|
|
|
|
|
| Sample: |
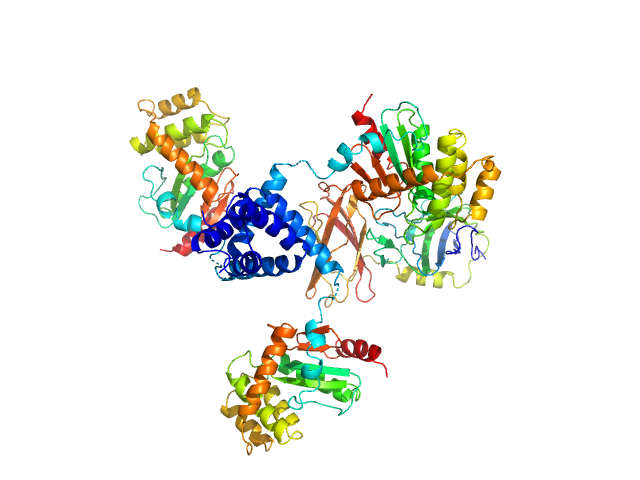
...protein trimer, 74 kDa Proteus mirabilis protein
...protein monomer, 30 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 Nov 2
|
...protein is primed by the tandem immunoglobulin-fold domain of ScsB.
J Biol Chem 293(16):5793-5805 (2018)
Furlong EJ, Choudhury HG, Kurth F, Duff AP, Whitten AE, Martin JL
|
| RgGuinier |
3.9 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
145 |
nm3 |
|