|
|
|
|
|
| Sample: |
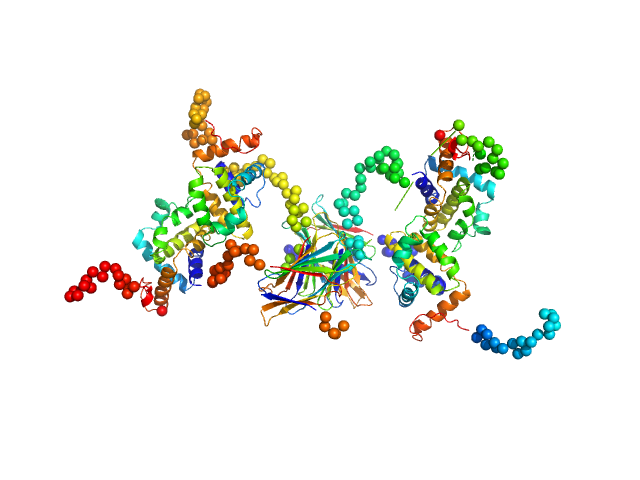
Protein A46 tetramer, 112 kDa Vaccinia virus protein
|
| Buffer: |
20 mM Tris-HCl, 10 mM DTT, pH: 8.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jun 25
|
Vaccinia Virus Immunomodulator A46: A Lipid and Protein-Binding Scaffold for Sequestering Host TIR-Domain Proteins.
PLoS Pathog 12(12):e1006079 (2016)
Fedosyuk S, Bezerra GA, Radakovics K, Smith TK, Sammito M, Bobik N, Round A, Ten Eyck LF, Djinović-Carugo K, Usón I, Skern T
|
| RgGuinier |
4.3 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
199 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
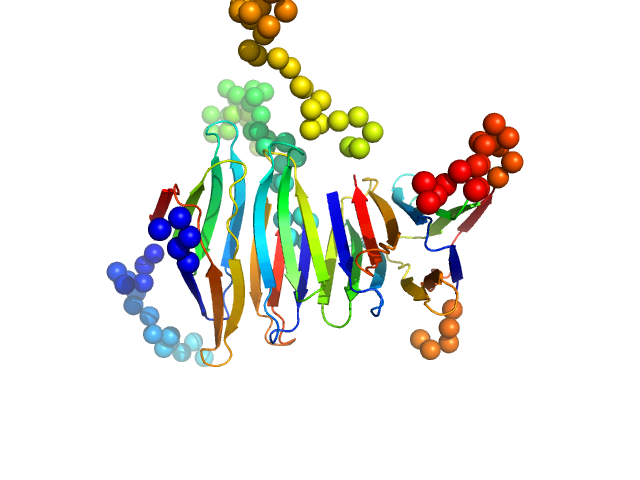
Protein A46 tetramer, 40 kDa Vaccinia virus protein
|
| Buffer: |
20 mM Tris-HCl, 10 mM DTT, pH: 8.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 Jun 25
|
Vaccinia Virus Immunomodulator A46: A Lipid and Protein-Binding Scaffold for Sequestering Host TIR-Domain Proteins.
PLoS Pathog 12(12):e1006079 (2016)
Fedosyuk S, Bezerra GA, Radakovics K, Smith TK, Sammito M, Bobik N, Round A, Ten Eyck LF, Djinović-Carugo K, Usón I, Skern T
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
68 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
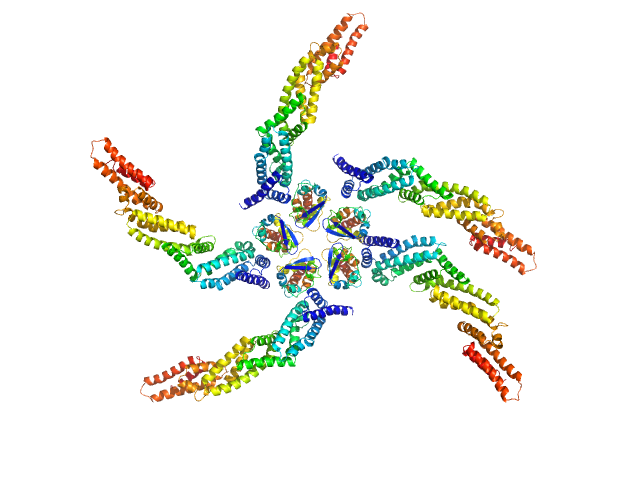
Cullin 3 pentamer, 60 kDa Drosophila melanogaster protein
Insomniac protein pentamer, 210 kDa Drosophila melanogaster protein
|
| Buffer: |
50 mM TrisHCl, 300 mM NaCl, 2 mM DTT, pH: 8
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2014 Nov 24
|
Proteins involved in sleep homeostasis: Biophysical characterization of INC and its partners
Biochimie 131:106-114 (2016)
Pirone L, Smaldone G, Esposito C, Balasco N, Petoukhov M, Spilotros A, Svergun D, Di Gaetano S, Vitagliano L, Pedone E
|
| RgGuinier |
6.7 |
nm |
| Dmax |
25.2 |
nm |
|
|
|
|
|
|
|
| Sample: |
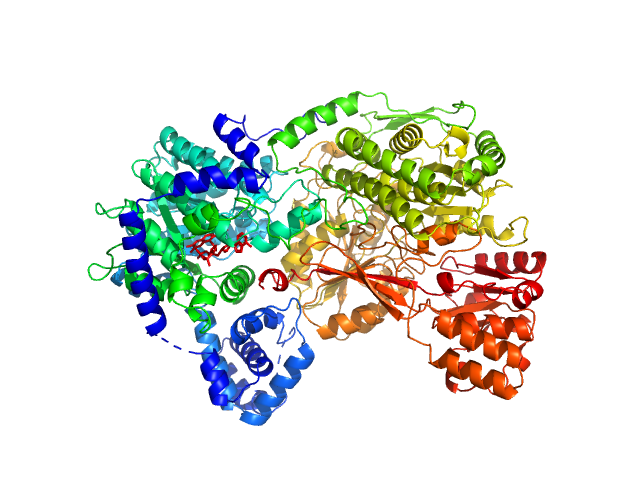
Sinorhizobium meliloti (SmPutA) monomer, 132 kDa Sinorhizobium meliloti protein
|
| Buffer: |
50 mM Tris, 1% (v/v) glycerol, 0.5 mM THP, and 50 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Mar 27
|
Structures of Proline Utilization A (PutA) Reveal the Fold and Functions of the Aldehyde Dehydrogenase Superfamily Domain of Unknown Function.
J Biol Chem 291(46):24065-24075 (2016)
Luo M, Gamage TT, Arentson BW, Schlasner KN, Becker DF, Tanner JJ
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
171 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
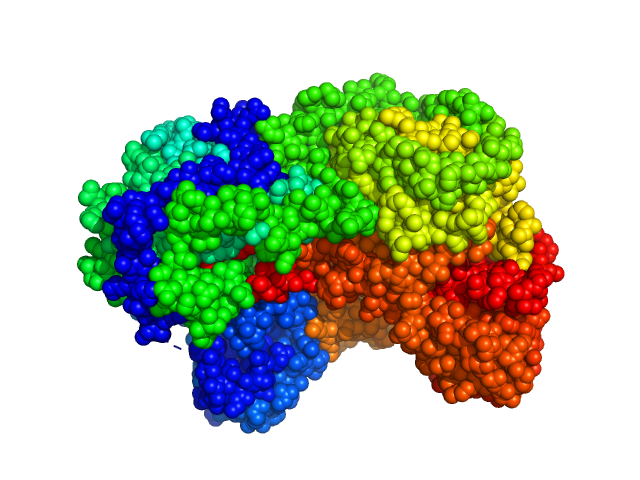
Sinorhizobium meliloti (SmPutA) monomer, 132 kDa Sinorhizobium meliloti protein
|
| Buffer: |
50 mM Tris, 1% (v/v) glycerol, 0.5 mM THP, and 50 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Mar 27
|
Structures of Proline Utilization A (PutA) Reveal the Fold and Functions of the Aldehyde Dehydrogenase Superfamily Domain of Unknown Function.
J Biol Chem 291(46):24065-24075 (2016)
Luo M, Gamage TT, Arentson BW, Schlasner KN, Becker DF, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
11.9 |
nm |
| VolumePorod |
225 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
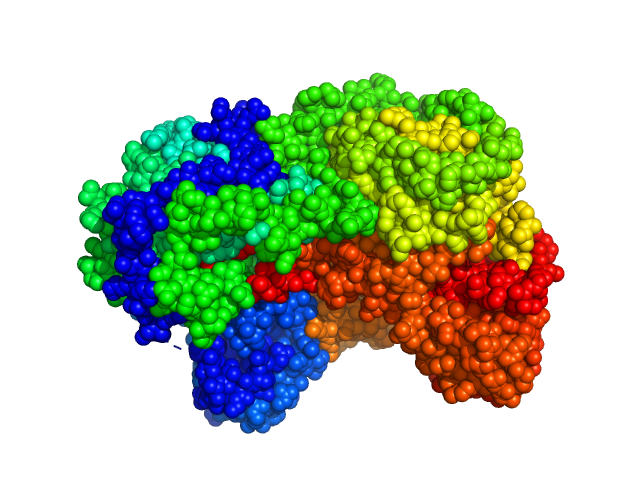
Sinorhizobium meliloti (SmPutA) monomer, 132 kDa Sinorhizobium meliloti protein
|
| Buffer: |
50 mM Tris, 1% (v/v) glycerol, 0.5 mM THP, and 50 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Mar 27
|
Structures of Proline Utilization A (PutA) Reveal the Fold and Functions of the Aldehyde Dehydrogenase Superfamily Domain of Unknown Function.
J Biol Chem 291(46):24065-24075 (2016)
Luo M, Gamage TT, Arentson BW, Schlasner KN, Becker DF, Tanner JJ
|
| RgGuinier |
3.8 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
248 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
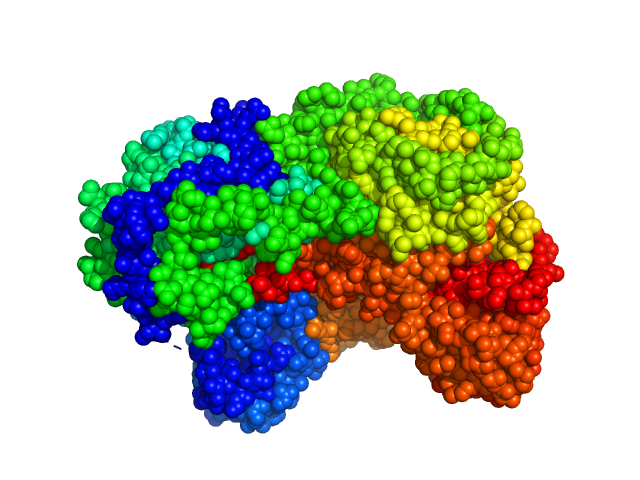
Sinorhizobium meliloti (SmPutA) monomer, 132 kDa Sinorhizobium meliloti protein
|
| Buffer: |
50 mM Tris, 1% (v/v) glycerol, 0.5 mM THP, and 50 mM NaCl, pH: 7.8
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2014 Mar 27
|
Structures of Proline Utilization A (PutA) Reveal the Fold and Functions of the Aldehyde Dehydrogenase Superfamily Domain of Unknown Function.
J Biol Chem 291(46):24065-24075 (2016)
Luo M, Gamage TT, Arentson BW, Schlasner KN, Becker DF, Tanner JJ
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.9 |
nm |
| VolumePorod |
277 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
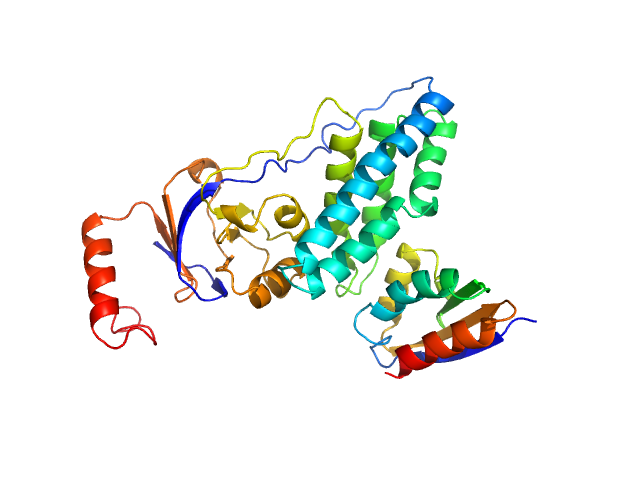
Phosphocarrier protein NPr monomer, 9 kDa Escherichia coli protein
Phosphoenolpyruvate-protein phosphotransferase PtsP monomer, 28 kDa Escherichia coli protein
|
| Buffer: |
10 mM Tris 150 mM NaCl 0.5 mM EDTA, pH: 7.5
|
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-1000, NIDDK, NIH on 2015 Jan 30
|
Structure of the NPr:EINNtr Complex: Mechanism for Specificity in Paralogous Phosphotransferase Systems.
Structure 24(12):2127-2137 (2016)
Strickland M, Stanley AM, Wang G, Botos I, Schwieters CD, Buchanan SK, Peterkofsky A, Tjandra N
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
|
|
|
|
|
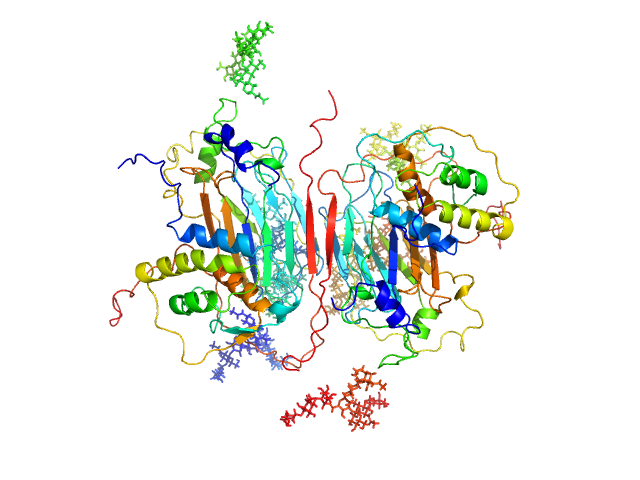
| Sample: |
GM-CSF/IL-2 inhibition factor tetramer, 120 kDa Orf virus protein
Granulocyte-macrophage colony-stimulating factor dimer, 29 kDa Ovis aries protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2014 Sep 10
|
Structural basis of GM-CSF and IL-2 sequestration by the viral decoy receptor GIF.
Nat Commun 7:13228 (2016)
Felix J, Kandiah E, De Munck S, Bloch Y, van Zundert GC, Pauwels K, Dansercoer A, Novanska K, Read RJ, Bonvin AM, Vergauwen B, Verstraete K, Gutsche I, Savvides SN
|
| RgGuinier |
3.8 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
231 |
nm3 |
|
|
|
|
|
|
|
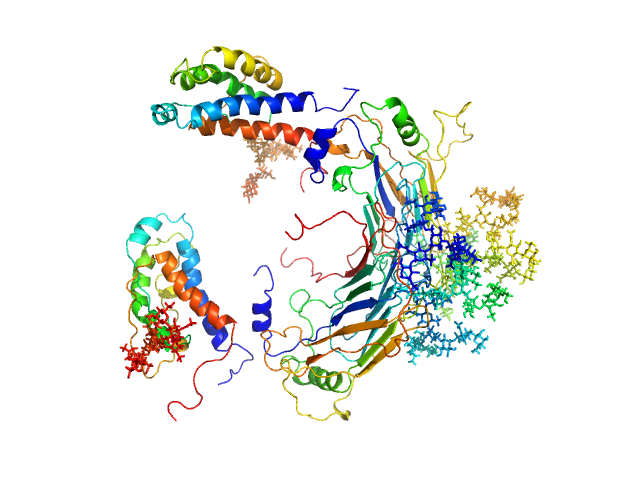
| Sample: |
GM-CSF/IL-2 inhibition factor tetramer, 120 kDa Orf virus protein
Interleukin-2 monomer, 16 kDa Ovis aries protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.4
|
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2014 Sep 10
|
Structural basis of GM-CSF and IL-2 sequestration by the viral decoy receptor GIF.
Nat Commun 7:13228 (2016)
Felix J, Kandiah E, De Munck S, Bloch Y, van Zundert GC, Pauwels K, Dansercoer A, Novanska K, Read RJ, Bonvin AM, Vergauwen B, Verstraete K, Gutsche I, Savvides SN
|
| RgGuinier |
4.1 |
nm |
| Dmax |
12.9 |
nm |
| VolumePorod |
253 |
nm3 |
|
|