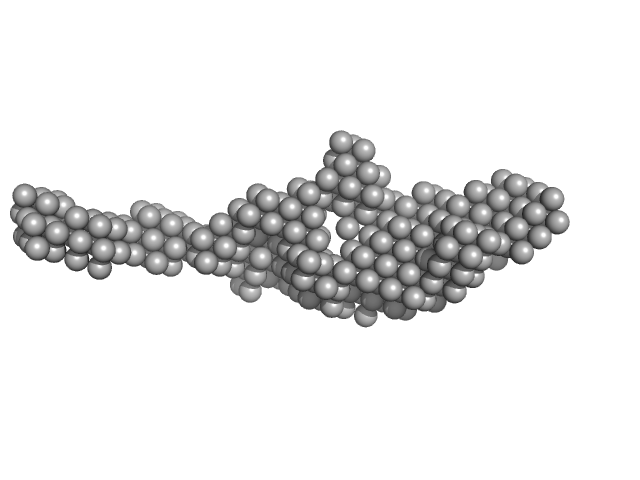
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP) monomer, 10 kDa Mus musculus protein
Brain and muscle ARNT-like 1 (G490-L625) including transactivation domain (TAD) monomer, 14 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
3.8 |
nm |
| Dmax |
17.5 |
nm |
|
|
|
|
|
|

|
| Sample: |
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
1-{4-[4-chloro-3-(trifluoromethyl)phenyl]-4-hydroxypiperidin-1-yl}-3-sulfanylpropan-1-one monomer, 0 kDa
|
| Buffer: |
25 mM Hepes, 150 NaCl, 5% Glycerol, pH: 7.2
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
1.7 |
nm |
| Dmax |
6.4 |
nm |
|
|
|
|
|
|
|
| Sample: |
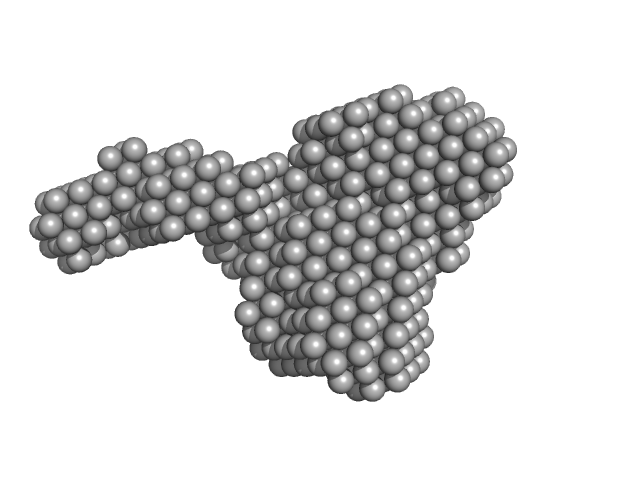
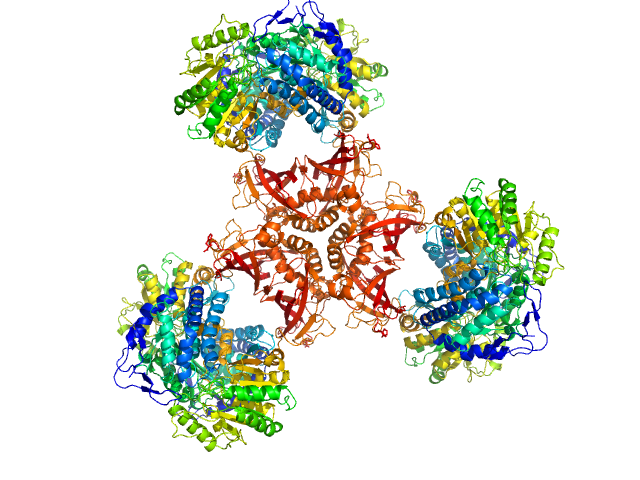
Bifunctional protein PaaZ hexamer, 438 kDa Escherichia coli protein
|
| Buffer: |
25 mM HEPES, 50 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Feb 24
|
Molecular basis for metabolite channeling in a ring opening enzyme of the phenylacetate degradation pathway.
Nat Commun 10(1):4127 (2019)
Sathyanarayanan N, Cannone G, Gakhar L, Katagihallimath N, Sowdhamini R, Ramaswamy S, Vinothkumar KR
|
| RgGuinier |
6.2 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
636 |
nm3 |
|
|
|
|
|
|

|
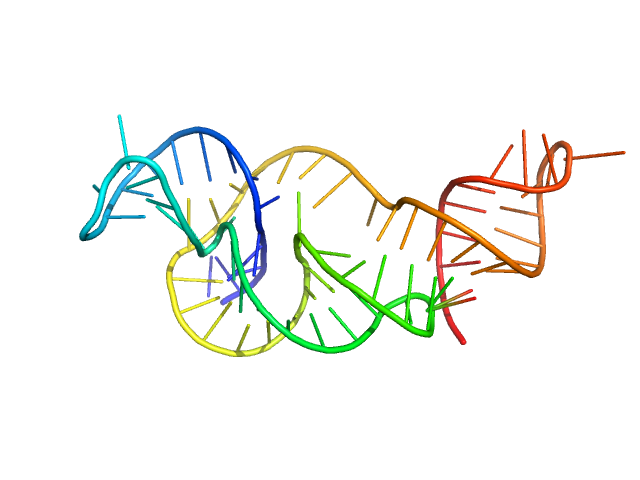
| Sample: |
Xrn1 resistance RNA2 from Zika virus monomer, 22 kDa Zika virus RNA
|
| Buffer: |
20mM Tris-HCl, 100mM NaCl, 5mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Dec 9
|
Long non-coding subgenomic flavivirus RNAs have extended 3D structures and are flexible in solution.
EMBO Rep 20(11):e47016 (2019)
Zhang Y, Zhang Y, Liu ZY, Cheng ML, Ma J, Wang Y, Qin CF, Fang X
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
34 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
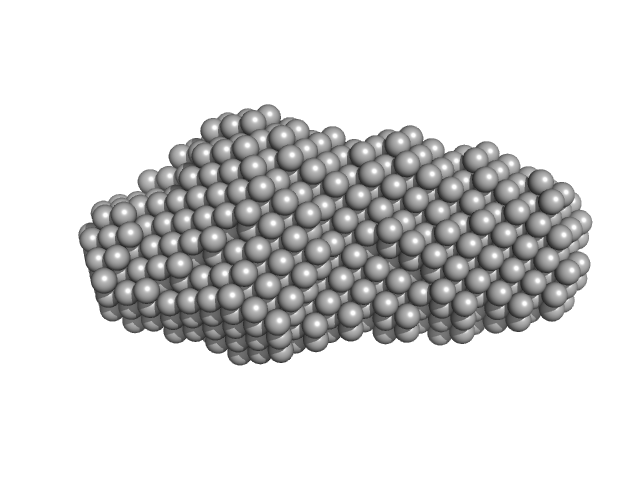
Xrn1 resistance RNA1 from Dengue virus 2 monomer, 21 kDa Dengue virus 2 RNA
|
| Buffer: |
20mM Tris-HCl, 100mM NaCl, 5mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 14-ID-B (BioCARS), Advanced Photon Source (APS), Argonne National Laboratory on 2016 Dec 9
|
Long non-coding subgenomic flavivirus RNAs have extended 3D structures and are flexible in solution.
EMBO Rep 20(11):e47016 (2019)
Zhang Y, Zhang Y, Liu ZY, Cheng ML, Ma J, Wang Y, Qin CF, Fang X
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
31 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
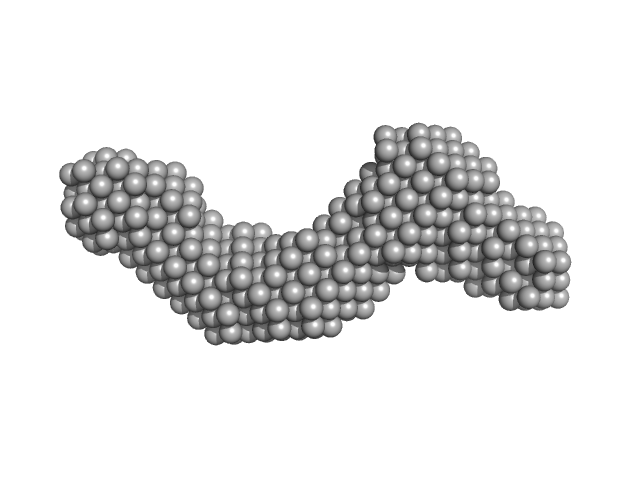
Xrn1 resistance RNA1-2 from Dengue virus 2 monomer, 46 kDa Dengue virus 2 RNA
|
| Buffer: |
20mM Tris-HCl, 100mM NaCl, 5mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2017 Apr 2
|
Long non-coding subgenomic flavivirus RNAs have extended 3D structures and are flexible in solution.
EMBO Rep 20(11):e47016 (2019)
Zhang Y, Zhang Y, Liu ZY, Cheng ML, Ma J, Wang Y, Qin CF, Fang X
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
67 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
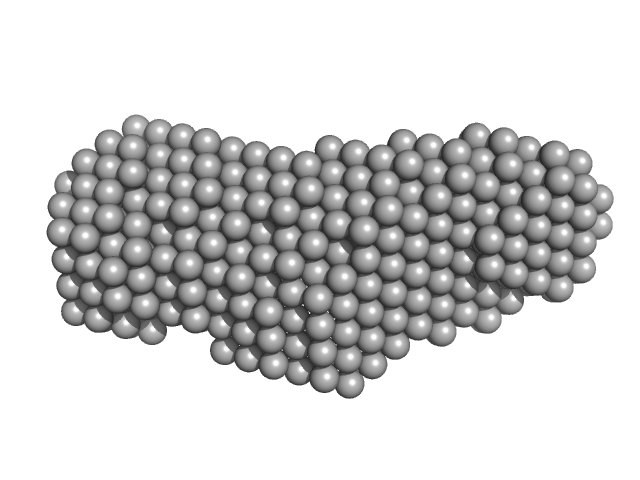
Xrn1 resistance RNA-1 from West Nile virus monomer, 25 kDa West Nile virus RNA
|
| Buffer: |
20mM Tris-HCl, 100mM NaCl, 5mM MgCl2, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Dec 9
|
Long non-coding subgenomic flavivirus RNAs have extended 3D structures and are flexible in solution.
EMBO Rep 20(11):e47016 (2019)
Zhang Y, Zhang Y, Liu ZY, Cheng ML, Ma J, Wang Y, Qin CF, Fang X
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
34 |
nm3 |
|
|