|
|
|
|
|
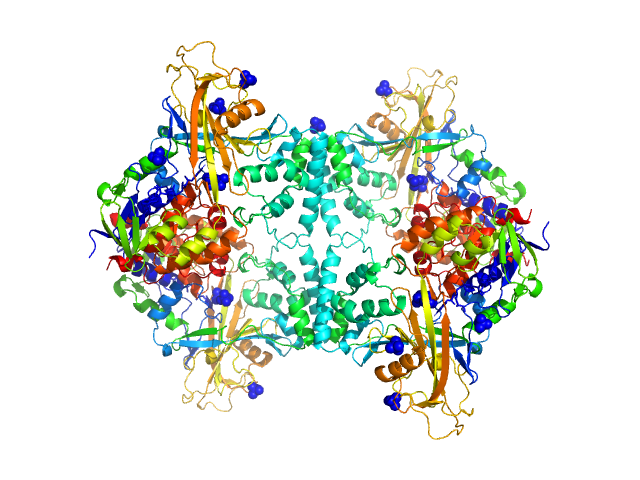
| Sample: |
Aspergillus fumigatus UDP galactopyranose mutase tetramer, 228 kDa protein
|
| Buffer: |
20 mM HEPES, 45 mM NaCl, 0.5 mM Tris(hydroxypropyl)phosphine, pH: 7.5
|
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Apr 19
|
Crystal structures and small-angle x-ray scattering analysis of UDP-galactopyranose mutase from the pathogenic fungus Aspergillus fumigatus.
J Biol Chem 287(12):9041-51 (2012)
Dhatwalia R, Singh H, Oppenheimer M, Karr DB, Nix JC, Sobrado P, Tanner JJ
|
| RgGuinier |
4.7 |
nm |
| Dmax |
14.7 |
nm |
| VolumePorod |
308 |
nm3 |
|
|
|
|
|
|
|
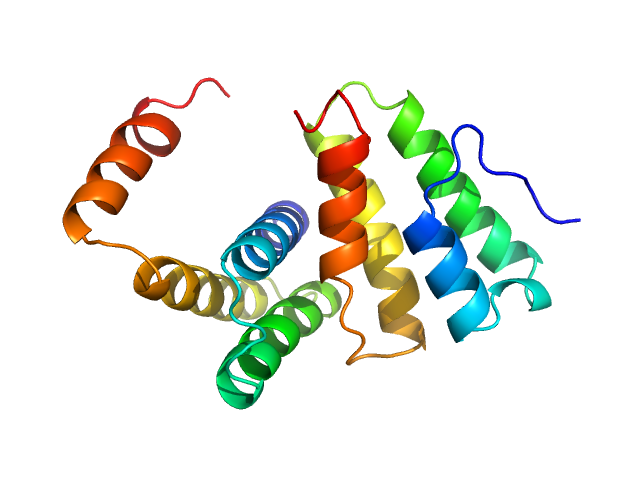
| Sample: |
Replicase polyprotein 1a dimer, 19 kDa Severe acute respiratory … protein
|
| Buffer: |
10 mM Tris-HCl, 200 mM NaCl, 5 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Apr 2
|
Nonstructural Proteins 7 and 8 of Feline Coronavirus Form a 2:1 Heterotrimer That Exhibits Primer-Independent RNA Polymerase Activity
Journal of Virology 86(8):4444-4454 (2012)
Xiao Y, Ma Q, Restle T, Shang W, Svergun D, Ponnusamy R, Sczakiel G, Hilgenfeld R
|
|
|
|
|
|
|
|
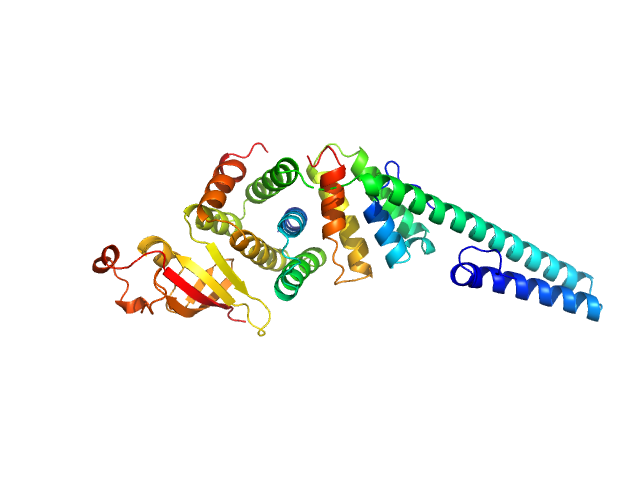
| Sample: |
Replicase polyprotein 1a dimer, 19 kDa Severe acute respiratory … protein
Replicase polyprotein 1a monomer, 22 kDa Severe acute respiratory … protein
|
| Buffer: |
10 mM Tris-HCl, 200 mM NaCl, 5 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Apr 2
|
Nonstructural Proteins 7 and 8 of Feline Coronavirus Form a 2:1 Heterotrimer That Exhibits Primer-Independent RNA Polymerase Activity
Journal of Virology 86(8):4444-4454 (2012)
Xiao Y, Ma Q, Restle T, Shang W, Svergun D, Ponnusamy R, Sczakiel G, Hilgenfeld R
|
|
|
|
|
|
|
|
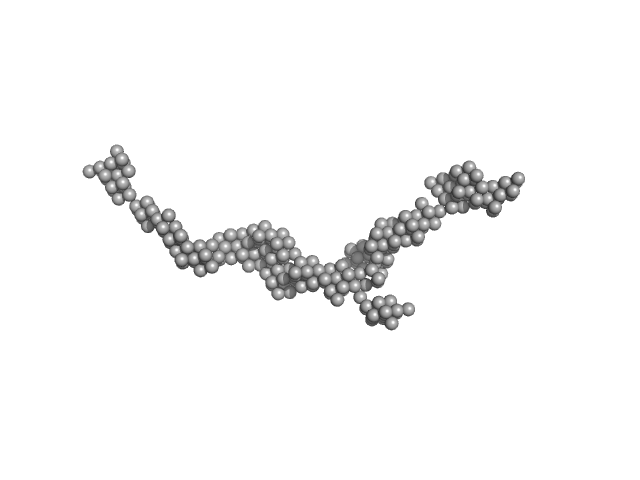
| Sample: |
Myomesin-1 dimer, 121 kDa Homo sapiens protein
|
| Buffer: |
25 mMTris-HCl, 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2005 May 10
|
Superhelical architecture of the myosin filament-linking protein myomesin with unusual elastic properties.
PLoS Biol 10(2):e1001261 (2012)
Pinotsis N, Chatziefthimiou SD, Berkemeier F, Beuron F, Mavridis IM, Konarev PV, Svergun DI, Morris E, Rief M, Wilmanns M
|
| RgGuinier |
9.6 |
nm |
| Dmax |
37.0 |
nm |
|
|
|
|
|
|
|
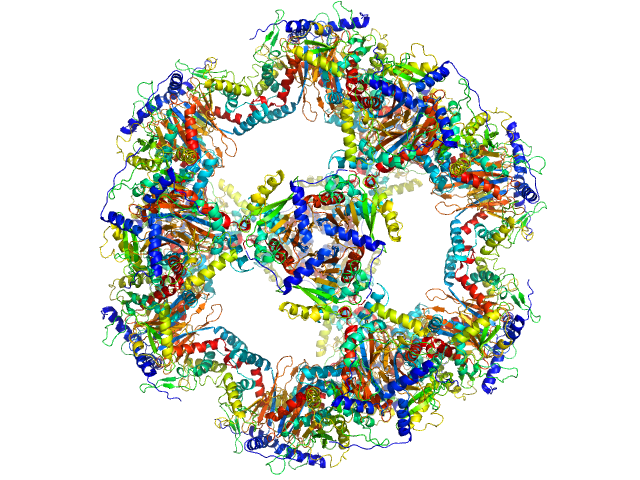
| Sample: |
Regulatory protein E2 , 1037 kDa Human papillomavirus type … protein
|
| Buffer: |
50 mM Tris ⁄ HCl, pH 8.8, 100 mM NaCl, pH: 8.8
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2009 Jul 13
|
The catalytic core of an archaeal 2-oxoacid dehydrogenase multienzyme complex is a 42-mer protein assembly
FEBS Journal 279(5):713-723 (2012)
Marrott N, Marshall J, Svergun D, Crennell S, Hough D, Danson M, van den Elsen J
|
| RgGuinier |
8.8 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
2473 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
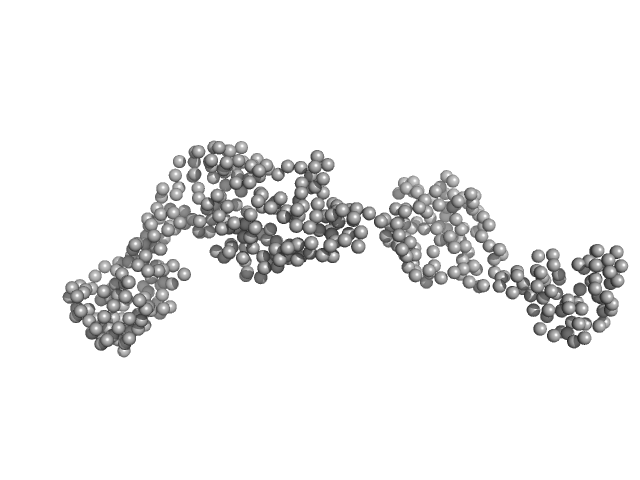
Full length GbpA monomer, 54 kDa Vibrio cholerae protein
|
| Buffer: |
25 mM Tris/HCl 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Oct 16
|
The Vibrio cholerae colonization factor GbpA possesses a modular structure that governs binding to different host surfaces.
PLoS Pathog 8(1):e1002373 (2012)
Wong E, Vaaje-Kolstad G, Ghosh A, Hurtado-Guerrero R, Konarev PV, Ibrahim AF, Svergun DI, Eijsink VG, Chatterjee NS, van Aalten DM
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
|
|
|
|
|
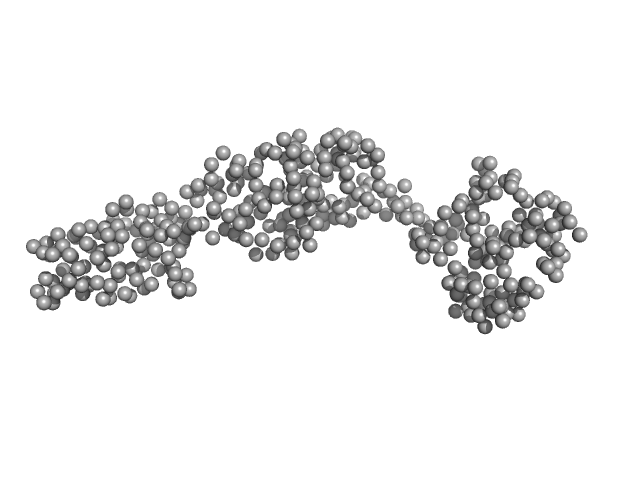
| Sample: |
Truncated GbpA monomer, 44 kDa Vibrio cholerae protein
|
| Buffer: |
25 mM Tris/HCl 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2007 Oct 16
|
The Vibrio cholerae colonization factor GbpA possesses a modular structure that governs binding to different host surfaces.
PLoS Pathog 8(1):e1002373 (2012)
Wong E, Vaaje-Kolstad G, Ghosh A, Hurtado-Guerrero R, Konarev PV, Ibrahim AF, Svergun DI, Eijsink VG, Chatterjee NS, van Aalten DM
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
|
|
|
|
|
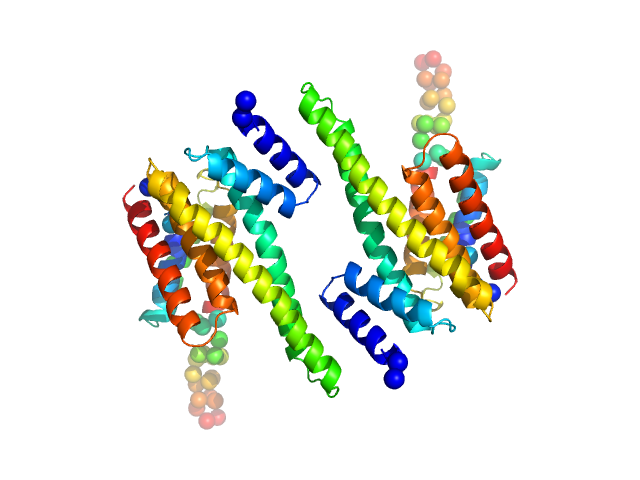
| Sample: |
14-3-3 protein beta/alpha dimer, 58 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2 mM 2-mercaptoethanol, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2011 Apr 7
|
The weak complex between RhoGAP protein ARHGAP22 and signal regulatory protein 14-3-3 has 1:2 stoichiometry and a single peptide binding mode.
PLoS One 7(8):e41731 (2012)
Hu SH, Whitten AE, King GJ, Jones A, Rowland AF, James DE, Martin JL
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
|
|
|
|
|
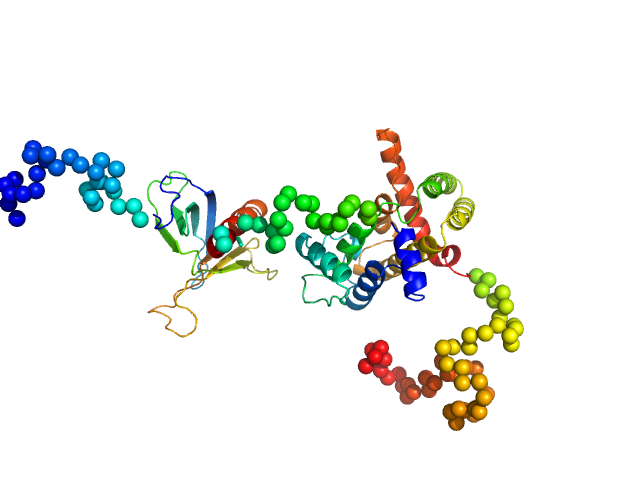
| Sample: |
Rho GTPase-activating protein 22 monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2 mM 2-mercaptoethanol, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2011 Apr 7
|
The weak complex between RhoGAP protein ARHGAP22 and signal regulatory protein 14-3-3 has 1:2 stoichiometry and a single peptide binding mode.
PLoS One 7(8):e41731 (2012)
Hu SH, Whitten AE, King GJ, Jones A, Rowland AF, James DE, Martin JL
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
88 |
nm3 |
|
|
|
|
|
|
|
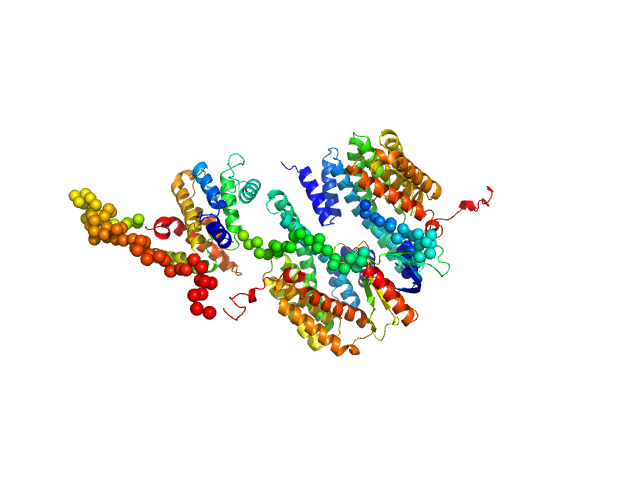
| Sample: |
14-3-3 protein beta/alpha dimer, 58 kDa Homo sapiens protein
Rho GTPase-activating protein 22 monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2 mM 2-mercaptoethanol, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2011 Apr 7
|
The weak complex between RhoGAP protein ARHGAP22 and signal regulatory protein 14-3-3 has 1:2 stoichiometry and a single peptide binding mode.
PLoS One 7(8):e41731 (2012)
Hu SH, Whitten AE, King GJ, Jones A, Rowland AF, James DE, Martin JL
|
| RgGuinier |
3.8 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
195 |
nm3 |
|
|